Key Points
The αIIbβ3 headpiece points away from the lipid bilayer, and the lower legs are either bent (αIIb) or freely coiled (β3).
The linking region between the ecto- and TM domains likely transmits the TM conformational changes associated with inside-out activation.
Abstract
Integrin αIIbβ3 plays a central role in hemostasis and thrombosis. We provide the first 3-dimensional reconstruction of intact purified αIIbβ3 in a nanodisc lipid bilayer. Unlike previous models, it shows that the ligand-binding head domain is on top, pointing away from the membrane. Moreover, unlike the crystal structure of the recombinant ectodomain, the lower legs are not parallel, straight, and adjacent. Rather, the αIIb lower leg is bent between the calf-1 and calf-2 domains and the β3 Integrin-Epidermal Growth Factor (I-EGF) 2 to 4 domains are freely coiled rather than in a cleft between the β3 headpiece and the αIIb lower leg. Our data indicate an important role for the region that links the distal calf-2 and β-tail domains to their respective transmembrane (TM) domains in transmitting the conformational changes in the TM domains associated with inside-out activation.
Introduction
Integrin receptors play important roles in a wide variety of physiological and pathological processes, including development, cell survival, immunity, malignancy, metastasis, and thrombosis.1 The platelet αIIbβ3 receptor is required for normal hemostasis and plays a permissive role in thrombosis; the latter has made it a validated target for preventing and treating heart attacks.2,3 Based on crystallographic studies of recombinant αIIbβ3 and αVβ3 ectodomains,4,5 as well as electron microscopic studies of the recombinant ectodomains,5-7 there is a broad consensus that unactivated αIIbβ3 has a bent conformation with straight, parallel, and adjacent lower leg domains. Uncertainties remain, however, about the orientation of the αIIbβ3 ectodomain relative to the transmembrane (TM) domain in full-length αIIbβ3 in a lipid bilayer and the conformations of the leg domains. In fact, studies of full-length αIIbβ3 in detergent using transmission cryo–electron microscopy (cryo-EM), small-angle neutron scattering (SANS), and small angle x-ray scattering (SAXS) have indicated different ectodomain orientations, with the cryo-EM data interpreted as showing the head region oriented away from the membrane, and the SANS and SAXS data interpreted as showing the head region oriented toward the membrane.8-10 Similarly, the cryo-EM data suggested that the lower leg domains were bent, whereas the SANS and SAXS studies were interpreted as consistent with the straight, parallel, and adjacent lower leg domain conformations observed in the crystal structure. In fact, it has been difficult to fit the crystallographic structure of the receptor with the data from cryo-EM, SAXS, or SANS.
Materials and methods
Materials
n-octyl-β-d-glucoside (OG), 1,2-dimyristoyl-sn-glycero-3-phosphocholine (DMPC), and 1,2-dimyristoyl-sn-glycero-3-phospho-(1′-rac-glycerol) (DMPG) were purchased from Anatrace. Prostaglandin E1 (PGE1), isopropyl β-D-1-thiogalactopyranoside, α-methyl glucoside, leupeptin, uranyl acetate, and Triton X-100 were from Sigma. Con A–Sepharose, Q-Sepharose, Nickel-Sepharose, Sephacryl S300 HR, Superdex 200, and N-hydroxysuccinimide (NHS)-activated Sepharose were from GE Healthcare. Bio-Bead SM-2 was from Bio-Rad. PAC-1 was from BD Biosciences. Four hundred mesh carbon-coated copper grids were from Electron Microscopy Sciences.
Purification of integrin αIIbβ3
We applied and modified the methods initially established by Ye et al.11 αIIbβ3 was purified from outdated single-donor platelet concentrates obtained from the New York Blood Center by washing platelets in the presence of PGE1, removing contaminating blood cells by centrifugation, lysing the resuspended platelets at 4°C in 5% (weight/volume [wt/vol]) OG in 150mM NaCl, 20mM N-2-hydroxyethylpiperazine-N′-2-ethanesulfonic acid (HEPES), 1mM CaCl2, 1mM MgCl2, 10μM leupeptin, pH 7.4; performing conconavalin A affinity chromatography (binding buffer: 150mM NaCl, 1% [wt/vol] OG, 20mM HEPES, 1mM CaCl2, 1mM MgCl2 and pH 7.4; washing buffer: binding buffer + 20mM α-methyl glucoside; elution buffer: binding buffer + 1M α-methyl glucoside); performing heparin affinity chromatography; applying the flow through fraction to Q-Sepharose (binding buffer: 75mM NaCl, 1% [wt/vol] OG, 10mM HEPES, 1mM CaCl2, 1mM MgCl2, pH 7.4; washing buffer: binding buffer + 200mM NaCl; elution buffer: binding buffer + 400mM NaCl); and performing gel-size exclusion chromatography on Sephacryl S300 HR (running buffer: 150mM NaCl, 1% (wt/vol) OG, 10mM HEPES, 1mM CaCl2, 1mM MgCl2, pH 7.4).
