Abstract
Treatment outcome in acute lymphoblastic leukemia (ALL) has improved over the past 30 years, with overall survival rates of ∼45% in adults and ∼85% in children. Gross cytogenetic abnormalities, including numerical changes and chromosomal translocations, are of considerable prognostic value in both pediatric and adult ALL. In addition, we and others have recently identified novel molecular markers associated with a poor outcome in ALL, including deletions of the lymphoid transcription factor IKZF1. In order to identify downstream signaling events associated with these genetic alterations, we performed an integrated analysis of genomic abnormalities, including copy number alterations, sequence mutations and chromosomal translocations, with alterations in protein expression and modification.
A cohort of 91 precursor B-ALL cases treated at M.D. Anderson Cancer Center in Houston, USA, including 82 newly diagnosed cases and 5 diagnosis-relapse pairs was used for this study. The cohort consisted of 6 children (age 1-6), 30 young adults (age 15-39) and 45 adults (age>39), and 20 patients carried a BCR-ABL1 chromosomal translocation. Copy number alterations in eight genes frequently deleted in ALL (IKZF1, PAX5, EBF1, RB1, CRLF2, CDKN2A/2B, BTG1, and ETV6) were determined by multiplex ligation-dependent probe amplification analysis. IKZF1 deletions were associated with relapse (Pearson's chi-square test, p=0.009), and the presence of BCR-ABL1 translocation (p=0.032). Protein expression and modification levels were determined by probing Reverse Phase Protein Arrays (RPPA) containing protein lysates of all above samples with 128 rigorously validated antibodies including 34 phospho-specific antibodies. Hierarchical clustering analysis was used to determine which (phospho)proteins are differently expressed in genetic subsets of ALL. The significance of correlations was determined using two-sample t-test, with correction for multiple testing (Beta-Uniform Mixture model).
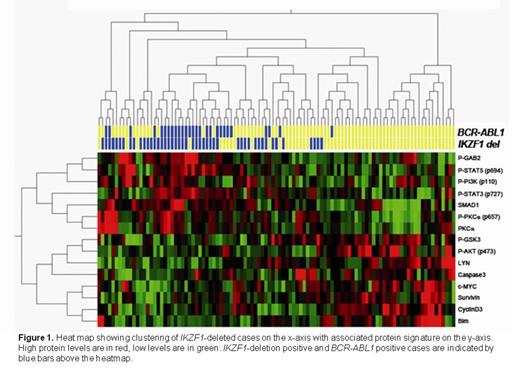
We identified clustering of cases with a BCR-ABL1 chromosomal translocation (p=0.01; false discovery rate (FDR)=0.1), IKZF1-deletions (p=0.01, FDR=0.072), RB1-deletions (p=0.03, FDR=0.43) and EBF1 deletions (p=0.05, FDR=0.63). As expected RB1 deletion positive cases were characterized by decreased levels of (phospho)-RB1 and increased levels of cyclin E, illustrating the validity of our approach. EBF1-deleted cases showed relatively high levels of SHIP1, SSBP2 and phospho-STAT5, and lower levels of FAK and LYN. The protein signatures of BCR-ABL1-positive cases and IKZF1-deletion positive cases largely overlapped, and were characterized by elevated levels of (phospho)PKCα, SMAD1, phospho-STAT3, and phospho-STAT5 and lower levels of LYN and cyclinD3 (Figure 1). In total 70% of the BCR-ABL1-positive cases carried an IKZF1 deletion and several BCR-ABL1-negative cases with similar RPPA signature could be identified, all of which were IKZF1-deletion positive. These cases may represent the “BCR-ABL1-like” cases that were previously identified using gene expression signatures (Mullighan et al. 2009, NEJM 360:470-480; Den Boer et al. 2009, Lancet Oncol. 10:125-134), and could reflect activation of cAbl or other cellular tyrosine kinases. Together, we conclude that integrated analysis of genetic and proteomic aberrations identified protein signatures downstream of recurrent mutational events in ALL, a strategy that promises to facilitate the discovery of novel therapeutic targets in ALL and may aid in the identification of (high risk) patients that would benefit from tyrosine kinase inhibition.
No relevant conflicts of interest to declare.
Author notes
Asterisk with author names denotes non-ASH members.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal