Abstract
To search for genes that promote hematopoietic development from human embryonic stem cells (hESCs) and induced pluripotent stem cells (iPSCs), we overexpressed several known hematopoietic regulator genes in hESC/iPSC-derived CD34+CD43− endothelial cells (ECs) enriched in hemogenic endothelium (HE). Among the genes tested, only Sox17, a gene encoding a transcription factor of the SOX family, promoted cell growth and supported expansion of CD34+CD43+CD45−/low cells expressing the HE marker VE-cadherin. SOX17 was expressed at high levels in CD34+CD43− ECs compared with low levels in CD34+CD43+CD45− pre-hematopoietic progenitor cells (pre-HPCs) and CD34+CD43+CD45+ HPCs. Sox17-overexpressing cells formed semiadherent cell aggregates and generated few hematopoietic progenies. However, they retained hemogenic potential and gave rise to hematopoietic progenies on inactivation of Sox17. Global gene-expression analyses revealed that the CD34+CD43+CD45−/low cells expanded on overexpression of Sox17 are HE-like cells developmentally placed between ECs and pre-HPCs. Sox17 overexpression also reprogrammed both pre-HPCs and HPCs into HE-like cells. Genome-wide mapping of Sox17-binding sites revealed that Sox17 activates the transcription of key regulator genes for vasculogenesis, hematopoiesis, and erythrocyte differentiation directly. Depletion of SOX17 in CD34+CD43− ECs severely compromised their hemogenic activity. These findings suggest that SOX17 plays a key role in priming hemogenic potential in ECs, thereby regulating hematopoietic development from hESCs/iPSCs.
Key Points
SOX17 plays a key role in priming hemogenic potential in endothelial cells during hematopoietic development from ES cells.
Introduction
During mammalian development, 2 waves of hematopoiesis occur in sequential stages: first, a transient wave of primitive hematopoiesis, followed by definitive hematopoiesis. These stages are temporally and anatomically distinct and involve unique cellular and molecular regulators. The formation of primitive blood cells occurs early during fetal life, with coordinated progression from extraembryonic to intraembryonic sites of hematopoiesis. Within the embryo, definitive hematopoiesis undergoes developmentally stereotyped transitions; hematopoietic stem cells (HSCs) arising from the aorta-gonad-mesonephros region migrate first to the placenta and fetal liver and then to the spleen. Eventually, hematopoiesis shifts to the BM, where homeostatic blood formation is maintained postnatally.1
During definitive fetal hematopoiesis, HSCs emerge directly from a small population of endothelial cells (ECs) in the conceptus, referred to as the “hemogenic endothelium” (HE).2-4 HE is located in all sites of HSC emergence, including the ventral aspect of the dorsal aorta, vitelline and umbilical arteries, yolk sac, and placenta. The process by which blood forms from HE involves an endothelial-to-hematopoietic cell transition during which individual cells bud out and detach from the endothelial layer.2-4 HE is distinguished from all other ECs by the presence of a transcription factor called Runx1.5 Runx1 is expressed in HE cells, in newly formed hematopoietic cell clusters, and in all functional HSCs.6,7 A similar process occurs during hemangioblast differentiation in primitive blood cell formation. The extraembryonic yolk sac is considered to be the first site of emergence of the “hemangioblast,” a mesodermal precursor with both endothelial and hematopoietic potential. Hemangioblasts differentiate into a HE intermediate, which gives rise to primitive hematopoietic cells but also definitive hematopoietic cells on activation of Runx1.8
Human embryonic stem cells (hESCs) and induced pluripotent stem cells (iPSCs) have been demonstrated to reproduce many aspects of embryonic hematopoiesis in stromal coculture or embryoid body (EB) culture. A recent study has provided evidence that hematopoietic differentiation of hESCs progresses through sequential stages: first is the HE, then primitive hematopoiesis, and finally definitive hematopoiesis, a process resembling the development of physiologic hematopoiesis.9 However, the induction of hematopoietic cells from hESCs/iPSCs is still inefficient. Significant innovations are required before it will be possible to obtain sufficient numbers of the specific types of hematopoietic cells needed for therapeutic uses.
Sry-related high-mobility group box 17 (SOX17) is a member of the SOX family of DNA-binding transcription factors. Sox17 participates in various developmental processes and biologic activities, such as formation of definitive endoderm10 and vascular development.11 Moreover, recent studies have shown that Sox17 also plays an important role in fetal hematopoiesis in the yolk sac and fetal liver, especially in the maintenance of fetal and neonatal HSCs, but not adult HSCs.12 Overexpression of Sox17 has also been shown to confer fetal HSC characteristics onto adult hematopoietic progenitors.13 Among SOX family members, Sox7, Sox17, and Sox18 are highly related and constitute the Sox subgroup F (SoxF). Sox7 and Sox18 are transiently expressed in hemangioblasts and hematopoietic precursors, respectively, at the onset of blood specification. Sustained expression of Sox7 and Sox18, but not Sox17, in early hematopoietic precursors from mouse ESCs and embryos enhances their proliferation while blocking their maturation.14,15 However, the role of Sox17 in early hematopoietic development, particularly from hESCs, has not yet been clarified. In the present study, we tested the effect of overexpression of known hematopoietic regulator genes in hiPSC-derived CD34+CD43− ECs enriched in HE to find genes that could be manipulated to efficiently produce hematopoietic cells from hESCs. We found that Sox17 promotes the expansion of HE-like cells. We demonstrate that SOX17 functions in HE and plays a role in the development of hematopoietic cells from hESCs/iPSCs.
Methods
Cell lines
H1 hESCs (WiCell Research Institute) and TkCBV4-7 hiPSCs generated from human cord blood (CB) CD34+ cells were maintained on irradiated murine embryonic fibroblasts in DMEM-F12 (Sigma-Aldrich) supplemented with 1× MEM nonessential amino acids (Gibco-Invitrogen), 1× GlutaMAX-I (Gibco-Invitrogen), 20% knockout serum replacement (Gibco-Invitrogen), 0.1mM 2-mercaptoethanol (Sigma-Aldrich), 1% penicillin/streptomycin solution (Sigma-Aldrich), and 5 ng/mL of human basic fibroblast growth factor (ReproCELL). Every 3-4 days, the cells were dissected into clumps of approximately 300-500 cells in a dissociation solution consisting of 0.25% trypsin, 20% knockout serum replacement, and 1mM CaCl2 in PBS and transferred to a new feeder layer to maintain them in an undifferentiated state. The OP9 stromal cell line was kindly provided by Toru Nakano (Osaka University, Osaka, Japan). OP9 cells were maintained in α-MEM (Gibco-Invitrogen) supplemented with 2.2 g/L of sodium bicarbonate, 20% FBS, and 1% l-glutamine and penicillin/streptomycin solution (Sigma-Aldrich).
EB differentiation
H1 hESCs or TkCBV4-7 hiPSCs were dissociated into single cells with Accumax (Innovative Cell Technologies). The cells were washed with DMEM-F12 and recultured at 1 × 106 cells per 60-mm Petri dish (Falcon) in 5-mL mTeSR1 (StemCell Technologies) supplemented with 10μM LY27632 (Cayman), 2 ng/mL of human Bone Morphogenetic Protein 4 (BMP4; PeproTech), and 2 ng/mL of human activin A (PeproTech). At day 2 of culture, EBs were split from 1 60-mm Petri dish to 2 60-mm Petri dishes and cultured in EB medium consisting of IMDM (Sigma-Aldrich) containing 15% FBS, 1× GlutaMAX I, 1% penicillin/streptomycin solution, 200 μg/mL of bovine holo transferrin (Bovogen), 50 μg/mL of ascorbic acid (Sigma-Aldrich), and 450μM 1-thioglycerol (Sigma-Aldrich) supplemented with 2 ng/mL of human BMP4 and 5 ng/mL of human VEGF (PeproTech). At day 4 of culture, medium conditions were changed as described in Figure 1B. LY363947 (Cayman) was used as an inhibitor of TGFβ signaling. EBs were constantly cultured on a shaker at 70 rpm.
