Key Points
EBV-coding miRNAs are transferred from infected into noninfected cells by exosome to regulate the function for the tumorigenesis.
Production of EBV-coding miRNAs will be an excellent diagnostic marker to separate patients with EBV+ diffuse large B-cell lymphoma into 2 groups.
Abstract
Epstein-Barr virus (EBV) causes various diseases in the elderly, including B-cell lymphoma such as Hodgkin’s lymphoma and diffuse large B-cell lymphoma. Here, we show that EBV acts in trans on noninfected macrophages in the tumor through exosome secretion and augments the development of lymphomas. In a humanized mouse model, the different formation of lymphoproliferative disease (LPD) between 2 EBV strains (Akata and B95-8) was evident. Furthermore, injection of Akata-derived exosomes affected LPD severity, possibly through the regulation of macrophage phenotype in vivo. Exosomes collected from Akata-lymphoblastoid cell lines reportedly contain EBV-derived noncoding RNAs such as BamHI fragment A rightward transcript (BART) micro-RNAs (miRNAs) and EBV-encoded RNA. We focused on the exosome-mediated delivery of BART miRNAs. In vitro, BART miRNAs could induce the immune regulatory phenotype in macrophages characterized by the gene expressions of interleukin 10, tumor necrosis factor-α, and arginase 1, suggesting the immune regulatory role of BART miRNAs. The expression level of an EBV-encoded miRNA was strongly linked to the clinical outcomes in elderly patients with diffuse large B-cell lymphoma. These results implicate BART miRNAs as 1 of the factors regulating the severity of lymphoproliferative disease and as a diagnostic marker for EBV+ B-cell lymphoma.
Introduction
Epstein-Barr virus (EBV) is an oncogenic human γ-herpes virus that causes various diseases such as B-cell lymphoma, T-cell and natural killer cell lymphoma, gastric cancer, nasopharyngeal carcinoma, and some autoimmune diseases.1,2 Among these diseases, B-cell lymphoma is 1 of the common forms in EBV-related cancer because of the strong tropism of EBV to B cells. EBV contains several oncogenes that encode proteins; these genes include latent membrane protein (LMP) and EBV nuclear antigen (EBNA). Once B cells are infected, expression of these genes induces immortalization and aberrant proliferation of the cells, leading to the development of lymphoblastoid cell lines (LCLs).1-4 Clinical studies have demonstrated that EBV+ cases show a poorer prognosis than EBV− cases in Hodgkin’s lymphoma and diffuse large B-cell lymphoma (DLBCL) of the elderly.5,6 Therefore, the establishment of a novel therapeutic strategy that specifically treats EBV+ B-cell lymphoma is required.
Presently, we found significant differences in the survival rate, macrophage infiltration, and EBV-positive tumor cells between mice infected with the Akata and B95-8 strains of EBV, regardless of the similar transforming ability of the strains in vitro, suggesting that these EBV strains have a different ability to form a tumor microenvironment.
Akata was originally isolated from Burkitt lymphoma. B95-8 was originally isolated from infectious mononucleosis and is maintained in marmoset LCLs. Hence, they are completely different strains, and so display several differences. We focused on the deletion in BamH1 fragment A rightward transcript (BART) region on the B95-8 genome as the most outstanding difference. Interestingly, 40 micro-RNAs (miRNAs) were reported to cluster and were transcribed from the BART region (BART miRNAs).7 miRNAs are small noncoding RNAs approximately 22 nucleotides in length that posttranscriptionally regulate the deadenylation, translation, and decay of their target messenger RNAs (mRNAs).8,9
EBV+ B-cell lymphoma is composed of a proportion of tumor cells and a large proportion of nontumor cells including immune cells, which is termed the inflammatory niche. This suggests that the inflammatory niche is necessary for the survival and growth of tumor cells. Although it is unclear whether BART miRNAs affect the formation of the tumor microenvironment, recently, Pegtel et al10 showed that EBV+ lymphoma cells secreted BART miRNAs through extracellular vesicles called exosomes and that monocyte-derived dendritic cells selectively incorporate the exosomes. Exosomes are cell-derived vesicles that function as communicators of various molecules, such as proteins, mRNAs, and miRNAs, from donor cells to recipient cells.11-13 In tumor studies, tumor-derived exosomes supported tumor development and metastasis through various mechanisms.11-15 It was reported that EBV+ lymphoma cells secrete exosomes and deliver EBV-derived noncoding RNA, such as BART miRNAs and EBV-encoded RNA (EBER),10,16 suggesting that tumor-derived exosomes could affect the development of EBV+ lymphoma. Exosome-mediated secretion of miRNAs also appears to be critical for the formation of the metastatic niche.15 In this study, we demonstrate the essential role of BART miRNAs for the formation of an inflammatory niche in EBV+ B-cell lymphoma. We compared lymphoma forming capacity between 2 EBV strains, Akata and B95-8, using a humanized mice model, and observed different lymphoma formation. Furthermore, we showed the role of exosomes for the development of EBV+ B-cell lymphoma in vivo, possibly through regulation of macrophage infiltration. Exosome-mediated delivery of BART miRNAs was critical for the induction of the immune regulatory phenotype in macrophages in vitro. Furthermore, analysis of EBV+ DLBCL in elderly patients revealed the strong correlation between the expression levels of BART miRNAs and clinical outcomes of the lymphoma. These results suggest that BART miRNAs could be a promising diagnostic tool as well as a novel therapeutic target of EBV+ lymphoma.
Results
Differential formation of lymphoproliferative diseases in vivo between Akata and B95-8 strains
The functional human immune system, including T cells, B cells, and natural killer lymphocytes, was reconstituted in nonobese diabetic (NOD)/shi-scid, interleukin (IL)-2Rγ null mice (NOG mice) that received hematopoietic stem cell transplants.17 These humanized mice recapitulate the human lymphoproliferative disease (LPD) induced by EBV infection.18 To date, a variety of EBV strains have been isolated from patients. Although all these strains equivalently transform human B ells in vitro, the potential for tumorigenicity in vivo involving nontumor cells is not well understood.
To clarify the differences in tumorigenicity between EBV strains, mice were inoculated by tail vein injection with a low (1 × 103) 50% transforming dose (supplemental Figure 1, available on the Blood Web site) of either the Akata or B95-8 EBV strain. Both are type I strains. Akata was isolated from EBV+ Burkitt lymphomas, in which EBV exhibits the type I form of latency. B95-8 was isolated from infectious mononucleosis and maintained in infected marmoset LCL, which shows the type III form of latency.19,20
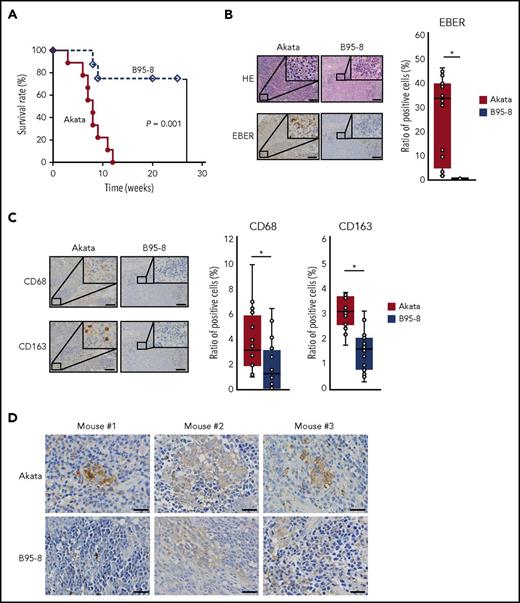
All mice infected with Akata died within 12 weeks with massive infiltration of EBV-encoded RNA (EBER)+ cells evident in the spleen (Figure 1A-B). In contrast, approximately 70% of mice infected with B95-8 survived (Figure 1A). These surviving mice had few EBER+ cells in the spleen (Figure 1B) and did not show symptoms of LPD, such as elevated viral load or body weight loss (data not shown). Pathological examination indicated increased infiltration of both CD68+ and CD163+ cells (markers for M1 and M2 macrophages, respectively) in the spleens of mice infected with Akata compared with those infected with B95-8 (Figure 1C). Cells were counted in 16 fields of 3 Akata-infected mice and 20 fields of 3 B95-8-infected mice. Notably, the distribution of CD68+ and CD163+ cells seemed not to overlap. CD68+ cells were sparse in the EBER+ tumor cell-rich area, whereas numerous CD163+ cells were located surrounding the tumor cell-rich area. Thus, the Akata and B95-8 strains showed different potentials for lymphomagenesis in vivo, despite having a similar titer in vitro. Consistent with increased infiltration of lymphoma cells and macrophages, increased expression of IL-10 was detected in the spleens of mice infected with Akata (Figure 1D), suggesting that infection with Akata could induce immune suppressive environment.