Preparation of αIIbβ3 nanodiscs
The MSP1D1 membrane scaffolding protein expression plasmid was purchased from Addgene and the protein was expressed and purified as described by Ritchie et al.12 The nanodisc lipid mixture (DMPC and DMPG) was prepared following the method of Ye et al,11 and solubilized in buffer (300mM NaCl, 100mM OG, 10mM cholate, 10mM HEPES, pH 7.4) to a 50mM final concentration of each lipid (molar ratio octyl glucoside:lipid = 2:1). The lipid solution and the purified membrane scaffold protein (1.6 mg/mL) were mixed and then purified αIIbβ3 (1.3 mg/mL) was added. The final molar ratio of lipids to MSP1D1 to αIIbβ3 was 100:1:0.05. The integrin nanodiscs were assembled by removing the sodium cholate and OG for 9 hours at room temperature with 0.3 g/mL macroporous polymeric beads of high surface area for adsorbing organics from aqueous solutions (Bio-Bead SM-2). Nanodiscs without and with αIIbβ3 were separated by gel filtration chromatography on Superdex 200 (running buffer: 150mM NaCl, 10mM HEPES, pH 7.4, 1mM CaCl2, and 1mM MgCl2). The monoclonal antibody (mAb) PAC-1 (200 μg), which reacts selectively with activated αIIbβ3,13 was immobilized on 1 mL of NHS-activated Sepharose according to the manufacturer’s instruction. The purified αIIbβ3 nanodiscs were chromatographed on a column containing the PAC-1 Sepharose to remove nanodiscs containing activated αIIbβ3. The flowthrough sample was used for the EM experiments.
Preparation and purification of mAb-bound αIIbβ3 nanodisc
αIIbβ3 nanodiscs (50 μg) were reacted with 0.1 mg of mAb 10E5, 7E3, or 7H2 for 1 hour at room temperature. Antibody molecules that did not bind to the αIIbβ3 nanodiscs were removed by size-exclusion chromatography on Superdex 200 (running buffer: 150mM NaCl, 10mM HEPES, pH 7.4, containing 1mM CaCl2 and 1mM MgCl2).
EM imaging
αIIbβ3 nanodiscs were loaded onto glow-discharged carbon-coated copper grids and stained with 2% uranyl acetate. Imaging of antibody-bound αIIbβ3 nanodiscs was performed using a JEOL JEM 100CX transmission electron microscope at 80 kV and magnifications of ×33 000 and ×50 000. EM images of αIIbβ3 nanodiscs in the absence of antibody were obtained on a Tecnai F20 transmission electron microscope (FEI Company) operating at either 120 kV or 200 kV either with or without tilting the specimen to 50°. The 50° (tilted) samples were acquired first, followed by an overlapping montage of 0° (untilted) images. The micrographs were recorded on a 4k × 4k CCD camera (Tietz; TVIPS GmbH) at ×29 000 or ×50 000 magnification, with a pixel size of 2.96 Å or 3.4 Å.
Image processing
Image processing was performed using the EMAN1.9 and SPIDER packages. Pairs of tilted and untilted views were identified in the micrographs and the tilt axis orientation (ψ) and tilt angle (θ) were determined. Untilted particles were subjected to reference-free alignment, and the aligned images were classified by principal component analysis and hierarchical clustering. Euler angles were assigned to each tilted particle by combining the tilt axis orientation and tilt angle with the in-plane rotation (Φ) of the corresponding untilted particle determined during classification and alignment. Random conical tilt (RCT) 3-dimensional (3D) reconstructions were performed on each of the final classes of tilted images. The average map of the RCT reconstructions was generated by aligning the RCT reconstructions according to the integrin location and weighted averaging for the size of each class. The refinement of Euler angles and image shifts was achieved by back projection of a reference volume using the assigned angles and alignment of each particle to the corresponding projection of the volume. The 50° tilt pair angle was calculated from aligned Euler angles of tilted and untilted particles by the following equation (1): Euler angles of untilted particles (ψ, θ, Φ) = Euler angles of tilted particles (ψ + tilt axis, θ + tilt angle, Φ) (1). The Chimera program from the University of California at San Francisco was used to visualize and align the maps and the crystal structure. The resolution of each volume was calculated from the Fourier shell correlation (FSC) between 2 different volumes, each of which was generated from one-half of the data set. The resolution was calculated from the spatial frequency at which the FSC equals 0.5.