Screening of genes that promote expansion of hematopoietic cells from hESCs/hiPSCs. (A) Hematopoietic fractions derived from hESCs in EB culture used in this study. (B) Schematic representation of the protocol modified for efficient induction of pre-HPCs/HPCs from hESCs/hiPSCs in EB culture. (C) Expression of BRACHYURY, RUNX1, and TAL1/SCL expression during differentiation of hESCs in EBs determined by quantitative RT-PCR analysis. mRNA levels were normalized to GAPDH expression. Expression levels relative to that in hESCs (day 0 of EB culture) are shown as the means ± SD for triplicate analyses. (D) Cell growth of CD34+CD43− cells from day 6 EBs and CD34+CD43+ cells from day 8 EBs. EBs were formed by suspension culture of hiPSCs. Sorted cells (2 × 104) were transduced with the indicated hematopoietic regulator genes and cultured on OP9 cells in the presence of 20 ng/mL of SCF and TPO. At day 14 of culture, the absolute numbers of cells were determined and are indicated in bars. Representative data from repeated experiments are shown. (E) Expression of SOX17 during differentiation of hESCs in EBs determined by quantitative RT-PCR analysis. mRNA levels were normalized to GAPDH expression. Expression levels relative to that in hESCs (day 0 of EB culture) are shown as the means ± SD for triplicate analyses. (F) Expression of SOX17, SOX7, and SOX18 in bulk EB cells, CD34+CD43− cells (ECs), CD34+CD43+CD45− cells (pre-HPCs), and CD34+CD43+CD45+ cells (HPCs) from day 8 EBs determined by quantitative RT-PCR analysis. mRNA levels were normalized to GAPDH expression. Expression levels relative to those in CB CD34+ cells are shown as the means ± SD for triplicate analyses.
Screening of genes that promote expansion of hematopoietic cells from hESCs/hiPSCs. (A) Hematopoietic fractions derived from hESCs in EB culture used in this study. (B) Schematic representation of the protocol modified for efficient induction of pre-HPCs/HPCs from hESCs/hiPSCs in EB culture. (C) Expression of BRACHYURY, RUNX1, and TAL1/SCL expression during differentiation of hESCs in EBs determined by quantitative RT-PCR analysis. mRNA levels were normalized to GAPDH expression. Expression levels relative to that in hESCs (day 0 of EB culture) are shown as the means ± SD for triplicate analyses. (D) Cell growth of CD34+CD43− cells from day 6 EBs and CD34+CD43+ cells from day 8 EBs. EBs were formed by suspension culture of hiPSCs. Sorted cells (2 × 104) were transduced with the indicated hematopoietic regulator genes and cultured on OP9 cells in the presence of 20 ng/mL of SCF and TPO. At day 14 of culture, the absolute numbers of cells were determined and are indicated in bars. Representative data from repeated experiments are shown. (E) Expression of SOX17 during differentiation of hESCs in EBs determined by quantitative RT-PCR analysis. mRNA levels were normalized to GAPDH expression. Expression levels relative to that in hESCs (day 0 of EB culture) are shown as the means ± SD for triplicate analyses. (F) Expression of SOX17, SOX7, and SOX18 in bulk EB cells, CD34+CD43− cells (ECs), CD34+CD43+CD45− cells (pre-HPCs), and CD34+CD43+CD45+ cells (HPCs) from day 8 EBs determined by quantitative RT-PCR analysis. mRNA levels were normalized to GAPDH expression. Expression levels relative to those in CB CD34+ cells are shown as the means ± SD for triplicate analyses.
Flow cytometric analysis and OP9 coculture
EBs were dissociated with 0.25% trypsin-EDTA solution (Sigma-Aldrich) and filtered through a nylon screen to obtain a single-cell suspension. Flow cytometric analysis and cell sorting were performed using a FACSAria II cell sorter (BD Biosciences) and the data were analyzed using FlowJo Version 9.5.3 software (TreeStar). The following Abs were used for the flow cytometric analysis: CD34 (clone 581; Alexa Fluor 647 or PE-Cy7), CD43 (clone CD43-10G7; PE), CD45 (clone HI30; PE-Cy7), CD11b (clone M1/70; Brilliant Violet 421), CD235a (clone HIR2; PE), CD144 (VE-Cad; clone 16B1; PE), and CD309 (KDR; clone HKDR-1; APC). Sorted cells were resuspended in hematopoietic medium (IMDM, 10% FBS, 1% l-glutamine and penicillin/streptomycin solution) supplemented with 20 ng/mL of human SCF and 20 ng/mL of human thrombopoietin (TPO; PeproTech), and transferred onto semiconfluent irradiated OP9 cells. For mature hematopoietic cell differentiation, sorted cells were resuspended in HE medium supplemented with 20 ng/mL of SCF, 20 ng/mL of TPO, 10 ng/mL of human IL-3 (PeproTech), and 3 units/mL of human erythropoietin.
Retrovirus and lentivirus vectors, virus production, and transduction
Mouse Sox17 fused to ERT with a 1× or 3× Flag tag was subcloned into the MIG retrovirus vector, which contains the long-terminal repeats from the murine stem cell virus and an internal ribosomal entry site upstream of the enhanced green fluorescent protein (GFP) as a marker gene. A recombinant vesicular stomatitis virus glycoprotein–pseudotyped high-titer retrovirus was generated using a 293gpg packaging cell line.16 The virus containing media from the 293gpg cell cultures was concentrated by centrifugation at 6000g for 16 hours. To knock down SOX17, lentiviral vectors (CS-H1-shRNA-EF-1α-EGFP) expressing shRNA against human SOX17 and luciferase were prepared. Target sequences were as follows; Sh-SOX17#1143; GCATGACTCCGGTGTGAAT, and Sh-SOX17#1273; GGCCAGAAGCAGTGTTACA. The viruses were produced as described previously.17 EBs at approximately day 5-12 of culture were dissociated and the indicated cell populations were sorted using a FACSAria II. Sorted cells were seeded onto semiconfluent irradiated OP9 cells and transduced with a SOX17-ERT retrovirus or a SOX17 knock-down virus. Transduced cells were cocultured with OP9 cells in the presence of the indicated cytokines. To induce nuclear translocation of SOX17-ERT, 4-hydroxy tamoxifen (4-OHT) was added to the medium to a concentration of 200nM on the following day.
Quantitative RT-PCR analysis
Total RNA was extracted using TRIzol reagent according to the manufacturer's instructions (Invitrogen). cDNA was synthesized from total RNA using ThermoScript RT-PCR System (Invitrogen). Quantitative RT-PCR was carried out using FastStart Universal Probe Master (Roche Applied Science), the Universal Probe Library (Roche Applied Science), and the Applied Biosystems 7300 Fast Real-Time PCR system (Applied Biosystems). Primer sequences and probe numbers used are listed in supplemental Methods (available on the Blood Web site; see the Supplemental Materials link at the top of the online article).
Colony-forming assay
Colony assays were performed in methylcellulose (StemCell Technologies) containing IMDM supplemented with 20 ng/mL of human SCF, 10 ng/mL of human IL-3, 10 ng/mL of human TPO, and 3 units/mL of human erythropoietin, and incubated at 37°C in a 5% CO2 atmosphere. The colonies were counted at day 12 of culture. Images were captured by BIOREVO BZ-9000 (KEYENCE) with CFI Plan Fluor ELWD DM 20×C (Nikon) and processed using Adobe Photoshop Elements 4.0.
Gene-expression microarray
Total RNA was extracted using TRIzol reagent according to the manufacturer's instructions (Invitrogen). Purified total RNA was amplified and labeled using the WT expression kit (Ambion) according to the manufacturer's instructions. The labeled samples were hybridized to Human Promoter Gene 1.0 ST GeneChip arrays (Affymetrix) to assess and compare overall gene-expression profiles as described previously.18 Microarray data were submitted to the Gene Expression Omnibus under accession number GSE38156. Expression profiles of the cells were clustered using hierarchical clustering. Distance between 2 samples was defined with the Pearson correlation using all or selected probes. Probes were selected using the Gene Ontology (GO) database or ChIP-on-chip data of Sox17.