Equivalent transformation units of Akata and B95-8 virus demonstrate differential formation of lymphoproliferative diseases in vivo. (A) Survival curve of Akata (n = 9, solid line) and B95-8-infected mice (n = 8, dashed line). P = .001, log-rank test. (B-C) Immunohistochemical staining of spleens from EBV-infected mice. (B) The spleens were stained with hematoxylin and eosin or treated for EBER in situ hybridization. Scale bar, 100 μm. (C) CD68 and CD163 immunohistochemical staining of the spleen. Scale bar, 100 μm. Cells were counted in 16 fields from 3 Akata-infected mice and 20 fields from 3 B95-8-infected mice, respectively. Ratio to total cells were calculated. *P < .01, Mann-Whitney U test. (D) Production of IL-10 was detected by immunohistochemical staining of spleens collected from EBV-infected mice, following the manufacturer’s protocol. Rabbit anti-human IL-10 (LS-B7432) was purchased from LifeSpan BioSciences, Inc. (Seattle, WA). Biopsies were collected from 3 Akata- and B95-8-infected mice. Scale bar, 25 μm.
Equivalent transformation units of Akata and B95-8 virus demonstrate differential formation of lymphoproliferative diseases in vivo. (A) Survival curve of Akata (n = 9, solid line) and B95-8-infected mice (n = 8, dashed line). P = .001, log-rank test. (B-C) Immunohistochemical staining of spleens from EBV-infected mice. (B) The spleens were stained with hematoxylin and eosin or treated for EBER in situ hybridization. Scale bar, 100 μm. (C) CD68 and CD163 immunohistochemical staining of the spleen. Scale bar, 100 μm. Cells were counted in 16 fields from 3 Akata-infected mice and 20 fields from 3 B95-8-infected mice, respectively. Ratio to total cells were calculated. *P < .01, Mann-Whitney U test. (D) Production of IL-10 was detected by immunohistochemical staining of spleens collected from EBV-infected mice, following the manufacturer’s protocol. Rabbit anti-human IL-10 (LS-B7432) was purchased from LifeSpan BioSciences, Inc. (Seattle, WA). Biopsies were collected from 3 Akata- and B95-8-infected mice. Scale bar, 25 μm.
EBV+ lymphoma-derived exosomes cause severe LPD in humanized mice model
Recently, the involvement of the exosome, 1 of the extracellular vesicles, has been reported in the development and metastasis of various tumors.11-15 The exosome is composed of a lipid bilayer with a diameter of 50 to 200 nm and is a carrier of biological molecules, including proteins, lipids, and nucleic acids. Importantly, EBV+ B-cell lymphoma cells secrete exosomes.10,16 To investigate the involvement of exosomes in the differential formation of LPD between the Akata and B95-8 strains, we intravenously injected exosomes into B95-8-infected humanized mice.
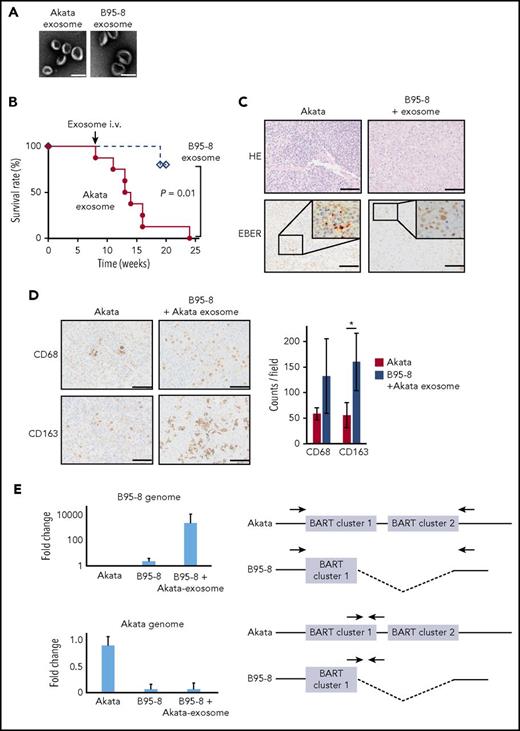
Exosomes were isolated from the culture supernatants of Akata- and B95-8-LCL cultures (Akata and B95-8 exosomes, respectively), using the differential centrifugation protocol.21 This method isolates and purifies exosomes from the supernatant of B-cell culture without EBV.10 EBV-infected cell-derived exosomes with a diameter of 10 to 100 nm were observed by electron microscopy (Figure 2A). The mice infected with B95-8 were injected with either Akata or B95-8 exosomes intravenously at 8 weeks postinfection. Surprisingly, the injection of Akata exosomes caused severe LPD in 6 of the 8 mice within 3 to 7 weeks after injection, but the injection of B95-8 exosomes did not do so in 5 mice (Figure 2B). Pathological analysis showed that the infiltration of EBER+ lymphoma cells in the spleen of B95-8-infected mice was recovered to a level that was comparable to that of the Akata-infected mice by the injection of Akata exosomes (Figure 2C). Furthermore, the infiltration of both CD68+ and CD163+ macrophages was increased 2- to 3-fold by the injection of Akata exosomes (Figure 2D). To count the number of CD68+ cells and CD163+ cells, 2 mice were analyzed in each group, and 3 fields were counted in each mouse.
Exosomes collected from Akata-LCLs accelerate LPD formation. (A) Electron microscope image of exosomes isolated from Akata-LCL (left) and B95-8-LCL culture medium by ultracentrifugation. Scale bar, 100 nm. (B) Survival curve of B95-8/miRNA-rich exosome-treated mice (n = 8, solid line) or B95-8/EBV-miRNA-deleted exosomes-treated mice (n = 5, dashed line). P = .01, log-rank test. (C) The spleens were stained with hematoxylin and eosin or treated for EBER in situ hybridization. Scale bar, 50 μm. (D) CD68 and CD163 immunohistochemical staining of the spleen (left). Scale bar, 50 μm. The cells positive for each macrophage marker were counted (right). Two mice were treated per group, and 3 fields were counted in each mouse. *P < .01, Student t test. (E, left top) qPCR revealed that the B95-8 virus-specific region was amplified in LPD in B95-8/miRNA-rich exosome-treated mice. (Left bottom) Akata virus-specific region was not amplified. (Right) The binding sites of primer pairs used to specifically amplify the B95-8 virus genome and Akata virus genome are indicated. Indicated primer pair detects B95-8 genome (top) and Akata genome (bottom), respectively. Half of the BART cluster 1 miRNAs, and all BART cluster 2 miRNAs were deleted in the B95-8, but not in Akata virus.
Exosomes collected from Akata-LCLs accelerate LPD formation. (A) Electron microscope image of exosomes isolated from Akata-LCL (left) and B95-8-LCL culture medium by ultracentrifugation. Scale bar, 100 nm. (B) Survival curve of B95-8/miRNA-rich exosome-treated mice (n = 8, solid line) or B95-8/EBV-miRNA-deleted exosomes-treated mice (n = 5, dashed line). P = .01, log-rank test. (C) The spleens were stained with hematoxylin and eosin or treated for EBER in situ hybridization. Scale bar, 50 μm. (D) CD68 and CD163 immunohistochemical staining of the spleen (left). Scale bar, 50 μm. The cells positive for each macrophage marker were counted (right). Two mice were treated per group, and 3 fields were counted in each mouse. *P < .01, Student t test. (E, left top) qPCR revealed that the B95-8 virus-specific region was amplified in LPD in B95-8/miRNA-rich exosome-treated mice. (Left bottom) Akata virus-specific region was not amplified. (Right) The binding sites of primer pairs used to specifically amplify the B95-8 virus genome and Akata virus genome are indicated. Indicated primer pair detects B95-8 genome (top) and Akata genome (bottom), respectively. Half of the BART cluster 1 miRNAs, and all BART cluster 2 miRNAs were deleted in the B95-8, but not in Akata virus.
To exclude the possibility of contamination of virus particles within the collected exosomes, the EBV genome was amplified by PCR. The B95-8 genome, but not the Akata genome, was detected at high levels in the mice injected with Akata exosomes after B95-8 infection, indicating that the proliferating cells were infected with B95-8, but not with Akata (Figure 2E). These results suggest that exosomes derived from Akata-infected lymphoma caused the severe LPD in the humanized mouse model, which was associated with increased infiltration of macrophages in lymphoma tissue.