Docking of αIIbβ3 crystal structure into the reconstruction
Correlation-based docking analysis was used to identify the αIIb and β3 domains in the 20.5-Å resolution EM map of αIIbβ3 nanodisc.14-16 First, the crystal structure of the αIIbβ3 ectodomain (Protein Data Bank [PDB]:3FCS) was separated into individual domains and the flexible linkers between the domains (supplemental Table 2, available on the Blood Web site), and then the protein component of each domain was converted into a 20-Å resolution EM map. Simultaneously, an anchor graph of the 20.5-Å resolution EM map of αIIbβ3 in the nanodisc was created using the MultiFit program of Chimera to discretize the map into regions and check the connectivity between them.16 The initial placement of domains into the map was guided by the results of the mAb binding studies in locating the αIIb cap and β3 βI and Plexin-Semaphorin-Integrin (PSI) domains, utilizing the anchor graph and considering the sequence of domains in each subunit and the distances between connected domains in the crystal structure. To minimize clashes between docked domains, clashes and contacts were continuously checked during fitting using Chimera’s “Find Clashes/Contacts” tool with the following settings: Van der Waals (VDW) overlap ≥2 Å; subtract 0.4 from overlap for potentially H-bonding pairs, and ignore contact pairs that are 2 or fewer bonds apart. Refinement of the placement of the initially docked domains was achieved by: (1) adding the carbohydrate components of the αIIbβ3 crystal structure to the protein domains in the appropriate locations, and (2) optimizing the cross-correlation and the atom inclusion of each domain in the map using Chimera’s “Fit in Map” tool, with the orientations of domains constrained by the size of the flexible loops connecting them. The cross-correlations, clashes, and orientations of individual domains in the EM map relative to the crystal structure are provided in supplemental Table 2.
Results
To study the structure of intact αIIbβ3 in a lipid bilayer and the orientation of the ectodomain relative to the membrane, we purified αIIbβ3 from platelets using OG and embedded it in a nanodisc lipid bilayer (supplemental Figure 1).11,12 Activated receptors were removed by affinity chromatography using the activation-specific mAb PAC1 (supplemental Table 1).13
Domain mapping of 2D EM images using mAbs
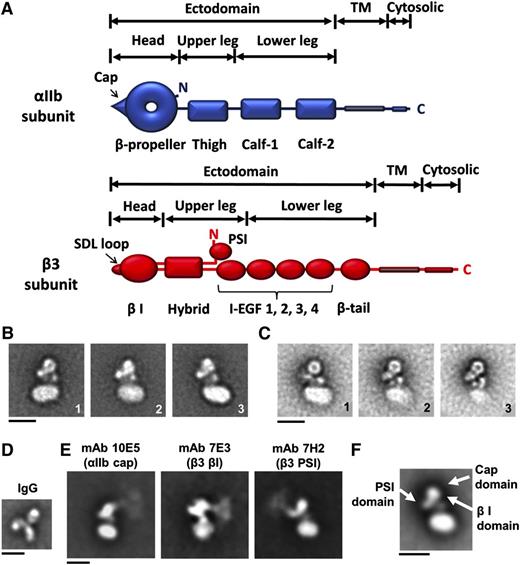
Figure 1A shows the domain structure of αIIb and β3, including the head domain, which contains a doughnut-shaped αIIb β-propeller and an adjacent cap subdomain.5,17 Using reference-free 2-dimensional (2D) alignment, class average EM images of negatively stained αIIbβ3 nanodiscs at 120 (Figure 1B) and 200 kV (Figure 1C) revealed the nanodisc and a compact αIIbβ3 ectodomain (Figure 1). We tentatively identified the αIIb head domain based on its characteristic doughnut shape and this assignment was confirmed with mAb 10E5 (Figure 1E), which binds to the αIIb cap.17,18 The β3 head containing the βI domain was identified adjacent to the αIIb head domain with mAb 7E3,19 and the β3 PSI domain in the hinge region was identified with mAb 7H2 (Figure 1E).20 Although the attachment point of each antibody to the αIIbβ3 is clear, the images blur with distance as expected based on other similar studies21,22 since, unlike the sharper images obtained with Fab fragments,23,24 the flexible immunoglobulin G (IgG) molecules adopt multiple orientations and there is always some heterogeneity in the orientation of the αIIbβ3 molecules. The most striking feature of our 2D images was the consistent orientation of the head domain away from the lipid bilayer. In addition, the lower legs adjacent to the lipid bilayer produced complex images, with 2 electron-dense circular-to-oblong structures between the αIIb β-propeller domain and the nanodisc.
2D electron microscopic images of αIIbβ3 embedded in a phospholipid bilayer nanodisc. (A) αIIb contains 7 domains and β3 contains 10 domains. The heterodimeric structure is maintained by the association of the αIIb β-propeller domain and the β3 β I domain, both of which contribute to the ligand-binding region, and the association of the αIIb and β3 TM domains. (B) Representative classified averages of αIIbβ3 nanodisc EM images obtained at 120 kV showing different views. (C) The averaged classes of αIIbβ3 nanodisc images at 200 kV. The circular-to-oblong structures between the β-propeller domain and the nanodisc are indicated by black arrows in the first image. (D) A representative class average of mouse mAb 10E5 (IgG). (E) Representative averaged images of the binding of mAb 10E5 (directed at the αIIb cap domain), mAb 7E3 (directed at β3 βI domain), and mAb 7H2 (directed at the β3 PSI domain). (F) The locations of the αIIb cap domain and the β3 PSI and βI domains (white arrows) according to the 3D map in Figure 2A. Note the correspondence between the mAb data and the 3D map data. Scale bars in panels B-F = 10 nm.