ChIP-on-chip experiment
CD34+CD43− cells from EBs at day 6 of culture were seeded on irradiated OP9 cells and transduced with a 3× Flag SOX17-ERT retrovirus. The cells were further cultured on OP9 cells in the presence of SCF, TPO (20 ng/mL), and 200nM 4-OHT. CD34+ cells were collected at day 27 of culture by magnetic cell sorting using magnetic beads conjugated with anti-CD34 Abs (Miltenyi Biotec) and subjected to a ChIP assay using an anti-FLAG Ab (M2, Sigma). ChIP was carried out as described previously.18 ChIP on chip analysis was carried out using the SurePrint G3 Human Promoter Kit, 1 × 1M (G4873A, Agilent Technologies). Purified immunoprecipitated and input DNA was subjected to T7 RNA polymerase-based amplification as described previously.19 Labeling, hybridization, and washing were carried out according to the Agilent mammalian ChIP-on-chip protocol (Version 9.0). Scanned images were quantified with Agilent Feature Extraction software under standard conditions. The assignment of regions bound by SOX17 around transcription start sites (TSSs) was carried out using direct sequence alignment on the human genome database (National Center for Biotechnology Information Version 36). The location of SOX17-bound regions was compared with a set of transcripts derived from the MGI database. Bound regions that were within −8.0 kb to +4.0 kb of the TSS were assigned. Alignments on the human genome and TSSs of genes were retrieved from Ensembl (http://www.ensembl.org). Intensity ratios (IP/input: fold enrichment) were calculated, and the maximum value for each promoter region of a gene was used to represent the fold enrichment of the gene. Fold enrichment was calculated only for probes for which signals both from IP and input DNA were significant (P < 10−3). ChIP-on-chip data were submitted to Gene Expression Omnibus under accession number GSE38156.
GO analysis
GO annotation was obtained using gene2go database (ftp://ftp.ncbi.nih.gov/gene/DATA/gene2go.gz) from Entrez (retrieved January 2012). Human genes were collected from the database and enrichment of SOX17-binding genes was distributed to 2 × 2 contingency tables for all GO terms (having/not having GO and binding/not binding to SOX17). We calculated P for each contingency table using hypergeometric distribution. The P value reflects the likelihood that we would observe the distribution by chance and significant GO terms were selected when P < .001.
Immunostaining
Sox17-ERT–transduced cells were sorted by flow cytometry and cultured on MAS-coated glass slides (Matsunami Glass Industries,) for 4 hours. The cells were then fixed with 2% paraformaldehyde and immunostained with an anti-laminin Ab (ab11575; Abcam) or an anti-FLAG Ab (M2; Sigma-Aldrich) for primary antibody reaction, and an Alexa Fluor 555 goat anti–rabbit IgG (Molecular Probes) or Alexa Fluor 555 goat anti–mouse IgG (Molecular Probes) for secondary antibody reaction, respectively. Images were captured by BIOREVO BZ-9000 (KEYENCE) with CFI Plan ApoVC 100×H (Nikon) and processed using Adobe Photoshop Elements 4.0.
Western blotting
Total cell lysate was resolved by SDS-PAGE and transferred to a PVDF membrane. The blots were probed with an anti-Sox17 Ab (09-038; Millipore) or an anti–α-tubulin (CP06; Calbiochem) and an HRP-conjugated secondary Ab. The protein bands were detected with SuperSignal West Pico Chemiluminescent Substrate (Thermo Scientific).
Results
Screening of genes that promote hematopoietic development from hESCs/iPSCs
Hematopoietic development from hESCs and hiPSCs recapitulates physiologic development, beginning in the conceptus and proceeding in a stepwise manner. CD34+CD43− endothelial cells (ECs) enriched in HE give rise to the earliest hematopoietic progenitors, pre-hematopoietic progenitor cells (pre-HPCs) with an immunophenotype of CD34+CD43+CD45−. Pre-HPCs then mature into CD34+CD43+CD45+ HPCs that express CD45, a marker antigen specific to hematopoietic cells (Figure 1A).20,21 We improved the conventional culture system to efficiently induce HPCs in EB culture by modifying cytokine conditions and adding an inhibitor of TGF-β signaling (Figure 1B). In our culture system, the expression of hematopoietic regulator genes such as RUNX1 and SCL/TAL1 increased in EBs after day 4 of culture accompanied by the decrease in expression of early mesodermal marker genes such as Brachyury (Figure 1C).
To identify genes that promote hematopoietic development from PSCs, we transduced hiPSC-derived ECs purified from day 6 EBs with several known hematopoietic regulator genes. We selected 13 genes that are known to play an important role in the development and/or maintenance of HSCs, including RUNX1, Scl/Tal1, Gata2, and HOXB4. The growth of the transduced cells was monitored in the presence of SCF and TPO for 14 days. Unexpectedly, most of these known regulator genes did not promote cell growth, but Sox17 did. A similar effect was observed when we transduced CD34+CD43+ pre-HPCs/HPCs from day 8 EBs (Figure 1D). To confirm these findings, we overexpressed Sox17 in hESC-derived ECs and pre-HPCs/HPCs. Overexpression of Sox17 also promoted cell growth of hESCs (data not shown). Based on these results, we decided to conduct a detailed analysis of the function of SOX17 using hESCs.
Sox17 promotes expansion of HE-like cells
SOX17 mRNA was highly expressed in EBs between days 2 and 4 of culture (Figure 1E). SOX17 has been described as one of the master regulator genes for endodermal development.10,22 High expression of SOX17 in EBs at early time points supposedly reflects the development of endodermal cells. In contrast, ECs emerged at approximately day 6 in our culture system at the same time as increased expression of hematopoietic regulator genes such as RUNX1 and SCL/TAL1 (Figure 1C). Therefore, the expression of SOX17 after day 6 may indicate a role of SOX17 in hematopoietic development (Figure 1E). Indeed, SOX17 was expressed at high levels in ECs, but at significantly lower levels in pre-HPCs, HPCs, and human CB CD34+ cells (Figure 1F). Other SOXF family genes, SOX7 and SOX18, showed a very similar pattern of expression profiles (Figure 1F).
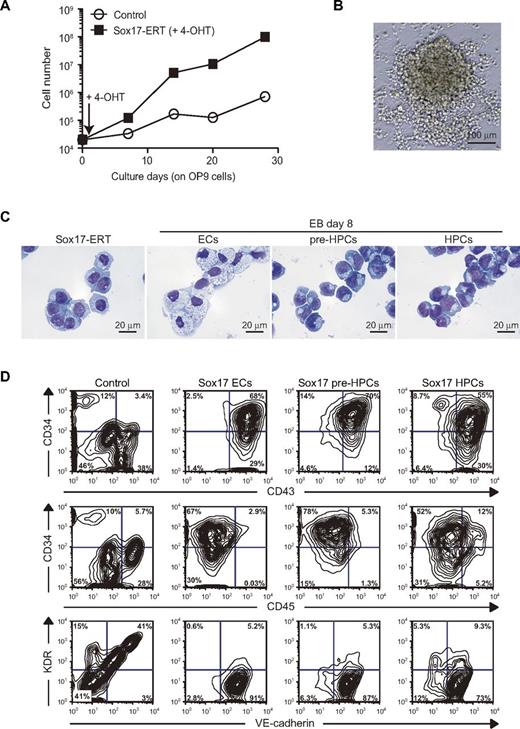
To evaluate the effect of overexpression of Sox17 in hematopoietic development in detail, we produced a retrovirus containing Sox17 fused to ERT (Sox17-ERT). We transduced ECs from day 6 EBs with the Sox17-ERT retrovirus on OP9 stromal cells and cultured them in the presence of SCF and TPO. The addition of 4-OHT, which induces nuclear translocation of ERT fusion protein, considerably stimulated cell growth (Figure 2A). Overexpression of Sox17-ERT promoted cell growth moderately even without 4-OHT, suggesting leaky translocation of Sox17-ERT (data not shown). Indeed, Sox17-ERT was detected in both the nucleus and cytoplasm without 4-OHT, whereas the addition of 4-OHT induced efficient nuclear translocation of Sox17-ERT (supplemental Figure 1A). Sox17-overexpressing cells formed semiadherent cell aggregates on OP9 cells (Figure 2B). Morphologic analysis revealed that they showed a monotonous morphology intermediate between ECs and pre-HPCs (Figure 2C). We performed further immunostaining with an anti-laminin Ab. After incubation in slide chambers for 4 hours, ECs attached to the slide glasses and stretched their cytoplasm out. In contrast, Sox17-overexpressing cells behaved like pre-HPCs and maintained a round shape, suggesting that Sox17-overexpressing cells do not retain strong adhesive properties of ECs, although they form semiadherent cell aggregates on OP9 cells (supplemental Figure 1B). Flow cytometric analysis demonstrated that Sox17-overexpressing cells expanded on OP9 cells were mostly CD34+CD43+ and did not express or expressed a low level of CD45 (CD45−/low). These cells coexpressed the HE producer VE-cadherin (Figure 2D). Interestingly, overexpression of Sox17 in pre-HPCs and HPCs from day 8 EBs similarly expanded CD34+CD43+CD45−/lowVE-cadherin+ cells (Figure 2D). Although the endothelial-specific marker KDR/FLK1 was expressed in the majority of ECs from day 6 and 8 EBs (data not shown), its expression was immediately down-regulated during differentiation into pre-HPCs and HPCs and also on activation of Sox17 (Figure 2D).
Sox17 promotes the expansion of CD34+CD43+CD45−/low cells. (A) Growth curve of ECs from day 6 EBs that were transduced with a Sox17-ERT or a control retrovirus. ECs (2 × 104) were transduced with the indicated retrovirus on OP9 cells and cultured in the presence of 20 ng/mL of SCF and TPO and 200nM 4-OHT. The absolute numbers of cells were determined and plotted. Representative data from repeated experiments are shown. (B) Appearance of a representative colony generated by Sox17-overexpressing cells in panel A observed under an inverted microscope. Images were collected using BIOREVO BZ-9000 (KEYENCE) with CFI Plan Fluor ELWD DM 20×C (Nikon). (C) Typical cell morphology of Sox17-overexpressing cells in panel A. Sorted cells were cytospun onto glass slides and observed after Wright-Giemsa staining. ECs, pre-HPCs, and HPCs from day 8 EBs served as controls. Images were collected using BIOREVO BZ-9000 (KEYENCE) with CFI Plan ApoVC 100×H (Nikon). (D) Flow cytometric analysis of expanded cells on overexpression of Sox17. ECs from day 6 EBs and pre-HPCs and HPCs from day 8 EBs were transduced with a Sox17-ERT or a control retrovirus cultured on OP9 in the presence of 20 ng/mL of SCF and TPO and 200nM 4-OHT for 10-15 days and then analyzed for their immunophenotypes.
Sox17 promotes the expansion of CD34+CD43+CD45−/low cells. (A) Growth curve of ECs from day 6 EBs that were transduced with a Sox17-ERT or a control retrovirus. ECs (2 × 104) were transduced with the indicated retrovirus on OP9 cells and cultured in the presence of 20 ng/mL of SCF and TPO and 200nM 4-OHT. The absolute numbers of cells were determined and plotted. Representative data from repeated experiments are shown. (B) Appearance of a representative colony generated by Sox17-overexpressing cells in panel A observed under an inverted microscope. Images were collected using BIOREVO BZ-9000 (KEYENCE) with CFI Plan Fluor ELWD DM 20×C (Nikon). (C) Typical cell morphology of Sox17-overexpressing cells in panel A. Sorted cells were cytospun onto glass slides and observed after Wright-Giemsa staining. ECs, pre-HPCs, and HPCs from day 8 EBs served as controls. Images were collected using BIOREVO BZ-9000 (KEYENCE) with CFI Plan ApoVC 100×H (Nikon). (D) Flow cytometric analysis of expanded cells on overexpression of Sox17. ECs from day 6 EBs and pre-HPCs and HPCs from day 8 EBs were transduced with a Sox17-ERT or a control retrovirus cultured on OP9 in the presence of 20 ng/mL of SCF and TPO and 200nM 4-OHT for 10-15 days and then analyzed for their immunophenotypes.
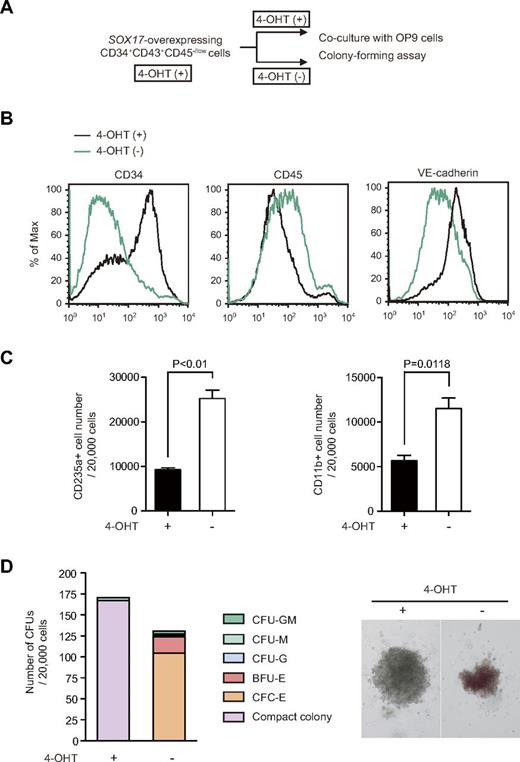
Comprehensive gene-expression analyses using microarrays were performed to investigate the developmental stage of the cells expanded on overexpression of Sox17. ECs from day 6 and 8 EBs, pre-HPCs from day 8 EBs, and HPCs from day 8 and 12 EBs were transduced with Sox17-ERT and cultured on OP9 cells. These cells were then treated with 4-OHT and the resulting CD34+CD43+CD45−/low cells were subjected to microarray analysis. Freshly isolated ECs from day 6, 8, and 12 EBs, pre-HSCs from day 8 EBs, and HPCs from day 8 and 12 EBs served as control samples. The CD34+CD43+CD45−/low cells overexpressing Sox17 appeared to express both EC-related genes such as VE-cadherin/CDH5 and ESAM and hematopoietic-related genes such as RUNX1 and SCL/TAL1 (supplemental Table 1). Hierarchical clustering of the cell populations based on the microarray data of total genes revealed that Sox17-overexpressing cells showed very similar profiles of gene expression irrespective of the cell sources (ie, ECs, pre-HPCs, and HPCs; Figure 3A). We next performed clustering using probes corresponding to genes identified as “Transcription factor” and “Hemopoiesis” from the GO database. Sox17-overexpressing cells were developmentally placed between ECs and pre-HPCs/HPCs (Figure 3B-C). These findings, together with the intermediate morphology between ECs and pre-HPCs, suggest that CD34+CD43+CD45−/low cells expanded on the overexpression of Sox17 are at a developmental stage between HE and early HPCs. To confirm this possibility, we then investigated whether the CD34+CD43+CD45−/low cells overexpressing Sox17 give rise to mature hematopoietic cells on inactivation of Sox17 (Figure 4A). As expected, after depletion of 4-OHT, CD34+CD43+CD45−/low cells lost expression of CD34 and VE-cadherin but gained a higher level of CD45 expression and gave rise to CD235a+ erythroblasts and CD11b+ myeloid cells more efficiently than they did in the presence of 4-OHT (Figure 4B-C). This trend was confirmed in colony-forming assays. We seeded CD34+CD43+CD45−/low cells overexpressing Sox17 in methylcellulose medium in the presence and absence of 4-OHT. Sox17-overexpressing cells in the presence of 4-OHT mainly formed compact colonies consisting of nonhemoglobinated cells with a morphology similar to ECs (Figure 2B), whereas they generated hemoglobinated erythroid colonies and myeloid colonies in the absence of 4-OHT (Figure 4D). These findings clearly indicate that the CD34+CD43+CD45−/low cells overexpressing Sox17 still retain hemogenic potential, which becomes apparent on removal of 4-OHT.
CD34+CD43+CD45−/low cells expanded on overexpression of Sox17 developmentally place between ECs and pre-HPCs/HPCs. Gene-expression patterns of wild-type and Sox17-overexpressing cells obtained in microarray analyses were clustered using hierarchical clustering. The distance between 2 samples was defined with the Pearson correlation using total genes (A) or certain probes selected from the GO database (B-C). “Transcription factor” represents genes that are located in the nucleus and have at least 1 of the GO terms “regulation of transcription, DNA-dependent,” “transcription factor activity,” or “transcription factor complex” (B). “Hemopoiesis” represents genes that are annotated with the GO terms “hemopoiesis,” “vasculogenesis,” “erythrocyte differentiation,” “erythrocyte maturation,” and/or “erythrocyte development” (C). The color of each cell represents the value of correlation indicated on the right side of the matrix.
CD34+CD43+CD45−/low cells expanded on overexpression of Sox17 developmentally place between ECs and pre-HPCs/HPCs. Gene-expression patterns of wild-type and Sox17-overexpressing cells obtained in microarray analyses were clustered using hierarchical clustering. The distance between 2 samples was defined with the Pearson correlation using total genes (A) or certain probes selected from the GO database (B-C). “Transcription factor” represents genes that are located in the nucleus and have at least 1 of the GO terms “regulation of transcription, DNA-dependent,” “transcription factor activity,” or “transcription factor complex” (B). “Hemopoiesis” represents genes that are annotated with the GO terms “hemopoiesis,” “vasculogenesis,” “erythrocyte differentiation,” “erythrocyte maturation,” and/or “erythrocyte development” (C). The color of each cell represents the value of correlation indicated on the right side of the matrix.