CD163+ macrophage depletion induces the elimination of EBER+ lymphoma cells in the humanized mouse model
A large number of infiltrated macrophages in LPDs suggest an essential role for macrophages in EBV-related lymphoma. Accumulating evidence has revealed the critical roles of macrophages for the tumor microenvironment.22,23
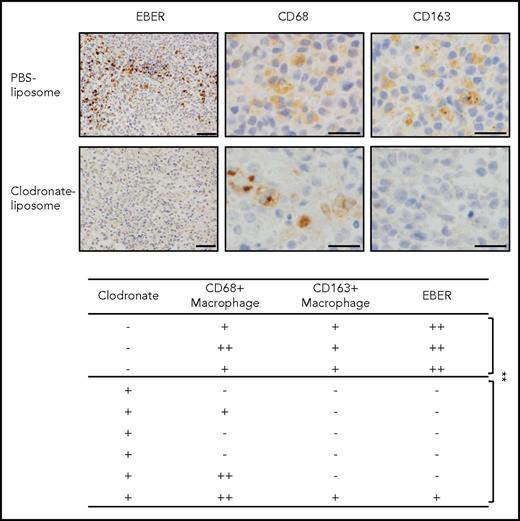
To determine the specific functions of macrophages in the tumor, the macrophages in the Akata-infected mice were depleted by clodronate liposomes, which specifically eliminated the macrophages. Surprisingly, a single injection of clodronate liposomes into mice with LPD eliminated CD163+ macrophages as well as the EBER+ lymphoma cells (Figure 3). Notably, although CD68+ macrophages were not completely cleared, EBER+ lymphoma cells completely disappeared in some of the mice. Three and 6 mice were treated with phosphate-buffered saline or clodronate liposomes, respectively. This may reflect the function of CD163+ macrophages in human patients. These results suggest that macrophages are essential for the development of EBV+ lymphoma, and especially suggest that CD163+ macrophages may act as key bystander cells.
Depletion of CD163+macrophages induces the elimination of EBER+lymphoma cells in humanize mice model. Clodronate liposome (300 μL/mouse) was injected into Akata-infected mice (intraperitoneally). After 5 days, the spleens were collected. EBER was detected by in situ hybridization (upper). Scale bar, 50 μm. CD68/163 was detected by immunohistochemical staining. Scale bar, 20 μm. The effects of clodronate liposome in mice (lower). EBER and CD68/CD163 expression was analyzed in each mouse spleen. -, no EBER or CD68/163-positive cells in 1 microscopic field; +, <10 positive cells; ++, >10 positive cells. **P < .05, Student t test.
Depletion of CD163+macrophages induces the elimination of EBER+lymphoma cells in humanize mice model. Clodronate liposome (300 μL/mouse) was injected into Akata-infected mice (intraperitoneally). After 5 days, the spleens were collected. EBER was detected by in situ hybridization (upper). Scale bar, 50 μm. CD68/163 was detected by immunohistochemical staining. Scale bar, 20 μm. The effects of clodronate liposome in mice (lower). EBER and CD68/CD163 expression was analyzed in each mouse spleen. -, no EBER or CD68/163-positive cells in 1 microscopic field; +, <10 positive cells; ++, >10 positive cells. **P < .05, Student t test.
Incorporation of exosomes into monocytes in an “eat me” signal-dependent manner
Exosomes derived from Akata-LCL increased the infiltration of macrophages, especially CD163+ macrophages. The phenotype of macrophages can be differently regulated by various cytokines, such as interferon-γ and IL-4, which are produced from T helper 1 and T helper 2 cells, respectively.24,25 Here, to investigate whether the macrophage phenotype could be regulated directly or indirectly by the lymphoma-derived exosomes, human peripheral blood mononuclear cells (PBMCs) were treated with exosomes in vitro.
Similar to Figure 2, exosomes were isolated from the culture supernatant of Akata- and B95-8-LCL by ultracentrifugation. Fractionation of isolated exosomes by iodixanol gradient centrifugation26 detected CD63, an exosome marker, from fractions 3 thorough 6 (Figure 4A), indicating that exosomes were successfully isolated from the culture supernatant. Because CD63 is highly glycosylated,27 the different sizes of CD63 between Akata and B95-8 exosomes suggest the existence of a different glycosylation machinery in Akata- and B95-8-LCLs. To monitor the transfer of the exosomes secreted from the lymphoma cells to immune cells, we labeled the exosomes collected from Akata- and B95-8-LCL with a red fluorescent lipid dye (PKH26). Labeled exosomes were captured on magnetic beads and confirmed by confocal microscopy (supplemental Figure 2D).
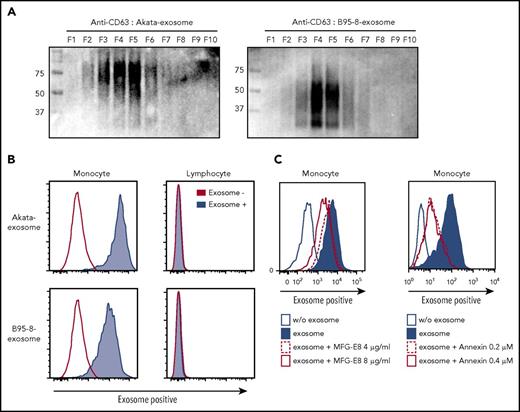
“Eat me” signal mediates exosome uptake by monocytes/macrophages. (A) Exosomes were collected from culture supernatant of Akata- and B95-8-LCLs by ultracentrifugation and further fractionated by iodixanol gradient centrifugation. CD63 expression in each exosome was detected by western blotting under nonreducing conditions. (B) PBMCs were treated with 2.5 mg/mL Akata and B95-8 exosomes for 48 hours. Flow cytometric analysis revealed strong incorporation of exosomes into monocytes, but not into lymphocytes. (C) Each dose of recombinant MFG-E8 and annexin V was added to the PBMCs and cultured for 4 days. Incorporation of PKH-labeled exosomes by monocytes was detected as described in Figure 4B. The inhibition of excess of MFG-E8 or annexin V on exosome uptake was repeated 5 times, using PBMCs derived from 2 individuals.
“Eat me” signal mediates exosome uptake by monocytes/macrophages. (A) Exosomes were collected from culture supernatant of Akata- and B95-8-LCLs by ultracentrifugation and further fractionated by iodixanol gradient centrifugation. CD63 expression in each exosome was detected by western blotting under nonreducing conditions. (B) PBMCs were treated with 2.5 mg/mL Akata and B95-8 exosomes for 48 hours. Flow cytometric analysis revealed strong incorporation of exosomes into monocytes, but not into lymphocytes. (C) Each dose of recombinant MFG-E8 and annexin V was added to the PBMCs and cultured for 4 days. Incorporation of PKH-labeled exosomes by monocytes was detected as described in Figure 4B. The inhibition of excess of MFG-E8 or annexin V on exosome uptake was repeated 5 times, using PBMCs derived from 2 individuals.
PBMCs were treated with the PKH26-labeled Akata and B95-8 exosomes for 48 hours. Flow cytometry analysis showed that almost all the monocytes incorporated both exosomes (Figure 4B), and that incorporation was dependent on the dose of the exosomes and accumulated with time (supplemental Figure 3). In contrast, incorporation of exosomes by lymphocytes was observed only when treated with a high dose of Akata exosomes for 6 hours, not with B95-8 exosomes (supplemental Figure 3). Exosome+ lymphocytes were not observed 24 and 48 hours after treatment with both Akata and B95-8 exosomes. These results suggest that EBV+ lymphoma-derived exosomes could transiently affect lymphocytes or attach to lymphocytes in high concentration, whereas substantial effects are expected on monocytes. The results indicate that exosomes are secreted from EBV-infected B cells and are then mainly incorporated into monocytes/macrophages, suggesting that lymphoma-derived exosomes could directly affect the macrophage phenotype.
Recognition of phosphatidylserine (PS), also known as the “eat me” signal, is a key step for the uptake of exosomes.16 The uptake of PKH-stained exosomes by monocytes was partially blocked by the excess amounts of PS-binding molecules, human recombinant milk fat globule epidermal growth factor VIII (MFG-E8) and annexin V, in a dose-dependent manner (Figure 4C, left and right, respectively). These results indicate that EBV+ lymphoma-derived exosomes are incorporated into monocytes in an eat me signal-dependent manner.
Treatment with exosomes shifts the phenotype of monocytes to the immune regulatory phenotype via BART miRNA delivery
To elucidate the direct effects of EBV+ lymphoma-derived exosomes on monocytes/macrophages, we investigated the effects of exosome uptake on monocytes phenotype in vitro. Akata exosomes showed enhanced CD69 expression in the CD14+ monocytes compared with B95-8 exosomes (Figure 5A). This result suggests the regulatory role of Akata exosomes in inflammatory responses of macrophages. Importantly, EBV+ lymphoma-derived exosomes deliver noncoding RNAs, such as BART miRNAs and EBER, and can affect immune regulation.10,16 We compared the expression level of EBER1 between Akata and B95-8 exosomes by reverse transcription quantitative polymerase chain reaction (RT-qPCR). The expression of EBER1 was slightly higher in B95-8 exosomes (about 1.6-fold) than in Akata exosomes, but was not significantly different (supplemental Figure 4).
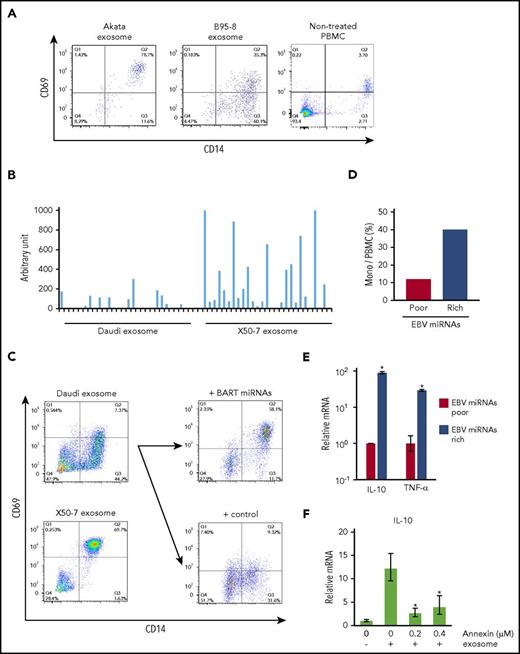
Treatment with exosomes shifts the phenotype of monocytes to the immune regulatory phenotype via BART miRNAs delivery. (A) PBMCs were cultured with PKH26-labeled Akata-LCL and B95-8-LCL exosomes. CD14 and CD69 expression was detected by flow cytometry. (B) Exosomes were harvested from Daudi and X50-7 culture supernatant. BART miRNA expression level in exosomes was measured by qPCR. Each column indicates each BART miRNA. (C) PBMCs were cultured with each type of exosome. CD14 and CD69 expression was detected by flow cytometry (left). Exogenous BART miRNAs were introduced into Daudi cells, and exosomes were harvested from cells transduced with control or BART miRNA-expressing vector. CD69 expression was upregulated on monocytes (right top) by exosomes collected from BART miRNA-transduced cells as compared with the control (right bottom). (D) Monocyte ratios in PBMCs were analyzed by flow cytometry. (E) Total RNA from PBMCs was collected. Each mRNA expression level was determined by qPCR. Fold-induction was calculated as the ratio of EBV miRNA-rich value to EBV miRNA-poor value. (F) mRNA expression level of IL-10 was detected by qPCR. The relative amount of IL-10 mRNA was standardized by the absolute amount of that the samples without exosome and annexin. Statistical significance was calculated by comparing that on 0 µg/µL of annexin. The triplicated value were analyzed twice. *P < .05, Student t test.
Treatment with exosomes shifts the phenotype of monocytes to the immune regulatory phenotype via BART miRNAs delivery. (A) PBMCs were cultured with PKH26-labeled Akata-LCL and B95-8-LCL exosomes. CD14 and CD69 expression was detected by flow cytometry. (B) Exosomes were harvested from Daudi and X50-7 culture supernatant. BART miRNA expression level in exosomes was measured by qPCR. Each column indicates each BART miRNA. (C) PBMCs were cultured with each type of exosome. CD14 and CD69 expression was detected by flow cytometry (left). Exogenous BART miRNAs were introduced into Daudi cells, and exosomes were harvested from cells transduced with control or BART miRNA-expressing vector. CD69 expression was upregulated on monocytes (right top) by exosomes collected from BART miRNA-transduced cells as compared with the control (right bottom). (D) Monocyte ratios in PBMCs were analyzed by flow cytometry. (E) Total RNA from PBMCs was collected. Each mRNA expression level was determined by qPCR. Fold-induction was calculated as the ratio of EBV miRNA-rich value to EBV miRNA-poor value. (F) mRNA expression level of IL-10 was detected by qPCR. The relative amount of IL-10 mRNA was standardized by the absolute amount of that the samples without exosome and annexin. Statistical significance was calculated by comparing that on 0 µg/µL of annexin. The triplicated value were analyzed twice. *P < .05, Student t test.
Although several differences in the genomes have been reported between Akata and B95-8,28 1 of the characteristic differences is a lack of the 12-kb BART locus in the B95-8 genome, where a number of BART miRNAs are encoded. Therefore, we focused on the effects of exosome-mediated delivery of BART miRNAs on macrophage phenotype.
Next, we examined whether similar effects were observed by treatment with exosomes collected from different cell lines. Daudi, Burkitt lymphoma cells, and the LCL X50-7 display different expression levels of BART miRNAs.29,30 Exosomes were collected from the culture supernatant of these cells, and the expression levels of BART miRNAs in each exosome were compared. BART miRNAs were highly expressed in X50-7-derived exosomes compared with Daudi-derived exosomes, reflecting the difference in BART miRNA levels between 2 cell lines (Figure 5B). As compared with Daudi-derived exosomes (BART miRNA-poor), treatment with X50-7-derived exosomes (BART miRNA-rich) upregulated CD69 expression in CD14+ monocytes (Figure 5C, left), similar to the results depicted in Figure 5A. In addition, treatment of PBMCs with BART miRNA-recovered exosomes, collected from Daudi cells exogenously introduced with BART miRNA-expressing vector, resulted in enhanced CD69 expression in CD14+ monocytes as compared with that in the control (Figure 5C, right panel). Furthermore, the survival rate of monocytes was increased (Figure 5D), and the expression of tumor necrosis factor-α (TNF-α) and IL-10 was consistently upregulated (Figure 5E) in cells treated with BART miRNA-rich X50-7-derived exosomes. Treatment with an overdose of annexin V, which binds to PS to block the eat me signal (Figure 4C) also inhibited IL-10 expression in monocytes (Figure 5F).
These results suggest that exosome-mediated delivery of BART miRNAs could contribute to the phenotypic changes of primary monocytes. In particular, induction of TNF-α and IL-10 expression suggests that BART miRNAs are key molecules to induce the immune regulatory phenotype.
BART miRNAs dramatically change gene expression in THP-1 cells
To explore how BART miRNAs affect the phenotypic changes in monocytes, we established an inducible BART miRNA expression system (Tet-Off system) in THP-1, human monocytic leukemia cells (BART/THP-1; Figure 6A).
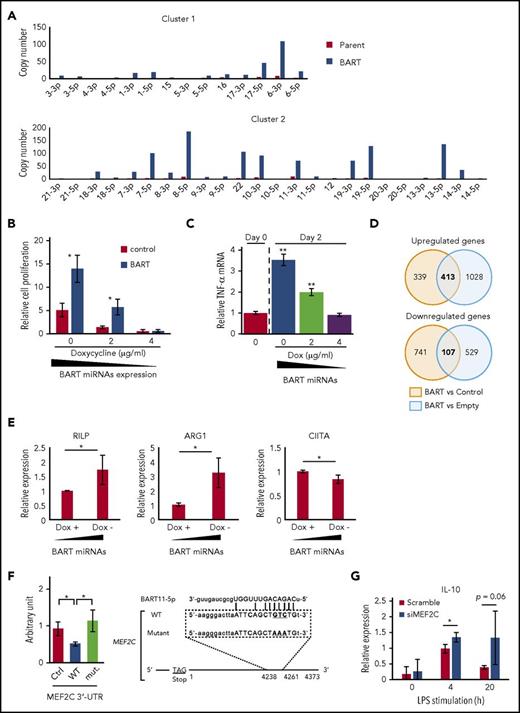
BART miRNAs alter gene expression in THP-1 cells. (A) BART miRNAs were overexpressed in THP-1 cells with the Tet-Off system. (B) BART miRNA expression was regulated by doxycycline. After 7 to 9 days of culture, the number of cells was counted. Cell proliferation is expressed as the fold-change relative to the cell number at day 0. *P < .05, using Student t test, compared with control values of each doxycycline concentration. (C) Total RNA was collected from BART/THP-1 cells at the indicated time and doxycycline concentration. TNF-α mRNA expression level was measured by qPCR. **P < .01, using Student t test, compared with day 0. (D) Microarray data revealed that about 400 genes were upregulated and 100 genes were downregulated by more than 2-fold in BART/THP-1 cells relative to both control cells treated with doxycycline (control) and cells treated with empty vector (Empty). (E) Microarray results were validated by RT-qPCR. Expression of ARG1 and RILP were upregulated by induction of BART miRNAs. Expression of CIITA was downregulated by induction of BART miRNAs. Expression levels were normalized to GAPDH. Averages from 3 independent experiments were shown. *P < .05, Student t test. (F) 3′-UTR luciferase assay was performed with HEK293T cells (left). Either the empty vector or BART miRNA overexpression vector was cotransfected along with the psiCHECKTM-2 vector, a luciferase reporter vector. Ctrl, psiCHECKTM-2 vector; WT, inserted wild-type 3′-UTR sequence of target genes; mut., including mutated target sites. *P < .05, Student’s t-test. Position of the predicted target sequences (miRANDA) of BART miRNAs in the 3′-UTR of human MEF2C mRNAs (right). (G) Downregulation of MEF2C by siRNA treatment enhanced LPS-stimulated expression of IL-10. Expression level was normalized to GAPDH. Three siRNAs were mixed to minimize off-target effects. An average of 3 independent experiments was shown. *P < .05, Student t test.