2D electron microscopic images of αIIbβ3 embedded in a phospholipid bilayer nanodisc. (A) αIIb contains 7 domains and β3 contains 10 domains. The heterodimeric structure is maintained by the association of the αIIb β-propeller domain and the β3 β I domain, both of which contribute to the ligand-binding region, and the association of the αIIb and β3 TM domains. (B) Representative classified averages of αIIbβ3 nanodisc EM images obtained at 120 kV showing different views. (C) The averaged classes of αIIbβ3 nanodisc images at 200 kV. The circular-to-oblong structures between the β-propeller domain and the nanodisc are indicated by black arrows in the first image. (D) A representative class average of mouse mAb 10E5 (IgG). (E) Representative averaged images of the binding of mAb 10E5 (directed at the αIIb cap domain), mAb 7E3 (directed at β3 βI domain), and mAb 7H2 (directed at the β3 PSI domain). (F) The locations of the αIIb cap domain and the β3 PSI and βI domains (white arrows) according to the 3D map in Figure 2A. Note the correspondence between the mAb data and the 3D map data. Scale bars in panels B-F = 10 nm.
Our ectodomain images are similar to some of those obtained by Zhu et al using a recombinant ectodomain with or without a C-terminal clasp5 and those of Eng et al using detergent-solubilized intact αIIbβ3 (if one rotates the images by ∼75°-90°),10 but differ from the previous report by Ye et al of images of αIIbβ3 inserted into nanodiscs11 in 2 major ways; we observed the nanodisc as having a disc shape (as has been reported by other investigators applying protein-containing nanodiscs to carbon-coated grids and using negative staining)25 rather than a circular shape and we found that the αIIbβ3 head domain containing the αIIb β-propeller oriented away from, rather than toward, the nanodisc. We excluded the possibility that the difference in detergent used by Ye et al to solubilize αIIbβ3 was responsible for the difference in αIIbβ3 orientation relative to the bilayer or the circular appearance of the nanodiscs by repeating our studies using the detergent they used (Triton X-100) and obtaining the same orientation and shape as with the detergent we used initially (OG). Technical differences in applying samples to the grid may account for the circular appearance of the nanodiscs studied by Ye et al because instead of directly applying the nanodiscs to a glow-discharged, carbon-coated grid, they adsorbed the negatively charged nanodiscs to a positively charged lipid monolayer supporting film.11,25 This may have led to preferential deposition of the flat surface of the nanodisc on the lipid, resulting in en face views of the nanodiscs. Alternatively, it is possible that the lipids in their nanodiscs were arrayed in a more spherical rather than flat orientation since this could also result in a more circular appearance.26
3D reconstruction of αIIbβ3 in lipid bilayer nanodiscs
We analyzed 12 504 pairs of untilted and 50° tilted 120-kV images using the RCT method, along with multiple refinements, to create 2 separate 3D reconstructions with complementary angular coverage (supplemental Figure 2). The images of extended αIIbβ3 (supplemental Figure 1F) or nanodiscs containing 2 αIIbβ3 molecules (supplemental Figure 1G) were manually excluded from the 3D reconstruction.
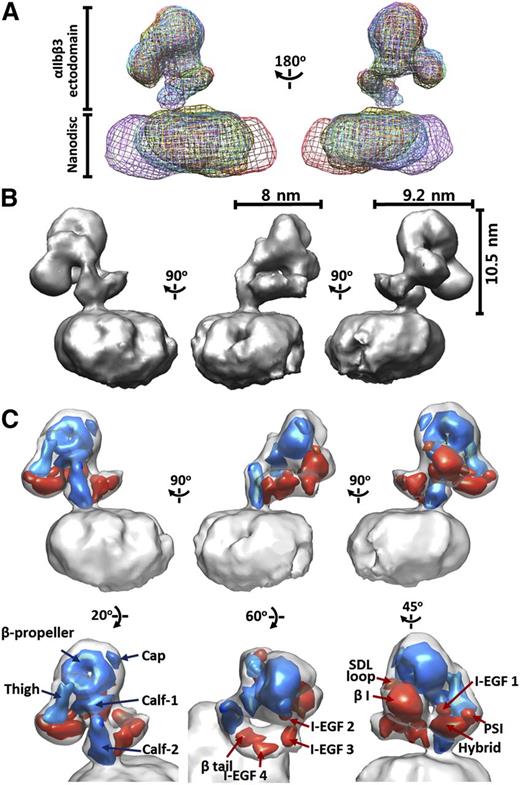
The αIIbβ3 3D maps of the resulting RCT classes were nearly identical, but the locations of the αIIbβ3 molecules in the nanodiscs varied considerably, suggesting that inactive intact αIIbβ3 has 1 consistent conformation that can diffuse laterally in the nanodisc lipid bilayer (Figure 2A). The 5 highest resolution reconstructions were combined by weighted averaging (based on the location of αIIbβ3 map) (supplemental Figure 4A) and then used as a reference for reference-based refinement. The Euler angles of the tilted images were reassigned by matching with the reference projections, and then applied to calculating the Euler angles of the untilted images. The back projection of tilted and untitled images using the assigned angles provided a map with a resolution of 28.5 Å determined from the 0.5 FSC (supplemental Figure 4B). To improve the resolution, we then obtained 6804 untilted EM images at 200 kV and performed 3D reconstruction of these images using the map as a reference (supplemental Figure 4C). The resulting map had a 0.5 FSC of 21.5 Å. To minimize artifacts resulting from having a limited and uneven angle of view, we reassigned the Euler angles of the 120-kV untilted and titled images using the higher resolution 200-kV EM map and combined the reconstructions of untilted and tilted images into a single map (supplemental Figure 5). As indicated in supplemental Figure 5A, the untilted and tilted images collectively reflect a complementary distribution of Euler angles. The final 3D reconstruction of the αIIbβ3 ectodomain showed dimensions of 8 × 9.2 × 10.5 nm with a resolution of 20.5 Å based on an FSC of 0.5 (Figure 2B).
3D reconstruction of integrin αIIbβ3 in lipid bilayer nanodisc. (A) Superimposition of 5 random conical tilt reconstructions of αIIbβ3 nanodiscs produced by back projection of tilted EM images obtained at 120 kV aligned according to the αIIbβ3 maps. Note the close alignment of the αIIbβ3 maps and the much more variable alignment of the nanodisc maps, indicating that the αIIbβ3 receptors have 1 consistent conformation but occupy different locations within the nanodiscs. (B) Three-dimensional surface views of αIIbβ3 inserted into lipid bilayer nanodiscs, along with the dimensions of the extracellular portions of αIIbβ3. (C) The 20-Å maps of the αIIb (blue) and β3 (red) domains from the crystal structure (3FCS) fitted into the αIIbβ3 nanodisc map by correlation-based docking.
3D reconstruction of integrin αIIbβ3 in lipid bilayer nanodisc. (A) Superimposition of 5 random conical tilt reconstructions of αIIbβ3 nanodiscs produced by back projection of tilted EM images obtained at 120 kV aligned according to the αIIbβ3 maps. Note the close alignment of the αIIbβ3 maps and the much more variable alignment of the nanodisc maps, indicating that the αIIbβ3 receptors have 1 consistent conformation but occupy different locations within the nanodiscs. (B) Three-dimensional surface views of αIIbβ3 inserted into lipid bilayer nanodiscs, along with the dimensions of the extracellular portions of αIIbβ3. (C) The 20-Å maps of the αIIb (blue) and β3 (red) domains from the crystal structure (3FCS) fitted into the αIIbβ3 nanodisc map by correlation-based docking.
Validation of the 3D reconstruction
The final 3D reconstruction (Figure 2B) was created by a series of refinements (supplemental Figures 2-5) and validated by comparing the reconstructions of tilt-pair EM images using the Euler angles calculated with the geometric relation between untilted and tilted EM images (equation 1 in “Materials and methods” and supplemental Figure 6). The reconstruction using the calculated angles had a resolution of 23.6 Å at 0.5 FSC, and a correlation of 21.4 Å with the refined reconstruction at 0.5 FSC (supplemental Figure 6C-D). We recognize the potential artifacts introduced by partial stain immersion and by flattening due to staining, drying, and attachment on a EM grid in negative stain EM studies.27 We specifically validated the integrity of the staining technique by observing essentially identical structures at different stain densities and with flip-flop images before the 3D reconstruction (supplemental Figure 3). The similarity of the 3D reconstructions based on the tilted and untilted images indicates that the protein did not undergo extensive flattening when attaching to the grid (supplemental Figures 5B and 6B).