CD34+CD43+CD45−/low cells expanded on overexpression of Sox17 retain hemogenic potential. (A) Experimental design to evaluate effects of withdrawal of 4-OHT on Sox17-overexpressing cells. ECs from day 6 EBs transduced with a Sox17-ERT retrovirus were cultured in the presence of 20 ng/mL of SCF and TPO and 200nM 4-OHT for 15 days. Then, the cells were subjected to coculture with OP9 cells and colony-forming assays. For coculture with OP9 cells, the cells were replated onto OP9 cells and cultured in the presence of 20 ng/mL of SCF and TPO, 10 ng/mL of IL-3, and 3 units/mL of erythropoietin with and without 4-oht. at day 7 of culture, the cells were analyzed for their immunophenotypes by Flow cytometry. For colony-forming assays, the cells were replated in methylcellulose in the presence of 20 ng/mL of SCF, 10 ng/mL of TPO and IL-3, and 3 units/mL erythropoietin with and without 4-OHT. At day 12 of culture, the colonies were counted. (B) Representative flow cytometric profiles of cells overexpressing Sox17-ERT before and after depletion of 4-OHT. (C) The absolute numbers of CD235+ erythroblasts and CD11b+ myeloid cells in culture at 7 days after depletion of 4-OHT. Data are shown as the means ± SD for triplicate cultures. (D) Ability of Sox17-ERT–overexpressing cells to form hematopoietic colonies in methylcellulose cultures with or without 4-OHT. The numbers of CFUs in culture are presented (left panel). CFU-GM, CFU-M, CFU-G, BFU-E, and CFU-E indicate CFU-granulocyte-macrophage, CFU-macrophage, CFU-granulocyte, burst-forming unit-erythroid, and colony-forming unit-erythroid, respectively. Compact colonies indicate colonies composed by HE cell–like cells. The appearance of a representative compact colony and an erythroid colony observed under an inverted microscope is depicted (right panel).
CD34+CD43+CD45−/low cells expanded on overexpression of Sox17 retain hemogenic potential. (A) Experimental design to evaluate effects of withdrawal of 4-OHT on Sox17-overexpressing cells. ECs from day 6 EBs transduced with a Sox17-ERT retrovirus were cultured in the presence of 20 ng/mL of SCF and TPO and 200nM 4-OHT for 15 days. Then, the cells were subjected to coculture with OP9 cells and colony-forming assays. For coculture with OP9 cells, the cells were replated onto OP9 cells and cultured in the presence of 20 ng/mL of SCF and TPO, 10 ng/mL of IL-3, and 3 units/mL of erythropoietin with and without 4-oht. at day 7 of culture, the cells were analyzed for their immunophenotypes by Flow cytometry. For colony-forming assays, the cells were replated in methylcellulose in the presence of 20 ng/mL of SCF, 10 ng/mL of TPO and IL-3, and 3 units/mL erythropoietin with and without 4-OHT. At day 12 of culture, the colonies were counted. (B) Representative flow cytometric profiles of cells overexpressing Sox17-ERT before and after depletion of 4-OHT. (C) The absolute numbers of CD235+ erythroblasts and CD11b+ myeloid cells in culture at 7 days after depletion of 4-OHT. Data are shown as the means ± SD for triplicate cultures. (D) Ability of Sox17-ERT–overexpressing cells to form hematopoietic colonies in methylcellulose cultures with or without 4-OHT. The numbers of CFUs in culture are presented (left panel). CFU-GM, CFU-M, CFU-G, BFU-E, and CFU-E indicate CFU-granulocyte-macrophage, CFU-macrophage, CFU-granulocyte, burst-forming unit-erythroid, and colony-forming unit-erythroid, respectively. Compact colonies indicate colonies composed by HE cell–like cells. The appearance of a representative compact colony and an erythroid colony observed under an inverted microscope is depicted (right panel).
We next compared the expression of globin genes in Sox17-overexpressing CD34+CD43+CD45−/low HE-like cells and their hematopoietic progeny with globin gene expression in CB CD34+ cells. CD34+CD43+CD45−/low cells expanded on overexpression of Sox17 were further cultured in the presence and absence of 4-OHT for 7 days and then GFP+ cells expressing Sox17-ERT were collected by cell sorting. RT-PCR analysis revealed that embryonic globin (ϵ) and fetal globin (γ), but not adult globin (β), were highly expressed in Sox17-overexpressing cells and/or their hematopoietic progeny (supplemental Figure 2). These results raise the possibility that the CD34+CD43+CD45−/low HE-like cells expanded on overexpression of Sox17 are a hemogenic intermediate differentiated from hemangioblasts that primarily give rise to yolk sac–type blood cells.8
SOX17 is essential for the hemogenic activity of HE cells
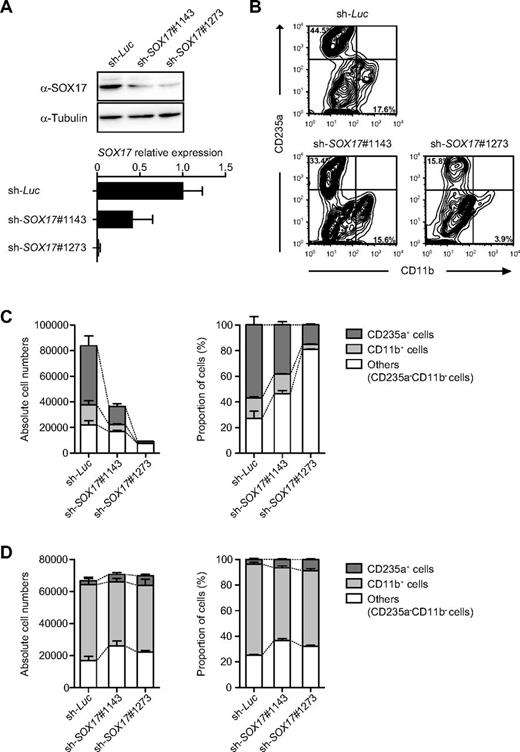
Our results so far indicate that the overexpression of Sox17 promotes the expansion of HE-like cells, but inhibits their hematopoietic differentiation into pre-HPCs. Because SOX17 is highly expressed in ECs enriched in HE, we examined the role of SOX17 by knock-down analysis. We transduced ECs from day 5 EBs with lentiviruses expressing shRNA against SOX17 on OP9 cells and allowed them to differentiate into hematopoietic cells for 9 days. The most effective shRNA, sh-SOX17#1273 (Figure 5A), suppressed the development and differentiation of hematopoietic cells including both erythroblasts and myeloid cells significantly, whereas it only moderately diminished the growth of CD235a−CD11b− nonhematopoietic cells, the majority of which do not express SOX17 even though approximately 25%-30% of these cells are SOX17+ ECs (Figure 5B-C). sh-SOX17#1143 similarly, albeit modestly, suppressed the production of hematopoietic cells. Similar results were obtained when we knocked down SOX17 in ECs from day 6 EBs (data not shown). However, hematopoietic differentiation was not affected on SOX17 knock-down in pre-HPCs from day 8 EBs (Figure 5D). These findings indicate that SOX17 plays a key role in the acquisition of hematopoietic potential in HE cells.
Hematopoietic differentiation from HE is inhibited by depletion of SOX17. (A) Knock-down efficiencies of shRNAs against SOX17. Western blot analysis of SOX17 in 293T cells transduced with shRNAs against SOX17 (top panel). α-Tubulin was used as the loading control. pre-HPCs from day 8 EBs were transduced with shRNAs against SOX17 on OP9 cells and cultured in the presence of 20 ng/mL of SCF and TPO, 10 ng/mL of IL-3, and 3 units/mL of erythropoietin (EPO) for 7 days. Levels of endogenous SOX17 were analyzed by quantitative RT-PCR analysis (bottom panel). mRNA levels were normalized to GAPDH expression. Expression levels relative to that in the control cells transduced with an shRNA against Luciferase are shown as the means ± SD for triplicate analyses. (B) Effects of depletion of SOX17 on hematopoietic development from HE cells. ECs from day 5 EBs were transduced with shRNAs against SOX17 on OP9 cells and were cultured in the presence of 20 ng/mL of SCF and TPO, 10 ng/mL of IL-3, and 3 units/mL of EPO for 9 days. Representative flow cytometric profiles of cells at day 9 of culture are depicted. (C) Absolute numbers and proportion of CD235a+ erythroblasts and CD11b+ myeloid cells in panel B at day 9 of culture. Data are shown as the means ± SD for 3 independent cultures. (D) Effects of depletion of SOX17 on pre-HPCs. Pre-HPCs from day 8 EBs were transduced with shRNAs against SOX17 on OP9 cells and were cultured in the presence of 20 ng/mL of SCF and TPO, 10 ng/mL of IL-3, and 3 units/mL of EPO for 7 days. Absolute numbers and proportion of CD235a+ erythroblasts and CD11b+ myeloid cells at day 7 of culture are presented. Data are shown as the means ± SD for triplicate cultures of 1 of 2 independent experiments that gave similar results.