BART miRNAs alter gene expression in THP-1 cells. (A) BART miRNAs were overexpressed in THP-1 cells with the Tet-Off system. (B) BART miRNA expression was regulated by doxycycline. After 7 to 9 days of culture, the number of cells was counted. Cell proliferation is expressed as the fold-change relative to the cell number at day 0. *P < .05, using Student t test, compared with control values of each doxycycline concentration. (C) Total RNA was collected from BART/THP-1 cells at the indicated time and doxycycline concentration. TNF-α mRNA expression level was measured by qPCR. **P < .01, using Student t test, compared with day 0. (D) Microarray data revealed that about 400 genes were upregulated and 100 genes were downregulated by more than 2-fold in BART/THP-1 cells relative to both control cells treated with doxycycline (control) and cells treated with empty vector (Empty). (E) Microarray results were validated by RT-qPCR. Expression of ARG1 and RILP were upregulated by induction of BART miRNAs. Expression of CIITA was downregulated by induction of BART miRNAs. Expression levels were normalized to GAPDH. Averages from 3 independent experiments were shown. *P < .05, Student t test. (F) 3′-UTR luciferase assay was performed with HEK293T cells (left). Either the empty vector or BART miRNA overexpression vector was cotransfected along with the psiCHECKTM-2 vector, a luciferase reporter vector. Ctrl, psiCHECKTM-2 vector; WT, inserted wild-type 3′-UTR sequence of target genes; mut., including mutated target sites. *P < .05, Student’s t-test. Position of the predicted target sequences (miRANDA) of BART miRNAs in the 3′-UTR of human MEF2C mRNAs (right). (G) Downregulation of MEF2C by siRNA treatment enhanced LPS-stimulated expression of IL-10. Expression level was normalized to GAPDH. Three siRNAs were mixed to minimize off-target effects. An average of 3 independent experiments was shown. *P < .05, Student t test.
Conditional induction of BART miRNAs using doxycycline (Dox) caused cell proliferation and TNF-α upregulation in a dose-dependent manner (Figure 6B-C). These results were consistent with those obtained in PBMCs (Figure 5D-E). Microarray analysis was performed to elucidate the comprehensive changes of gene expression. According to microarray data, about 400 genes were upregulated and about 100 genes were downregulated by more than 2-fold in BART miRNA-expressing cells (BART) relative to both cells treated with Dox (Control) and the cells treated with empty vector (Empty) (Figure 6D). Among the upregulated genes, arginase 1 (ARG1), a tumor-associated macrophage marker, was prominently upregulated by BART miRNAs (supplemental Figure 5A). In addition, the expression levels of several molecules involved in maturation and trafficking of lysosomes, such as ATP6V1B1 and RILP,31,32 were upregulated by BART miRNAs, 1 of the characteristic features of the M2-like macrophage.33 HLA-DR, DM, and CIITA, which is a trans-activator of HLA-class II expression, were downregulated, suggesting the inhibitory role of BART miRNAs on acquired immune responses (supplemental Figure 5B). Among them, upregulation of ARG1 and RILP and downregulation of CIITA were also validated by RT-qPCR (Figure 6E).
In silico prediction identified MEF2C and CD1c as targeted genes by several BART miRNAs (Figure 6F; supplemental Figure 5C), which are a transcription factor involved in the development of various cell lineage and regulation of apoptosis of macrophage and a surface molecule involved in presentation of lipid antigen, respectively.34-38 Downregulation of MEF2C and CD1c by BART miRNAs was validated using a 3′-UTR luciferase reporter assay in HEK293T cells (Figure 6F; supplemental Figure 5C).
To investigate the role of these genes in the macrophage phenotype, we performed a knockdown experiment using small interfering (si)RNA. THP-1 cells were transduced with siRNAs for 48 hours and subsequently stimulated with lipopolysaccharide (LPS). LPS-induced expression of TNF-α and IL-10 was examined by RT-qPCR at early (4 h) and later (20 h) points. Decreased expression of MEF2C (20% to approximately 50%, data not shown) led to the enhanced expression of IL-10 (Figure 6G), consistent with the data in Figure 5E. Expression of TNF-α was not affected by MEF2C, suggesting that other targeted molecules are involved in the regulation of TNF-α. The collective results suggest that BART miRNAs could regulate gene expression, which involves in immune regulation in THP-1 cells.
Amount of tumor-produced BART miRNAs correlates with clinical outcomes in EBV+ DLBCL of elderly patients
Finally, to investigate the significance of BART miRNA expression in EBV-induced lymphomagenesis, BART13, a BART miRNA that is abundantly expressed in L591 cells derived from patients with EBV+ lymphoma, was analyzed in 13 EBV+ DLBCL biopsies obtained from elderly patients by fluorescent in situ hybridization (Figure 7A). This result was also obtained by RT-qPCR in the representative cases (supplemental Figure 6).
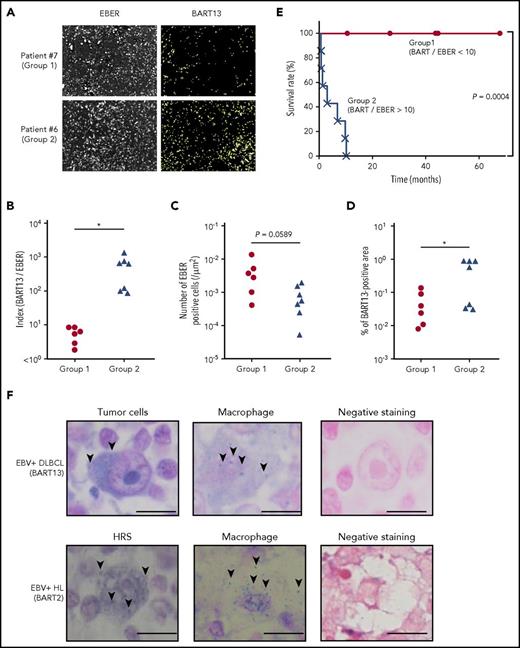
Amounts of tumor-produced BART miRNAs in EBV+DLBCL in elderly patients. (A) EBV-encoded noncoding RNA (EBER) served as a highly sensitive marker to detect EBV infection. EBER and BART13 were detected from DLBCL biopsy samples by fluorescence in situ hybridization. Representative results are shown. The definition of group 1 (n = 6) and 2 (n = 7) is indicated in B. (B) Comparison of the index of BART13+ area/number of EBER+ cells. Group 1, BART/EBER <10; gGroup 2, BART/EBER >10. *P < .05. (C-D) The numbers of EBER+ cells and BART13+ cell areas were automatically counted. Comparison of the number of EBER+ cells (C) and BART13+ cell areas (D) in the tissue area on a slide. *P < .05. (E) Survival ratio of patients with DLBCL. *P = .0004, log-rank test. (F) BART miRNAs were detected from EBV+ DLBCL (top) and EBV+ Hodgkin’s lymphoma (bottom) biopsy samples by in situ hybridization. (Left) Tumor cells; (middle) macrophages; (right) negative staining (using scramble miRNAs for the probe). (Top) BART-13 was detected; (bottom) BART-2. Representative BART miRNA signals (blue dots) are indicated by arrows. Scale bar, 10 μm.
Amounts of tumor-produced BART miRNAs in EBV+DLBCL in elderly patients. (A) EBV-encoded noncoding RNA (EBER) served as a highly sensitive marker to detect EBV infection. EBER and BART13 were detected from DLBCL biopsy samples by fluorescence in situ hybridization. Representative results are shown. The definition of group 1 (n = 6) and 2 (n = 7) is indicated in B. (B) Comparison of the index of BART13+ area/number of EBER+ cells. Group 1, BART/EBER <10; gGroup 2, BART/EBER >10. *P < .05. (C-D) The numbers of EBER+ cells and BART13+ cell areas were automatically counted. Comparison of the number of EBER+ cells (C) and BART13+ cell areas (D) in the tissue area on a slide. *P < .05. (E) Survival ratio of patients with DLBCL. *P = .0004, log-rank test. (F) BART miRNAs were detected from EBV+ DLBCL (top) and EBV+ Hodgkin’s lymphoma (bottom) biopsy samples by in situ hybridization. (Left) Tumor cells; (middle) macrophages; (right) negative staining (using scramble miRNAs for the probe). (Top) BART-13 was detected; (bottom) BART-2. Representative BART miRNA signals (blue dots) are indicated by arrows. Scale bar, 10 μm.