Fitting the αIIbβ3 crystal structure into the 3D reconstruction
The β-propeller in the headpiece could be tentatively identified by its doughnut shape and the binding of mAb 10E5. Using the locations of the αIIb headpiece and calf-2 domains and the β3 βI, PSI, and β-tail domains as reference points, we fit the individual αIIb and β3 domains from the crystal structure into our EM map using correlation-based docking (Figure 2; supplemental Figures 7 and 8).14-16 Supplemental Table 2 details the differences in axes, tilt angles, and rotary angles of the docked domains in the EM map relative to the ectodomain crystal structure, along with the cross-correlations and clashes. As in the 2D images, the compact headpiece is located on the top of the structure and oriented away from the lipid bilayer (Figure 2). The crystal structure of the unliganded ectodomain reveals a bent conformation with a head region joined to lower legs that are nearly straight, parallel, and adjacent, with the β3 integrin-epidermal growth factor (I-EGF) 2 to 4 domains originating in a crevice between the upper β leg on one side and the extended calf-1 and calf-2 domains on the other, and extending in an almost straight orientation.5 In sharp contrast, we found: (1) nearly a right angle (91°) between the calf-1 and calf-2 domains as a result of both bending and rotation, with the calf-2 domain oriented nearly perpendicular to the lipid bilayer (Figure 3; supplemental Table 2), and (2) that the β3 lower leg is not in the crevice identified in the crystal structure, but rather follows a separate, coiled path with fewer contacts with neighboring structures, joining αIIb immediately above the lipid bilayer. Because the purified αIIbβ3 complex we studied differs from the isolated αIIbβ3 ectodomain studied by x-ray crystallography in containing the TM domains, it most likely that the coiled conformation is maintained by the orientation of the αIIbβ3 TM domains (Figure 4). Indirect support for the coiling of the β3 lower leg and its separation from the αIIb lower leg that we found in our 3D reconstruction comes from EM studies of extended αIIbβ3, which consistently show a “crossed leg” appearance rather than straight legs.10,11,28
Comparison of the EM map of intact αIIbβ3 embedded in a nanodisc with the crystal structure of the recombinant αIIbβ3 ectodomain. (A) The headpiece in the ectodomain crystal structure (ribbon diagram; PDB: 3FCS) compared with correlation-based docking of headpiece domains into the EM map. Note that the αIIbβ3 headpiece in the 3D reconstruction points away from the membrane and has a more compact conformation than the recombinant ectodomain. (B-C) The overall conformations of the leg domains of αIIb (blue) (B) and β3 (red) (C) in the EM map compared with the crystal structure (ribbon diagram; PDB: 3FCS) displayed by matching the αIIb thigh and β3 hybrid domains, respectively. Note that the αIIb and β3 lower legs in the 3D reconstruction are not adjacent, parallel, and straight as in the crystal structure. Instead, the αIIb calf-1 and calf-2 domains are bent and rotated and the β3 I-EGF 2 to 4 domains are freely coiled. (D) The structure of the calf-2 domain superimposed on the docked calf-2 domain. The disulfide-linked β-tail in the crystal structure (ribbon diagram; PDB: 3FCS) and the β-tail docked in the map are displayed. The distances between the ends of the structured residues in the calf-2 and β-tail domains (αIIb:G954 [blue sphere]; β3:C687 [red sphere]) were measured in the crystal structure (1 = 19.6 Å) and in the fitted structure (2 = 11.7 Å).
Comparison of the EM map of intact αIIbβ3 embedded in a nanodisc with the crystal structure of the recombinant αIIbβ3 ectodomain. (A) The headpiece in the ectodomain crystal structure (ribbon diagram; PDB: 3FCS) compared with correlation-based docking of headpiece domains into the EM map. Note that the αIIbβ3 headpiece in the 3D reconstruction points away from the membrane and has a more compact conformation than the recombinant ectodomain. (B-C) The overall conformations of the leg domains of αIIb (blue) (B) and β3 (red) (C) in the EM map compared with the crystal structure (ribbon diagram; PDB: 3FCS) displayed by matching the αIIb thigh and β3 hybrid domains, respectively. Note that the αIIb and β3 lower legs in the 3D reconstruction are not adjacent, parallel, and straight as in the crystal structure. Instead, the αIIb calf-1 and calf-2 domains are bent and rotated and the β3 I-EGF 2 to 4 domains are freely coiled. (D) The structure of the calf-2 domain superimposed on the docked calf-2 domain. The disulfide-linked β-tail in the crystal structure (ribbon diagram; PDB: 3FCS) and the β-tail docked in the map are displayed. The distances between the ends of the structured residues in the calf-2 and β-tail domains (αIIb:G954 [blue sphere]; β3:C687 [red sphere]) were measured in the crystal structure (1 = 19.6 Å) and in the fitted structure (2 = 11.7 Å).
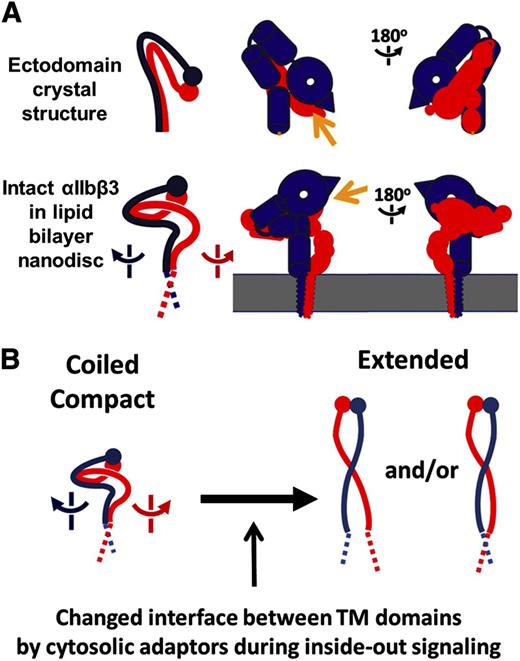
Conformational transition model of αIIbβ3 by the separation of TM domains. (A) To facilitate comparison, simplified figures demonstrating the path of linearly linked extracellular domains in αIIb (blue) and β3 (red) in the crystal structure and in the EM map from this study are displayed as the solid lines (legs) and the closed circles (head domains). Yellow arrows indicate the ligand-binding site. (B) Proposed model of conformational αIIbβ3 transition during inside-out signaling.