Hematopoietic differentiation from HE is inhibited by depletion of SOX17. (A) Knock-down efficiencies of shRNAs against SOX17. Western blot analysis of SOX17 in 293T cells transduced with shRNAs against SOX17 (top panel). α-Tubulin was used as the loading control. pre-HPCs from day 8 EBs were transduced with shRNAs against SOX17 on OP9 cells and cultured in the presence of 20 ng/mL of SCF and TPO, 10 ng/mL of IL-3, and 3 units/mL of erythropoietin (EPO) for 7 days. Levels of endogenous SOX17 were analyzed by quantitative RT-PCR analysis (bottom panel). mRNA levels were normalized to GAPDH expression. Expression levels relative to that in the control cells transduced with an shRNA against Luciferase are shown as the means ± SD for triplicate analyses. (B) Effects of depletion of SOX17 on hematopoietic development from HE cells. ECs from day 5 EBs were transduced with shRNAs against SOX17 on OP9 cells and were cultured in the presence of 20 ng/mL of SCF and TPO, 10 ng/mL of IL-3, and 3 units/mL of EPO for 9 days. Representative flow cytometric profiles of cells at day 9 of culture are depicted. (C) Absolute numbers and proportion of CD235a+ erythroblasts and CD11b+ myeloid cells in panel B at day 9 of culture. Data are shown as the means ± SD for 3 independent cultures. (D) Effects of depletion of SOX17 on pre-HPCs. Pre-HPCs from day 8 EBs were transduced with shRNAs against SOX17 on OP9 cells and were cultured in the presence of 20 ng/mL of SCF and TPO, 10 ng/mL of IL-3, and 3 units/mL of EPO for 7 days. Absolute numbers and proportion of CD235a+ erythroblasts and CD11b+ myeloid cells at day 7 of culture are presented. Data are shown as the means ± SD for triplicate cultures of 1 of 2 independent experiments that gave similar results.
SOX17 regulates directly the transcription of key regulator genes for HE cells
A ChIP-on-chip analysis was conducted to identify the direct target genes of SOX17 in HE cells. We transduced ECs cells from day 6 EBs with a 3xFlag-Sox17-ERT retrovirus and expanded CD34+CD43+CD45−/low HE-like cells on OP9 cells. At day 27 of culture, 94.2% of the expanded cells were positive for CD34. CD34+ cells were further enriched (99.8%) by magnetic cell sorting using magnetic beads conjugated with anti-CD34 Abs, and these purified cells were then subjected to ChIP-on-chip analysis.
The ChIP-on-chip analysis was performed with human promoter microarrays containing approximately 21 000 probe sets covering from −8.0 kb upstream to +4.0 kb downstream of the TSS of RefSeq genes. 3xFlag-Sox17 was cross-linked to DNA and precipitated using the anti-FLAG M2 Ab. Gene promoters bound by Sox17 were ranked according to fold enrichments calculated in comparison with signals obtained with the input DNA. Of the 19 457 gene promoter regions analyzed, 182 and 98 regions showed Sox17 binding with an enrichment greater than 2- and 3-fold, respectively (full data are listed in supplemental Table 2). The functional annotation of the genes bound by Sox17 with a fold enrichment greater than 3 was performed based on GO and showed significant enrichment for genes that fell into categories such as “vasculogenesis,” “hemopoiesis,” and “positive regulation of erythrocyte differentiation” (Figure 6A, Table 1, and supplemental Table 2). The genes bound by Sox17 include genes well characterized as regulators of hematopoietic development from HE cells, such as VE-cadherin/CDH5, RUNX1, SCL/TAL1, and HHEX (Table 1 and supplemental Table 2). VE-cadherin, an endothelial marker antigen, is expressed by HE cells and by early HSCs, which appear in the yolk sac and the aorta-gonad-mesonephros region, as well as by a transient HSC population of the fetal liver.23-25 RUNX1, SCL/TAL1, and HHEX encode transcription factors essential for the development of HSCs from HE cells or hemangioblasts.5,26,27 The distribution of SOX17 signals at these genes varied greatly across the promoter region (Figure 6B). SOX17 is reported to bind to the consensus motif of “ATTGT.”22 The VE-cadherin/CDH5 promoter contains the consensus motif between −107 bp and −103 bp from the TSS. It was recently reported that this site is conserved in mouse and Sox7 binds directly to it to activate transcription.28 Our ChIP-on-chip data showed that Sox17 binds to this site as well (Figure 6B arrowhead). These findings clearly indicate that SOX17 regulates directly the expression of a set of key genes for hematopoietic development in HE cells.
Targets of SOX17 detected by ChIP-on-chip analysis. (A) GO analysis of the Sox17 targets detected by ChIP-on-chip analysis. CD34+CD43− cells from day 6 EBs were transduced with a 3× Flag SOX17-ERT retrovirus. The cells were further cultured on OP9 cells in the presence of 20 ng/mL of SCF and TPO and 200nM 4-OHT. CD34+ cells were collected at day 27 of culture and subjected to ChIP-on-chip analysis. P for each GO term is indicated. (B) ChIP-on-chip profile of SOX17 occupancy at genes related to hematopoietic development from HE cells. Plot under the x-axis shows the position of probe sets. Arrowhead at the VE-cadherin/CDH5 promoter indicate consensus motif of Sox17-binding site. (C) Gene-expression patterns of wild-type and engineered cells obtained in microarray analyses clustered using hierarchical clustering. Distance between 2 samples was defined with the Pearson correlation of Sox17 target genes with Sox17 binding more than 3-fold in the ChIP-on-chip analysis presented in supplemental Table 2. The color of each cell represents the value of correlation indicated on the right side of the matrix. (D) Comparative analysis of ChIP-on-chip and microarray data. Venn diagrams showing the number of genes bound by Sox17 (> 3-fold enrichment) and the number of genes up-regulated in expression more than 2-fold in at least 1 cell type among ECs, pre-HPCs, and HPCs on overexpression of Sox17 (Sox17-overexpressing CD34+CD43+CD45−/low cells compared with those of respective fresh controls; left panel). The percentages of overlapping and nonoverlapping bound genes are indicated in parentheses. Shown are the correlation of Sox17 binding (fold enrichment) in ChIP-on-chip analysis and the fold changes in expression during differentiation of ECs to HPCs (right panel). The fold enrichments and fold changes in expression were plotted for 84 genes of 98 showing enrichments greater than 3-fold (the microarray data were not available for the remaining 14 genes). Correlation coefficients (R) are indicated for genes with fold enrichment greater than 3- and 6-fold, respectively.
Targets of SOX17 detected by ChIP-on-chip analysis. (A) GO analysis of the Sox17 targets detected by ChIP-on-chip analysis. CD34+CD43− cells from day 6 EBs were transduced with a 3× Flag SOX17-ERT retrovirus. The cells were further cultured on OP9 cells in the presence of 20 ng/mL of SCF and TPO and 200nM 4-OHT. CD34+ cells were collected at day 27 of culture and subjected to ChIP-on-chip analysis. P for each GO term is indicated. (B) ChIP-on-chip profile of SOX17 occupancy at genes related to hematopoietic development from HE cells. Plot under the x-axis shows the position of probe sets. Arrowhead at the VE-cadherin/CDH5 promoter indicate consensus motif of Sox17-binding site. (C) Gene-expression patterns of wild-type and engineered cells obtained in microarray analyses clustered using hierarchical clustering. Distance between 2 samples was defined with the Pearson correlation of Sox17 target genes with Sox17 binding more than 3-fold in the ChIP-on-chip analysis presented in supplemental Table 2. The color of each cell represents the value of correlation indicated on the right side of the matrix. (D) Comparative analysis of ChIP-on-chip and microarray data. Venn diagrams showing the number of genes bound by Sox17 (> 3-fold enrichment) and the number of genes up-regulated in expression more than 2-fold in at least 1 cell type among ECs, pre-HPCs, and HPCs on overexpression of Sox17 (Sox17-overexpressing CD34+CD43+CD45−/low cells compared with those of respective fresh controls; left panel). The percentages of overlapping and nonoverlapping bound genes are indicated in parentheses. Shown are the correlation of Sox17 binding (fold enrichment) in ChIP-on-chip analysis and the fold changes in expression during differentiation of ECs to HPCs (right panel). The fold enrichments and fold changes in expression were plotted for 84 genes of 98 showing enrichments greater than 3-fold (the microarray data were not available for the remaining 14 genes). Correlation coefficients (R) are indicated for genes with fold enrichment greater than 3- and 6-fold, respectively.