The index of BART13+ area/number of EBER+ cells (BART/EBER), indicating the amount of BART13 production in EBER+ lymphoma cells, significantly differed between the 2 groups (Figure 7B). The number of EBER+ cells did not significantly differ between the groups (Figure 7C). In contrast, the proportion of the BART13+ area relative to the negative area was higher in samples of group 2 patients (Figure 7D). Surprisingly, patients with low BART/EBER indices (<10; group 1) had higher survival rates compared with those with high BART/EBER indices (>10; group 2; Figure 7E). Overall, our results demonstrated that the amount of BART miRNA produced by EBV+ lymphoma cells differed between the 2 groups, suggesting that BART miRNAs play critical roles in tumorigenesis.
Furthermore, in EBV+ DLBCL biopsy samples, BART13 was detected in EBER+ lymphoma cells with large nuclei, and also in a few macrophage cells with small nuclei and large volume of cytoplasm (Figure 7F, upper). In addition, expression of the BART2 miRNA was detected in morphologically macrophages with small nuclei and large cytoplasm, as well as in Hodgkin/Reed-Sternberg cells with large nuclei in EBV+ Hodgkin’s lymphoma biopsy (Figure 7F, lower).
These results suggest that BART miRNAs derived from EBV+ tumor cells are likely to be transferred from lymphoma cells to non-EBV-infected macrophages in EBV-related lymphomas.
Discussion
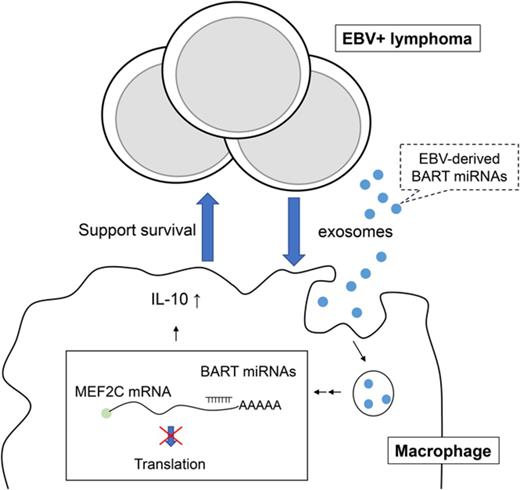
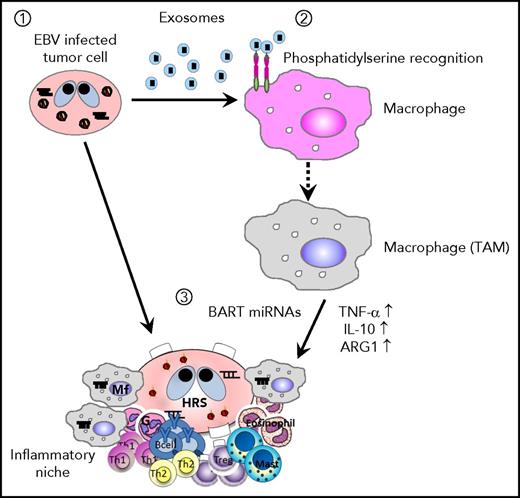
We demonstrated that EBV+ lymphoma cells secrete exosomes to support tumor cell survival through the regulation of inflammatory responses in macrophages both in vivo and in vitro (Figure 8). In detail, we found that: exosomes are an important factor for the formation of EBV+ lymphoma in a humanized mouse model comparing the Akata and B95-8 strains; exosomes collected from lymphoma cells could regulate the activity of macrophages and induce the immune regulatory phenotype in vitro characterized by the enhanced expression of TNF-α, IL-10, and ARG1, which were partly regulated by BART miRNAs; and the amount of lymphoma-produced BART miRNAs correlates with clinical outcomes in EBV+ DLBCL in elderly patients.
Scheme of EBV+B-cell lymphoma microenvironment establishment. (1) EBV-infected cells release exosomes containing EBV-miRNAs, which are incorporated into macrophages. (2) Lymphoma-derived exosomes alter gene expression and convert the macrophages into “tumor associated macrophages.” (3) Accumulation of BART miRNAs and upregulation of tumor-supporting molecules, TNF-α, IL-10, and ARG1, enhance the development of EBV+ B-cell lymphoma.
Scheme of EBV+B-cell lymphoma microenvironment establishment. (1) EBV-infected cells release exosomes containing EBV-miRNAs, which are incorporated into macrophages. (2) Lymphoma-derived exosomes alter gene expression and convert the macrophages into “tumor associated macrophages.” (3) Accumulation of BART miRNAs and upregulation of tumor-supporting molecules, TNF-α, IL-10, and ARG1, enhance the development of EBV+ B-cell lymphoma.
Since Valadi et al reported that exosomes contain abundant miRNAs and transfer them from cell to cell,12 several studies have revealed the physiological significances and roles of exosomes in the development of diseases, especially in the tumor development and metastasis.11-15 Although it has been already reported that EBV+ lymphoma cells secrete exosomes, their roles in the development of lymphoma remained to be elucidated.10,16 In this study, we demonstrate the significant role of tumor-derived exosomes for the development of EBV+ lymphoma, using a humanized mouse model.
CD163+ macrophages, which are thought to be M2-like macrophages, were completely eliminated in all 6 mice by clodronate liposomes, whereas CD68+ macrophages were not (Figure 3). One possible explanation for this result is that M2 macrophages, which support tumor growth, are more sensitive to clodronate than M1 macrophages, which attack and eliminate tumors.39,40 Accordingly, the selective elimination of CD163+ macrophages may alter the tumor microenvironment, making it more difficult for EBER+ tumor cells to survive than that caused when both CD163+ and CD68+ macrophages are eliminated by clodronate liposomes.
In vitro studies demonstrated that exosomes were mainly incorporated into monocytes/macrophages. Incorporation of exosomes by monocytes increased in dose- and time-dependent manners, whereas that by lymphocytes was transiently detected in only high-dose treatment, suggesting that the activity of monocytes could be mainly regulated by tumor-derived exosomes, but that of lymphocytes might possibly still be regulated in the local environment in vivo. We evaluated the overlapping mechanism of the engulfment of apoptotic cells and intake of the tumor-derived exosomes by monocytes/macrophages. To maintain homeostasis, phagocytes engulf dead cells expressing the eat me signal. Considering previous results indicating that PS is exposed by both apoptotic cells and exosomes,16,41-43 and on the basis of our results, the mechanism of macrophage-specific uptake of tumor-derived exosomes can be considered to overlap with that of engulfment of apoptotic cells by macrophages through the eat me signal. However, the apoptotic cells engulfed by monocytes/macrophages should enter the lysosomal pathway, whereas the exosome should not. Hence, an exosome-specific mechanism for incorporation after the eat me signal pathway may exist.
In this study, we focused on the delivery of BART miRNAs by exosomes. Pegtel et al reported that BART miRNAs in the exosome can be transferred into monocyte-derived dendritic cells.10 Similarly, we detected BART miRNAs in monocytes treated with lymphoma-derived exosomes by next-generation sequencing and RT-qPCR (supplemental Figure 7). Importantly, B95-8 has a deletion in the BART region and lacks the expression of the majority of BART miRNAs, which is thought as the most significant difference between the Akata and B95-8 strains. Exosomes collected from EBV+ lymphoma cells have also been reported to carry EBER and drive antiviral responses in dendritic cells.16 Thus, we compared the expression level of EBER1 between exosomes collected from Akata- and B95-8-LCL by RT-qPCR. Although the expression level of EBER1 was not significantly different between exosomes collected from Akata- and B95-8-LCL (supplemental Figure 4B), its general inflammatory function could work in the phenotype caused by the exosomes derived from Akata-infected cells.
In addition to BART miRNAs, the Akata and B95-8-strains might differentially induce the expression of EBV-coding genes in the infected cells.28 Therefore, different gene expression in tumor cells might be reflected in exosomes secreted from Akata- and B95-8-infected tumor cells. Although the expression of LMP-1 and EBNA3B, which is reportedly involved in the transformation and immortalization of B cells,3,4,44 were not significantly different between Akata- and B95-8-LCL (supplemental Figure 4A,C), we could not exclude the possibility that the different expression of these genes and components in exosomes affects tumorigenesis. Treatment with exosomes containing high levels of BART miRNAs enhanced survival and activation in monocytes/macrophages. In particular, the enhanced expression of TNF-α and IL-10 was a characteristic feature of the immune regulatory phenotype. Furthermore, inducible expression of BART miRNAs in THP-1 also showed similar results, supporting the idea that BART miRNAs could regulate inflammatory responses in macrophages and induce the immune regulatory phenotype. Although treating monocytes with exosomes in vitro for 6 hours induced the production of TNF-α, the production of IL-10 was not induced, regardless of elevated mRNA level (data not shown; Figure 5). This discrepancy could be caused by the different kinetics of translation between TNF-α and IL-10. Production of IL-10 in vivo was elevated in the spleens of Akata-infected mice compared with B95-8-infected mice (Figure 1D), suggesting that the long-time exposure to exosomes produces IL-10 in macrophages.