Conformational transition model of αIIbβ3 by the separation of TM domains. (A) To facilitate comparison, simplified figures demonstrating the path of linearly linked extracellular domains in αIIb (blue) and β3 (red) in the crystal structure and in the EM map from this study are displayed as the solid lines (legs) and the closed circles (head domains). Yellow arrows indicate the ligand-binding site. (B) Proposed model of conformational αIIbβ3 transition during inside-out signaling.
Discussion
The upward orientation of the αIIbβ3 head region we observed in our study is consistent with the cryo-EM reconstruction,8 and although the SANS shape reconstruction was interpreted as demonstrating a head-down orientation,9 our head-up reconstruction appears to fit the SANS shape as well or better than the head-down crystal structure. Our data are also consistent with the platelet binding data of the 3 mAbs we studied. The 7H2 and 10E5 epitopes are both freely available in our reconstruction and both mAbs bind equally well to unactivated and activated platelets.18,20 If the headpiece were pointed toward the membrane, one would expect that the access of mAb 10E5 to its epitope might be limited by the adjacent membrane. The 7E3 epitope in our reconstruction is oriented such that the intact antibody closely approaches the lipid bilayer, which is consistent with it being able to bind more rapidly to activated than unactivated platelets and the ability of smaller fragments of 7E3 to bind more rapidly to platelets than the intact antibody or multimeric forms of the antibody.29,30 Because mAb 7E3 binds very close to the ligand-binding pocket,19 its binding properties indicate that even with the head-up orientation in our reconstruction, access of large ligands to the binding pocket may be limited by the latter’s proximity to the membrane. The relationship between the αIIb calf-2 and β3 β-tail domains differs in our 3D reconstruction compared with the crystal structure of the recombinant ectodomain, and this may reflect the effects of the TM domains present in our αIIbβ3 preparation in contrast to the recombinant ectodomain used for that study.5 The ectodomain construct also contained an engineered disulfide bond (α959C/β688C) that retained the receptor in the bent conformation and partially reduced ligand binding in the presence of Mn2+ and the activating mAb PT25-2.5 The crystal structure of αIIbβ3 suggests considerable flexibility in this region, with large unstructured loops in calf-2 of αIIb and weak or missing densities in the β3 β-tail.5 Moreover, based on cysteine scanning and the variation in orientation of the calf-2 and β-tail domains in their crystal molecules 1 and 2, Zhu et al suggested that a range of orientations of the calf-2 and β-tail domains can occur on the cell surface.5 Our 3D reconstruction also differs from the crystal structure and a 3D EM reconstruction of the ectodomain of αVβ3 in this region, even though the construct used for those studies did not include an engineered disulfide bond.4,7 As noted by Zhu et al,5 however, the β-tail region of αVβ3 was the site of a large crystal lattice contact that may have had an impact on the structure. In addition, since the ectodomain was studied, it did not include the TMs in a lipid bilayer and the resolution of the 3D reconstruction (∼30Å) was not sufficient for fine localization of secondary structures.7
Our reconstruction has implications for the mechanism of αIIbβ3 activation. The deadbolt model, which was based on the αVβ3 crystal structure, suggests that access to the ligand-binding site is limited in the inactive receptor by an interaction between the β3 β-tail domain and the β3 βI domain.31 In our reconstruction, however, the β3 β-tail domain and the β3 βI domains are not adjacent to each other. The switchblade hypothesis of inside-out activation has 3 elements: (1) the inactive receptor is bent and undergoes extension with activation, (2) the αIIbβ3 head region points toward the platelet membrane, and (3) extension is achieved by separation of the lower α and β subunit legs, leading to destabilization of their interfaces with head and upper legs, ultimately leading to a change in the fulcrum between the upper and lower legs of αIIb and β3 involving the αIIb genu and the region between the β3 I-EGF1 and 2 domains.32 Our finding of a compact conformation for inactive αIIbβ3 is consistent with a model wherein extension contributes to activation, but our structure requires modification of the switchblade hypothesis since we found that the head region is pointing away from the lipid bilayer. Moreover, because we found that the lower legs have complex conformations, receptor extension requires changes in multiple interdomain articulations rather than a change in a single fulcrum. One of the major conceptual challenges posed by the αIIbβ3 ectodomain crystal structure has been understanding how inside-out signaling (which evidence indicates is triggered by changes in the interactions of cytoplasmic proteins with the integrin cytoplasmic domains resulting in unwinding of the coiled coil TM domains)33-37 can transmit a signal to the ectodomain that is sufficient to induce the β3 I-EGF 2 to 4 domains to exit from its crevice. Our data indicate, however, that the β3 I-EGF 2 to 4 domains are not in a crevice and do not make extensive contacts with neighboring structures. Our data also suggest an important role for the region that links the distal calf-2 and β-tail domains to their respective TM domains in transmitting the conformational changes in the TM domains associated with inside-out activation, a region previously implicated in affecting ligand binding.5
The electron microscopy map of the integrin αIIbβ3 nanodisc has been deposited under accession number EMD-2281 in the Electron Microscopy Data Bank, and the fitted map has been deposited under ID code 4cak in the Protein Data Bank.