Candidate SOX17 target genes according to ChIP-chip scores
| Rank . | Symbol . | Gene name . | Fold enrichment . | GO term . | Fold difference . | ||||
|---|---|---|---|---|---|---|---|---|---|
| Vasculogenesis . | Positive regulation of erythrocyte differentiation . | Hemopoiesis . | Sox17 ECs d6/ECs d6 . | Sox17 pre-HPCs d8/pre-HPCs d8 . | Sox17 HPCs d8/HPCs d8 . | ||||
| 1 | GPSM3 | G-protein signaling modulator 3 | 48.50 | No | No | No | 1.38 | 2.42 | 2.56 |
| 2 | MAST4 | Microtubule associated serine/threonine kinase family member 4 | 41.62 | No | No | No | 1.02 | 3.22 | 4.75 |
| 3 | TXLNB | Taxilin beta, muscle-derived protein 77 | 38.05 | No | No | No | 1.18 | 2.89 | 4.29 |
| 4 | EGOT | Eosinophil granule ontogeny transcript | 37.53 | No | No | No | ND | ND | ND |
| 5 | PPBPL1 | Pro-platelet basic protein-like 1 | 35.26 | No | No | No | ND | ND | ND |
| 6 | MFSD6 | Major facilitator superfamily domain containing 6 | 30.27 | No | No | No | 19.79 | 14.24 | 19.26 |
| 7 | CDH5 | Cadherin 5, type 2 | 29.86 | No | No | No | 0.70 | 9.07 | 10.81 |
| 8 | BCL6B | B-cell CLL/lymphoma 6, member B | 28.05 | No | No | No | 1.11 | 5.73 | 8.50 |
| 9 | C1orf55 | Chromosome 1 open reading frame 55 | 25.81 | No | No | No | 2.23 | 1.77 | 1.44 |
| 10 | PTTG1IP | Pituitary tumor-transforming 1 interacting protein | 25.28 | No | No | No | 0.93 | 1.49 | 1.67 |
| 11 | TRIM67 | Tripartite motif containing 67 | 24.08 | No | No | No | 4.16 | 4.29 | 5.03 |
| 12 | MYCT1 | myc target 1myc target 1 | 21.26 | No | No | No | 0.74 | 2.90 | 7.66 |
| 13 | CD40LG | CD40 ligand | 14.12 | No | No | No | 6.65 | 11.57 | 16.69 |
| 14 | SCOC | Short coiled-coil protein | 13.45 | No | No | No | 0.97 | 0.79 | 1.04 |
| 15 | PPP1R16B | Protein phosphatase 1, regulatory subunit 16B | 13.18 | No | No | No | 0.70 | 1.83 | 1.15 |
| 19 | ACVR2A | Activin A receptor, type IIA | 9.45 | No | Yes | No | 0.15 | 0.42 | 0.67 |
| 26 | HHEX | Hematopoietically expressed homeobox | 6.87 | Yes | No | Yes | 0.66 | 1.44 | 0.89 |
| 37 | RUNX1 | runt-related transcription factor 1 | 5.46 | No | No | Yes | 5.02 | 1.25 | 1.12 |
| 41 | TAL1/SCL | T-cell acute lymphocytic leukemia 1 | 5.28 | No | Yes | Yes | 1.82 | 1.76 | 1.92 |
| 50 | EGFL7 | EGF-like-domain, multiple 7 | 4.69 | Yes | No | No | 0.67 | 2.11 | 1.92 |
| 69 | JUNB | jun B protooncogene | 3.78 | Yes | No | No | 3.50 | 3.87 | 3.24 |
| Rank . | Symbol . | Gene name . | Fold enrichment . | GO term . | Fold difference . | ||||
|---|---|---|---|---|---|---|---|---|---|
| Vasculogenesis . | Positive regulation of erythrocyte differentiation . | Hemopoiesis . | Sox17 ECs d6/ECs d6 . | Sox17 pre-HPCs d8/pre-HPCs d8 . | Sox17 HPCs d8/HPCs d8 . | ||||
| 1 | GPSM3 | G-protein signaling modulator 3 | 48.50 | No | No | No | 1.38 | 2.42 | 2.56 |
| 2 | MAST4 | Microtubule associated serine/threonine kinase family member 4 | 41.62 | No | No | No | 1.02 | 3.22 | 4.75 |
| 3 | TXLNB | Taxilin beta, muscle-derived protein 77 | 38.05 | No | No | No | 1.18 | 2.89 | 4.29 |
| 4 | EGOT | Eosinophil granule ontogeny transcript | 37.53 | No | No | No | ND | ND | ND |
| 5 | PPBPL1 | Pro-platelet basic protein-like 1 | 35.26 | No | No | No | ND | ND | ND |
| 6 | MFSD6 | Major facilitator superfamily domain containing 6 | 30.27 | No | No | No | 19.79 | 14.24 | 19.26 |
| 7 | CDH5 | Cadherin 5, type 2 | 29.86 | No | No | No | 0.70 | 9.07 | 10.81 |
| 8 | BCL6B | B-cell CLL/lymphoma 6, member B | 28.05 | No | No | No | 1.11 | 5.73 | 8.50 |
| 9 | C1orf55 | Chromosome 1 open reading frame 55 | 25.81 | No | No | No | 2.23 | 1.77 | 1.44 |
| 10 | PTTG1IP | Pituitary tumor-transforming 1 interacting protein | 25.28 | No | No | No | 0.93 | 1.49 | 1.67 |
| 11 | TRIM67 | Tripartite motif containing 67 | 24.08 | No | No | No | 4.16 | 4.29 | 5.03 |
| 12 | MYCT1 | myc target 1myc target 1 | 21.26 | No | No | No | 0.74 | 2.90 | 7.66 |
| 13 | CD40LG | CD40 ligand | 14.12 | No | No | No | 6.65 | 11.57 | 16.69 |
| 14 | SCOC | Short coiled-coil protein | 13.45 | No | No | No | 0.97 | 0.79 | 1.04 |
| 15 | PPP1R16B | Protein phosphatase 1, regulatory subunit 16B | 13.18 | No | No | No | 0.70 | 1.83 | 1.15 |
| 19 | ACVR2A | Activin A receptor, type IIA | 9.45 | No | Yes | No | 0.15 | 0.42 | 0.67 |
| 26 | HHEX | Hematopoietically expressed homeobox | 6.87 | Yes | No | Yes | 0.66 | 1.44 | 0.89 |
| 37 | RUNX1 | runt-related transcription factor 1 | 5.46 | No | No | Yes | 5.02 | 1.25 | 1.12 |
| 41 | TAL1/SCL | T-cell acute lymphocytic leukemia 1 | 5.28 | No | Yes | Yes | 1.82 | 1.76 | 1.92 |
| 50 | EGFL7 | EGF-like-domain, multiple 7 | 4.69 | Yes | No | No | 0.67 | 2.11 | 1.92 |
| 69 | JUNB | jun B protooncogene | 3.78 | Yes | No | No | 3.50 | 3.87 | 3.24 |
ND indicates no data.
We next performed hierarchical clustering of the cell populations based on the microarray data using “SOX17 targets” that we arbitrarily selected as genes with Sox17 binding greater than 3-fold over the input levels in the ChIP-on-chip analysis (supplemental Table 2). As expected, Sox17-overexpressing cells were again developmentally placed between ECs and pre-HPCs/HPCs (Figure 6C).