In silico prediction and 3′-untranslated region (UTR) luciferase assay determined MEF2C as a target gene of BART miRNAs, which is a transcription factor involved in the development of various cell lineages and apoptosis of macrophages.34-36 Downregulation of MEF2C by siRNA enhanced the LPS-stimulated IL-10 expression in THP-1, indicating the regulatory role in inflammatory responses. In contrast, expression of TNF-α and ARG1 was not affected by downregulation of MEF2C. Production of ARG1 in vivo was detected only in 1 of 3 Akata-infected mice and no B95-8-infected mice (supplemental Figure 8), suggesting that other molecules were involved in the expression of these genes. Because exosomes can transfer not only miRNAs but also various biomolecules, such as proteins and lipids,45 it is possible that these molecules might synergistically induce the macrophage phenotype with BART miRNAs. Although downregulation of MEF2C was insufficient in monocytes treated with exosomes for 48 hours (data not shown), elevated production of IL-10 in Akata-infected mice suggests that longer exposure to exosomes is needed for the induction of IL-10-producing macrophages in vivo (Figure 1D).
The roles of BART miRNAs in tumorigenesis are complicated. It has been reported that BART miRNAs suppress apoptosis by inhibiting Bim, a pro-apoptotic protein, and LMP1.46,47 In addition, an in vivo xenograft model of EBV-driven carcinogenesis using nasopharyngeal carcinoma cells demonstrated that the BART miRNAs potentiate tumor growth and development in epithelial cells.48 In contrast, Lin et al reported that BART miRNAs suppress B-cell tumorigenesis by inhibiting BZLF1.49 We think these opposite functions of BART miRNAs reflect the use of EBV strain M81. M81 was isolated from a patient with nasopharyngeal carcinoma and has a strong tropism to epithelial cells,50 which basically differs from strains Akata and B95-8. Furthermore, M81 contains the mutation in BZLF1, which makes the infected cells constitutively active in replication instead of formation of latent infection. Once Akata and B95-8 infect the cells, they establish a markedly more latent infection in these cells.51 Consistently, production of BZLF1 was detected in neither the spleens of Akata- nor B95-8-infected mice in our model (data not shown), suggesting that the different tumorigenic activity between the 2 strains was not related to levels of BZLF1.
Analysis of BART miRNAs expression in EBV+ DLBCL of elderly patients indicated that the potency of the production of BART miRNAs by EBV+ tumor cells was significantly correlated with clinical outcomes (Figure 7A-E). Although the differences in the biology and pathogenesis between the 2 groups should be further investigated, the production of BART miRNAs by tumor cells may be useful for the diagnosis of the 2 clinical groups of EBV+ DLBCL in elderly patients. Recently, the expression of EBV-coding miRNAs in several EBV-related lymphomas was investigated by next-generation sequencing.52 The expression profile was quite different among the types of lymphoma. EBV+ DLBCL of the elderly significantly expressed BART 13-3p, which is consistent with our results of in situ hybridization and RT-qPCR, using lymphoma sections (Figure 7A-D; supplemental Figure 6).
It is expected that viral miRNAs could be easily distinguished from endogenous miRNAs,53 which may lead to fewer adverse effects of the therapy. Furthermore, it is reported that some BART miRNAs share seed sequence homology among themselves,54 suggesting that targeting the seed sequences of BART miRNAs may be more efficient. However, at present, further clinical information is required; for example, which miRNAs are strongly expressed in lymphoma tissue, and the generality of their expression among patients. Further studies are warranted to determine the efficient and effective strategy to target BART miRNAs for the therapy of EBV+ lymphomas.
Methods
EBV
Akata and B95-8 strains were prepared as described previously.55 Virus production by EBV-infected Akata cells was stimulated by the brief treatment with anti-IgG antibody (Dako, Carpinteria, CA), and the culture fluid was used as the inoculum after filtration through a 0.45-µm membrane filter.56
For virus titration, cord blood lymphocytes were plated at a density of 2 × 105 cells per well in 6-well plates and then inoculated with serial 10-fold dilutions of the virus preparation. The number of wells containing proliferating lymphocytes was counted 6 weeks after infection, and the titer of the virus in 50% transforming dose was determined using the Reed-Muench method (supplemental Figure 1).
Humanized mice
NOG mice were purchased from the Central Institute for Experimental Animals (Kanagawa, Japan). The detailed method of humanization of NOG mice has been described previously.17 In brief, 1 × 105 CD34+ cord blood cells were administered intravenously. After about 3 months, peripheral blood was collected from the mice, and human CD21 expression was checked for humanization. All experiments were approved by the Institutional Review Board of Tokai University. The animals received humane care as required by the institutional guidelines for animal care and treatment in experimental investigations.
Mouse experiments
Akata, B95-8, EBV miRNA-rich Akata exosomes, and EBV-miRNA-deleted B95-8 exosomes were injected intravenously into the hematopoietically humanized NOG mice. When the animal weight reached the determined value, the mice were killed, and histological analysis was performed on the obtained spleens. Clodronate liposome injection trial was performed with EBV-infected mice. Mice were intraperitoneally injected with clodronate- or phosphate buffered saline-liposomes (ClodronateLiposomes.org, Amsterdam, The Netherlands).
EBV viral load determination by qPCR
Total DNA was extracted from the whole blood of EBV-infected mice with the DNeasy Blood and Tissue kit (QIAGEN, Hilden, Germany), according to the manufacturer’s instructions. EBV DNA loads were determined by the qPCR assay. The PCR was performed with the THUNDERBIRD SYBR qPCR Mix (Toyobo, Osaka, Japan), following the manufacturer’s protocol. The primers used for detection of B95-8 were 5′-ATCGACGTATCGCTGGAAAC-3′ and 5′-AGTCCTGATCGTCCTCCTC-3′. The primers used for detection of Akata were 5′-ACACCAAGATCACCACCCTC-3′ and 5′-TTAGGGTGCCACATCCTGTTC-3′.
BART miRNA expression analysis
RNAs were isolated from cells or exosomes using Sepasol-RNA I Super G (Nacalai Tesque, Kyoto, Japan). Reverse transcription was performed with the TaqMan MicroRNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA). qPCR was performed with TaqMan Fast Universal PCR Master Mix, using StepOnePlus Real-Time PCR System (Applied Biosystems). The sequences of primers and probes were described previously.10 RNA isolation, reverse transcription, and qPCR were performed following each manufacturer’s protocol.
Cells
Akata and Daudi cells from EBV+ patients with Burkitt lymphoma, B95-8 cells from EBV-transformed marmoset lymphocytes, THP-1 cells from a patient with acute monocytic leukemia, and LCLs (Akata-LCL, B95-8-LCL, and X50-7) from an EBV-transformed umbilical cord blood B lymphoblast line were cultured in RPMI 1640 medium (Nacalai Tesque) supplemented with 10% (vol/vol) fetal bovine serum (FBS), 100 U/mL penicillin, and 100 μg/mL streptomycin (Life Technologies, Carlsbad, CA) in 50-mL flasks.
Lentiviral transfer
High-titer lentiviral supernatant was observed after cotransfection of a miRNA expression vector and pCAG-KGP1R for gag protein, pCAG-4RTR2 for Rev/tat, and pCMV-VSVG viral packaging construct into 293T cells using ExtremeGENE (Roche Diagnostic, Basel, Switzerland). The cells were plated at a concentration of 4 to 5 × 105/mL in a 24-well plate, and spin-infected with the desired lentiviral supernatant for 2 hours.
Exosome isolation
Akata-LCL, B95-8-LCL, Daudi, and X50-7 were incubated in RPMI-1640 medium supplemented with 10% FBS (vol/vol), and cell supernatants were harvested. Exosomes were collected from the supernatants by ultracentrifugation at 110 000 × g for 70 minutes.21 Collected exosomes were labeled with PKH26 dye (SIGMA-ALDRICH, Saint Louis, MO), following the manufacturer’s protocol.
Exosome fractionation
In some experiments, exosomes were separated by density gradient centrifugation after ultracentrifugation, as reported previously.26 Briefly, 10%, 20%, and 30% Iodixanol solutions were prepared by mixing Optiprep (Axis-Shield, Oslo, Norway) with buffer containing 0.25 M sucrose, 10 mM Tris at pH 8.0, and 1 mM EDTA, final pH set to 7.4. Exosome pellets were resuspended in 3 mL of 30% Iodixanol solution. Subsequently, 1.3 mL of 20% and 1.2 mL of 10% Iodixanol solutions were carefully layered on top of the suspension.