The online version of this article contains a data supplement.
There is an Inside Blood commentary on this article in this issue.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.
Acknowledgments
We thank Drs Seth Darst, Natacha Opalka, Brian Bae, and Kunihiro Uryu for valuable suggestions, Claudia Gelfond for assistance with EM imaging, and Suzanne Rivera for administrative assistance.
This work was supported, in part, by grants HL19278 from the National Institutes of Health, National Heart, Lung, and Blood Institute; Clinical and Translational Science Award grant ULRR024143 from the National Institutes of Health, National Center for Research Resources and the Center for Advancing Translational Sciences; funds from Stony Brook University; and a National Research Foundation of Korea grant funded by the Korean Government (NRF-2009-352-E00042). The investigation was conducted in a facility constructed with support from Research Facilities Improvement Program grant number C06RR017528-01-CEM from the National Institutes of Health, National Center for Research Resources.
Authorship
Contribution: W.-S.C., W.J.R., D.L.S., and B.S.C. designed the study; W.-S.C. and W.J.R. performed the research; W.-S.C. and W.J.R. analyzed data; and W.-S.C. and B.S.C. prepared the figures and wrote the manuscript.
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Correspondence: Barry S. Coller, Allen and Frances Adler Laboratory of Blood and Vascular Biology, The Rockefeller University, 1230 York Ave, New York, NY 10065; e-mail: collerb@rockefeller.edu.

![Figure 3. Comparison of the EM map of intact αIIbβ3 embedded in a nanodisc with the crystal structure of the recombinant αIIbβ3 ectodomain. (A) The headpiece in the ectodomain crystal structure (ribbon diagram; PDB: 3FCS) compared with correlation-based docking of headpiece domains into the EM map. Note that the αIIbβ3 headpiece in the 3D reconstruction points away from the membrane and has a more compact conformation than the recombinant ectodomain. (B-C) The overall conformations of the leg domains of αIIb (blue) (B) and β3 (red) (C) in the EM map compared with the crystal structure (ribbon diagram; PDB: 3FCS) displayed by matching the αIIb thigh and β3 hybrid domains, respectively. Note that the αIIb and β3 lower legs in the 3D reconstruction are not adjacent, parallel, and straight as in the crystal structure. Instead, the αIIb calf-1 and calf-2 domains are bent and rotated and the β3 I-EGF 2 to 4 domains are freely coiled. (D) The structure of the calf-2 domain superimposed on the docked calf-2 domain. The disulfide-linked β-tail in the crystal structure (ribbon diagram; PDB: 3FCS) and the β-tail docked in the map are displayed. The distances between the ends of the structured residues in the calf-2 and β-tail domains (αIIb:G954 [blue sphere]; β3:C687 [red sphere]) were measured in the crystal structure (1 = 19.6 Å) and in the fitted structure (2 = 11.7 Å).](https://ash.silverchair-cdn.com/ash/content_public/journal/blood/122/26/10.1182_blood-2013-04-499194/4/m_4165f3.jpeg?Expires=1769188158&Signature=Rj-FNHqQJkFOHEj6tU4nzR4eYwofjzDpq2lArx44M0QT~w9Kr1VF28mAJdk2JMrfqRjqQ-MuR8UJDYAYlEK7qSnJGtF1tSDrUfNWDf1D6yd0F9gncacvpZNuPlO8V68hr74z-JOlgpDI9SI5mdRY~qV9dmza4~u6WKHNvVBxpjDnwWELo~l~8kFQsF7Ga2pRkccB2T-SVp2R3-9GN8PRxt0T~sT7~ztynv6ZfmPoKUIuZ67PQwRjFwbKyDs04Nqp-U-XMFBHRHm5PaklrMt7bsUx-okKGrAPcDNZOLfnw8moa2l5IR~YAoGTCIfWfVy6YEqIe-DIB77vqzF~NIF1PA__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)
This feature is available to Subscribers Only
Sign In or Create an Account Close Modal