Comparison of gene lists between the ChIP-on-chip and microarray assays
We next examined the changes in expression of the 98 genes bound by Sox17 (> 3-fold enrichment in the ChIP-on-chip analysis) on Sox17 overexpression. The microarray data of Sox17-overexpressing CD34+CD43+CD45−/low cells shown in Figure 3 were compared with those of respective fresh controls. Sox17 is thought to activate the transcription of target genes.22 As expected, 36 of the 98 genes showed up-regulation in expression of more than 2-fold in at least in 1 cell type (Figure 6D, Table 1, and supplemental Table 2). This tendency was evident in the top 15 genes with Sox17 binding (Table 1 and supplemental Table 2), although in 4 genes of 15, the effects of overexpression of Sox17 were obvious only in pre-HPCs and HPCs, but not in ECs, which possess a high level of endogenous SOX17 (supplemental Table 2). Similarly, a negative correlation was detected between the levels of Sox17 binding in ChIP-on-chip analysis and the fold changes in expression during differentiation of ECs (SOX17+) to HPCs (SOX17−; Figure 6D). Among genes bound by Sox17, EGFL7 and VE-cadherin/CDH5 have been shown to be up-regulated in BM HPCs transduced with Sox17.13 However, in our ChIP-on-chip analysis, Sox17 did not show any binding to the genes directly regulated by Sox17 during differentiation of ES cells into extraembryonic endoderm.22 These data suggest that SOX17 regulates different targets in hematopoietic and endodermal development.
Discussion
In the present study, we found that all of the SOXF subfamily genes, SOX7, SOX17, and SOX18, are highly expressed in hESC-derived ECs enriched in HE and markedly down-regulated in pre-HPCs and HPCs to the levels comparable to that in CB CD34+ cells. Overexpression of Sox17 in ECs resulted in expansion of monotonous cells with a CD34+CD43+CD45−/low immunophenotype. These cells coexpressed hematopoietic marker antigens such as CD43 and a low level of CD45, as well as the HE marker VE-cadherin. These unique characteristics of Sox17-overexpressing ECs are reminiscent of HE cells. Overexpression of Sox17 inhibited the hematopoietic differentiation of both pre-HPCs and HPCs and reprogrammed them into HE-like cells. In contrast, depletion of SOX17 in pre-HPCs did not affect their hematopoietic differentiation. These findings suggest that SOX17 is one of the master regulators that define HE but must be down-regulated during the development of pre-HPCs to allow hematopoietic differentiation.
The effects of overexpression of SOX17 in hESC-derived ECs and HPCs are very similar to that of overexpression of Sox7 and Sox18 in early hematopoietic precursors from mouse embryos and mouse ESCs.14,15,28 However, it has been reported in mice that Sox17 remains marginally expressed during blood specification and the overexpression of Sox17 in early hematopoietic precursors induces massive apoptosis.14 The contrasting effects of Sox17 between humans and the mouse is somewhat surprising but could be partially attributed to the difference in expression during early hematopoiesis (see previous paragraph). The expression of all of the SOXF subfamily genes in ECs evokes the possibility that they have redundant function in the development of hematopoiesis from hESCs, as they do in postnatal angiogenesis in mice.11 Nonetheless, the effects of knock-down of Sox7 in mice and SOX17 in the present study are different. Sox7 knock-down in Brachyury+Flk1− mesodermal precursors, which give rise on further differentiation to Flk1+ cells containing hemangioblast precursors, profoundly inhibited the production of both hematopoietic progenitors and endothelial progenitors, leaving open the possibility that Sox7 inhibits the production of hematopoietic progenitors through inhibiting the formation of HE or hemangioblasts. In contrast, SOX17 knock-down in ECs enriched in HE in the present study mainly compromised the development of mature hematopoietic cells and only mildly affected the proliferation of nonhematopoietic cells. Furthermore, depletion of SOX17 in pre-HPCs did not significantly affect their hematopoietic differentiation. Therefore, the role of SOX17 at the developmental stage of blood specification could be more specific to the establishment of a hemogenic program in mesodermal or endothelial precursors compared with that of Sox7. Although we did not detect the effects of SOX17 knock-down in pre-HPCs, recent studies have shown that Sox17 also plays an important role in the maintenance of fetal and neonatal HSCs, but not adult HSCs.12 Sox17 has also been demonstrated to confer fetal HSC characteristics to adult hematopoietic progenitors.13 SOX17 may again exert its critical function at a stage later than the pre-HPC stage, when pre-HPCs/HPCs differentiate into embryonic HSCs.
Very similar results have been demonstrated in the murine system with the transcription factor HoxA3. HoxA3 is a gene uniquely expressed in the embryonic vasculature, but not in the yolk sac vasculature. HoxA3 restrains hematopoietic differentiation of the earliest endothelial progenitors and can induce reversion of the earliest hematopoietic progenitors into CD41-negative endothelial cells.29 This reversible modulation of endothelial-hematopoietic state is accomplished by down-regulation of key hematopoietic transcription factors. Among these, Runx1 is able to erase the endothelial program set up by HoxA3 and promote hematopoietic differentiation. Sox17 was listed as one of the targets regulated by HoxA3. Given that SOX17 appeared to regulate directly the expression of RUNX1 in this study, it could be assumed that HoxA3 functions as an apical regulator of HE, eventually activating the transcription of Runx1 via up-regulation of Sox17 to initiate hematopoietic differentiation. It would be intriguing to address this question.
The direct targets for Sox17 have been characterized during endodermal differentiation of mouse ESCs using ChIP-on-chip analysis. The Sox17-binding consensus motif has also been identified using de novo motif analysis from the ChIP-on-chip data.22 As expected, the genes bound by Sox17 in Sox17-overexpressing HE-like cells were quite different from those detected during endodermal differentiation and were related to the GO terms “vasculogenesis,” “hemopoiesis,” or “positive regulation of erythrocyte differentiation.” Among these genes, VE-cadherin/CDH5 encodes one of the well-known marker antigens of HE and is also expressed by embryonic HSCs.24,25 Sox17 appears to bind directly to the promoters of RUNX1, SCL/TAL1, and HHEX, which encode key transcription factors essential to the development of HSCs from HE cells or hemangioblasts.4,26,27 Other target genes included BAZF/BCL6B, JUNB, and EGFL7, which encode a POZ/BTB zinc finger protein, a basic HLH transcription factor, and a secreted angiogenic factor, respectively. These genes have been implicated in vasculogenesis and/or angiogenesis and BAZF/BCL6B and JUNB have also been implicated in hematopoiesis.30-34 The profiles of these Sox17 targets during early hematopoietic development further support the critical role of SOX17 in the regulation of HE.
The results of the present study have unveiled a novel function of SOX17 in hematopoietic development. Because the overexpression of Sox17 expands HE-like cells, it is possible that conditional expression of SOX17 in hESC-derived endothelial progenitors facilitates hematopoietic development. Therefore, SOX17 could be a novel target for manipulation to improve the yield of hematopoietic progenies from hESCs for regenerative cell therapies.
There is an Inside Blood commentary on this article in this issue.
The online version of this article contains a data supplement.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.
Acknowledgments
The authors thank Toru Nakano for providing OP9 cells; Makiko Yui and Atsunori Saraya for technical assistance; George Wendt for critical reading of the manuscript; and Mieko Tanemura and Akemi Matsumura for laboratory assistance.
This work was supported in part by Grants-in-Aid for Scientific Research (21390289 and 23659483) and the Global Center for Education and Research in Immune System Regulation and Treatment, MEXT, Japan; a Grant-in-Aid for Core Research for Evolutional Science and Technology (CREST) from the Japan Science and Technology Corporation (JST); a grant from the Astellas Foundation for Research on Metabolic Disorders; and a grant from the Tokyo Biochemical Research Foundation.
Authorship
Contribution: Y. N.-T. performed the experiments, analyzed the results, produced the figures, and wrote the manuscript; M. Osawa, M. Oshima, H.T., and S.M. assisted with the experiments including the hematopoietic analyses; M.E., T.A.E., T.T, and H.K. performed the microarray and ChIP-on-chip analyses; N.T., K.E., and H.N. generated the iPSCs; M. Osawa and A.I. conceived of and directed the project; and A.I. secured the funding and wrote the manuscript.
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Correspondence: Mitsujiro Osawa, PhD, or Atsushi Iwama, MD, PhD, 1-8-1 Inohana, Chuo-ku, Chiba 260-8670, Japan; e-mail: mitsujiro.osawa@faculty.chiba-u.jp or aiwama@faculty.chiba-u.jp.




This feature is available to Subscribers Only
Sign In or Create an Account Close Modal