Constructs
Genomic DNA from L591, derived from an EBV+ patient with Hodgkin’s lymphoma, was extracted using the DNeasy Tissue extraction kit (QIAGEN). The genomic sequences of the segments in BART cluster 1 and 2 were amplified by genomic PCR, using Pfx polymerase (Invitrogen, Carlsbad, CA). The oligonucleotide sequences used for PCR were Bartcluster1-NotI-F, 5′gsstgcggccgcattgctcaggccaaagtt3′; Bartcluster1-BamH1-R, 5′gaatggatcctgaaacccaagtttccttgc3′; Bartcluster2-NotI-F, 5′aattgcggccgctgccattattcccttga3′; and Bartcluster2-NotI-R, 5′aattgcggccgcattaacgggggaaggaa3′.
The PCR products were purified with a PCR purification kit (QIAGEN) and sequenced. The purified genomic PCR products of BART cluster 1 and cluster 2 were serially cloned into the mammalian lentiviral expression vector, pCS2-Ires-GFP.
For the Tet-off inducible system, we used the Lenti-X Tet-Off Advanced Inducible Expression System (Clontech, Mountain View, CA) and the associated products.
Flow cytometry analysis of exosome-treated PBMCs
The supernatants of EBV-infected cells were harvested. Exosomes were isolated from these supernatants and added to PBMCs. CD14 and CD69 expressions on the cell surface were detected by flow cytometry. Pacific Blue anti-human CD14 antibody (clone: HCD14, Biolegend, San Diego, CA) and PE/Cy7 anti-human CD69 antibody (clone: FN50, Biolegend) were used.
Quantitative PCR
Total RNAs of PBMCs or THP-1 cells were isolated using Sepasol-RNA I Super G. RT-PCR was performed with the High-Capacity Reverse Transcription Kit (Applied Biosystems), and qPCR was performed with the THUNDERBIRD SYBR qPCR Mix, following each manufacturer’s protocol. The primers were as follows:
IL-10 5′-CGGCGCTGTCATCGATTT-3′
5′-GAGTCGCCACCCTGATGTCT-3′
TNF-α 5′-TCTGGCCCAGGCAGTCAGATCAT-3′
5′-CGGCGGTTCAGCCACTGGAG-3′
ARG1 5′-GGAAACTTGCATGGACAACC-3′
5′-TCCACGTCTCTCAAGCCAAT-3′
RILP 5′-CAAGATGTTAGGGACACCAGAG-3′
5′-CAGCTTTACCCCGATACCATAG-3′
CIITA 5′-AGCTGAAGTCCTTGGAAACC-3′
5′-CGTCGCAGATGCAGTTATTG-3′
MEF2C 5′-TGTAACACATCGACCTCCAAG-3′
5′-TGTTCAAGTTACCAGGTGAGAC-3′
EBER1 5′-AGCACCTACGCTGCCCTAGA-3′
5-AAAACATGCGGACCACCAGC-3′
EBNA3B 5′-CCACGCTGTCTATGATTCCA-3′
5′-CGATGTTCAGGTTTTGCTCA-3′
GAPDH 5′-CTGCACCACCAACTGCTTAG-3′
5′-TTCAGCTCAGGGATGACCTTG-3′
Western blotting
Protein concentration in exosomes was measured by absorbance at 280 nm. Proteins were separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis and transferred onto polyvinylidene fluoride membranes. The membranes were exposed to mouse anti-LMP1 antibody (ab78113, clone: CS 1-4; Abcam, Cambridge, UK) or mouse anti-CD63 antibody (sc-5275, clone: MX-49. 129. 5; Santa Cruz Biotechnology, Santa Cruz, CA). The protein signals were detected using a ChemiDoc Touch System (BIO-RAD, Hercules, CA).
THP-1 cell functional assay
BART clusters 1 and 2 were overexpressed in THP-1 cells with the Tet-Off system (BART/THP-1). BART miRNA expression was regulated by doxycycline. Ctrl/THP-1 and BART/THP-1 cells were normally cultured with 10% FBS RPMI medium with 1 μg/mL doxycycline. For proliferation assay, cells were precultured with 4 μg/mL doxycycline and cultured with 0, 2, or 4 μg/mL doxycycline in 0.1% FBS RPMI medium. After 7 to 9 days of culture, the number of cells was counted. Cell proliferation is expressed as fold change relative to the cell number on day 0. Total RNA was collected from BART/THP-1 cells at the time and doxycycline concentration indicated in Figure 6.
3′-UTR luciferase assay
3′-UTR luciferase assay was performed in HEK293T cells. Either the empty vector or the BART miRNA overexpression vector was cotransfected, along with the psiCHECK-2 vector (Promega, Madison, WI), using X-tremeGENE (Roche). The wild-type 3′-UTR sequence of each gene was inserted into the vector. For the mutation experiment, we introduced a 3-base change in the wild-type sequence, predicted as a BART miRNA target site by MiRanda, using the Site-Directed Mutagenesis kit (New England Biolabs, Ipswich, MA). The Dual-Luciferase Reporter Assay System (Promega) was used for detection, following the manufacturer’s protocol. Renilla luciferase activity was calibrated with firefly luciferase activity.
siRNA transfection
siRNA was transfected by Neon electroporation system (Invitrogen) following the manufacturer’s protocol. In brief, 1 × 105 THP-1 cells were resuspended in 10 μL buffer containing 5 μM siRNA. Electroporation was performed under the condition with 1300 V, 10 milliseconds of pulse width and 3 pulses. Predesigned siRNA against human MEF2C was purchased from Bioneer Corporation (Daejeon, Korea). To minimize the off-target effects, 3 different siRNAs were mixed and introduced into the cells simultaneously.
Patients and specimens
The Institutional Review Board of Tokai University, School of Medicine, approved this study, and all human samples were handled accordingly. The biopsy samples of patients with DLBCL and the patient prognosis information were provided by Sato, Kashiwagi, Matsui, Okamoto, and Nakamura.
In situ hybridization of BART miRNAs
In situ hybridization was performed as described previously.57 We purchased the following 5′ digoxigenin-labeled locked nucleic acid probes (miRCURY LNA Detection probe, Exiqon, Demmark): ebv-miR-BART2-5p, 5′-(digoxigenin)GCAAGGGCGAATGCAGAAAATA-3′; ebv-miR-BART13, 5′-(DIG)TCAGCCGTCCCTGGCAAGTTACA-3′. Slides were photographed using a BZ-X700 All-in-1 fluorescence microscope (Keyence, Osaka, Japan). The threshold values were determined by our observations. Each positive signal decision was then made automatically, using Keyence software.
DNA microarray
BART/THP-1 cells and ctrl/THP-1 cells (with empty vector) were cultured without doxycycline for 4 days, and total RNA was harvested. Reverse-transcribed samples were hybridized to Whole Human Genome DNA microarray 4 × 44K (Agilent Technologies, Santa Clara, CA) and analyzed according to the manufacturer’s protocol.
Statistics
Statistical significance was determined using Student t test or Mann-Whitney U-test. P < .05 was considered statistically significant. Survival curves were analyzed with GraphPad Prism software (GraphPad Software, Inc., La Jolla, CA), using the log-rank test.
Study approval
All animal experiments were approved by the Institutional Review Board of Tokai University. The animals received humane care, as required by the institutional guidelines for animal care and treatment in experimental investigations. The Institutional Review Board of Tokai University, School of Medicine, approved this study, and all human samples were handled accordingly. The participants were identified by number.
The online version of this article contains a data supplement.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.
Acknowledgments
The authors thank Kayoko Iwao, Tomoko Uno, Yukie Y. Kikuchi, Masayuki Tanaka, Hideki Hayashi, and the Support Center for Medical Research and Education, Tokai University, for technical assistance; Kenzo Takada for his kind gift of EBV-infected cell lines; Mamoru Ito for supplying the material; and Junko Sasaki for her critical advice regarding the in vivo experiments.
This study was supported by Research and Study Program of Tokai University Educational System General Research Organization (H. Higuchi); Tokai Scholarship Award (N.Y.); and Precursory Research for Embryonic Science and Technology, AMED-PRIME, and the Research Program on Hepatitis from Japan Agency for Medical Research and Development (16fk0210114h0001; A.K.).
Authorship
Contribution: H. Higuchi, N.Y., K.-I.I., T.Y., R.K., J.O., M. Kakizaki, K.F., J.L., K.Y., K.O., A.S., M.T., N.K., S.M.A., A.A., T. Kitamura, T. Kashiwagi, T.M., A.O., H. Handa, M. Kuroda, N.N., and A.K. performed the experiments described in this manuscript; R.H. and T.W. provided precious materials used in this study; T. Kashiwagi, T.M., A.O., K.A., and N.N. collected patient samples; and H. Higuchi, N.Y., K.-I.I., and A.K. wrote the manuscript.
Conflict-of-interest disclosure: A.K. received consultant fees from Janssen Pharmaceutical K.K. The remaining authors declare no competing financial interests.
Correspondence: Ai Kotani, 143 Shimokasuya, Isehara-shi, Kanagawa 259-1193, Japan; e-mail: aikotani@k-lab.jp.
Reference
Author notes
H.H., N.Y., and K.-I.I. contributed equally to this study.