Abstract
The receptor tyrosine kinase Flt3 is expressed and functionally important in early myeloid progenitor cells and in the majority of acute myeloid leukemia (AML) blasts. Internal tandem duplications (ITDs) in the juxtamembrane domain of the receptor occur in 25% of AML cases. Previously, we have shown that these mutations activate the receptor and induce leukemic transformation. In this study, we performed genome-wide parallel expression analyses of 32Dcl3 cells stably transfected with either wild-type or 3 different ITD isoforms of Flt3. Comparison of microarray expression analyses revealed that 767 of 6586 genes differed in expression between FLT3-WT– and FLT3-ITD–expressing cell lines. The target genes of mutationally activated Flt3 resembled more closely those of the interleukin 3 (IL-3) receptor than those of ligand-activated Flt3. The serine-threonine kinase Pim-2 was up-regulated on the mRNA and the protein level in Flt3-ITD–expressing cells. Further experiments indicated that Pim-2 function was important for clonal growth of 32D cells. Several genes repressed by the mutations were found to be involved in myeloid gene regulation. Pu.1 and C/EBPα, both induced by ligand-activation of wild-type Flt3, were suppressed in their expression and function by the Flt3 mutations. In conclusion, internal tandem duplication mutations of Flt3 activate transcriptional programs that partially mimic IL-3 activity. Interestingly, other parts of the transcriptional program involve novel, IL-3–independent pathways that antagonize differentiation-inducing effects of wild-type Flt3. The identification of the transcriptional program induced by ITD mutations should ease the development of specific therapies.
Introduction
The receptor tyrosine kinase (RTK) Flt3 is expressed in early hematopoietic progenitor cells and mediates important functions for proliferation and survival.1 The receptor shows considerable functional overlap with c-kit, the closely related stem cell factor receptor. In synergism with lineage-specific cytokines, Flt3 activation enhances colony-forming capacity of all hematopoietic lineages.
In acute myeloid leukemia (AML), the receptor is expressed on leukemic blasts in the majority of cases. A series of mutations of the receptor has been reported that results in the insertion of several amino acids in its juxtamembrane domain.2,3 These mutations activate Flt3 kinase activity. We and other researchers have shown that internal tandem duplication (ITD) mutations cause malignant transformation of myeloid cells in vitro and in vivo.4-7 The mutations are present in about 20% to 25% of AML cases, and in several patient series they were associated with decreased survival.8-13In addition, we previously described an activating point mutation in the kinase domain of the receptor14 that has recently been shown to occur in another 7% of patients with AML.15 16Thus, activating mutations of Flt3 are the most frequent genetic alterations in AML and present a promising therapeutic target for treatment of this disease.
Previously, we analyzed signaling properties and biologic functions of the ITD mutations in myeloid progenitor cell lines.6 In contrast to ligand-activated wild-type receptors, mutation-activated Flt3 induced colony growth of the mouse myeloid progenitor cell line 32D as well as constitutive and aberrant activation of the signal transducer and activator of transcription 3 (STAT3) and STAT5 transcription factors. However, the mutations only marginally activated signaling cascades that are induced by ligand-activated wild-type receptor.
Little is known about the effects of Flt3 mutations on the transcriptional program of myeloid cells. Also, no data are available on the influence of these mutations on the hematopoietic differentiation program. Its disturbance is one of the hallmarks of AML, associated with the inhibition of several myeloid transcription factors by the products of recurrent chromosomal translocations, often in a dominant-negative fashion.17 Currently, no data on the interference of Flt3 mutations with the expression and function of these transcription factors are known.
Here, we identified target genes of the Flt3 mutations by microarray expression profiling. ITD mutations induced transcriptional programs that partially mimicked IL-3 activity—many genes being specifically regulated by the mutations but not by ligand-activated wild-type Flt3. We found profound induction of Pim-2, a member of a family of myb-activating kinases to be specifically associated with the mutations. Cotransfection experiments with kinase-defective forms of Pim-2 revealed the functional relevance of this kinase for ITD-mediated transformation.
Interestingly, other parts of the mutation-induced transcriptional program involved novel pathways, antagonizing the function elicited by ligand-activated Flt3. In the presence of the ITD mutations, the expression and function of several myeloid transcription factors was significantly repressed, in contrast to the induction caused by activation of wild-type Flt3. These results indicated that the ITD mutations not only constitutively activated the Flt3 kinase activity, but they also induced aberrant receptor functions with influence not only on proliferation and survival but also on myeloid differentiation programs.
Patients, materials, and methods
Reagents and cell lines
Recombinant human Flt3 ligand (FL) and recombinant murine interleukin 3 (IL-3) were purchased from Pepro Tech (Rocky Hill, NJ). Phycoerythrin (PE)–labeled monoclonal rat antimouse and mouse antihuman Flt3 antibodies as well as appropriate isotype controls were obtained from Pharmingen (San Diego, CA). Polyclonal rabbit antibodies for Pim-2, Pu.1, C/EBPα, and Actin were purchased from Santa Cruz Biotechnology (Santa Cruz, CA). The IL-3–dependent murine myeloid cell line 32Dcl3 (kindly provided by Dr Felicitas Rosenthal, Freiburg, Germany) and the murine lymphoid cell line Ba/F3 were cultured in RPMI 1640 supplemented with 10% WEHI-conditioned medium as a source of IL-3, 10% fetal calf serum, and antibiotics at 37°C with 5% CO2. The tyrosine kinase inhibitor D-65476 and its activity toward Flt3 were described previously.18
Patient samples
All patients, whose samples were analyzed, were enrolled into the treatment optimization trial of the AML Cooperative Group (AMLCG) in Germany.19 The 7 control samples were obtained from patients with nonhematologic diseases whose bone marrow aspirates were obtained for diagnostic reasons. Written consent was obtained from all patients.
cDNA construction and gene expression
The construction of murine Flt3 containing the human sequences of the juxtamembrane region has previously been described.6 Briefly, ITD mutations from 3 mutated Flt3 receptor sequences detected in AML blasts as well as wild-type human Flt3 derived from the Oci-AML5 cell line were amplified by reverse transcription–polymerase chain reaction (RT-PCR). PCR products spanning nucleotides 1567 to 2077 of the human Flt3 sequence were substituted into the sequence of murine Flt3. These chimera constructs were named ITD1, ITD2, and ITD3, corresponding to 3 different ITD mutants.6 For the construction of ITD5 and WT2, the full-length coding sequence of Flt3 from a patient with ITD mutation as well as the full-length wild-type sequence from the Oci-AML5 cell line were amplified and cloned as described.18 The constructs were cloned into an expression vector (pAL) under the control of the 5′ long terminal repeat (LTR) of the Moloney murine sarcoma virus (MoMuSV). The constructs were stably transfected into 32Dcl3 cells. Polyclonal cell lines were used for further experiments. The murine Pim-2 construct was obtained from Dr Berns (Netherlands Cancer Institute).20To make a kinase-defective Pim-2, Lys120Met was accomplished by site-directed mutagenesis (Quickchange, Stratagene). These Pim-2 constructs were subcloned into the pAL vector.
Microarray analysis
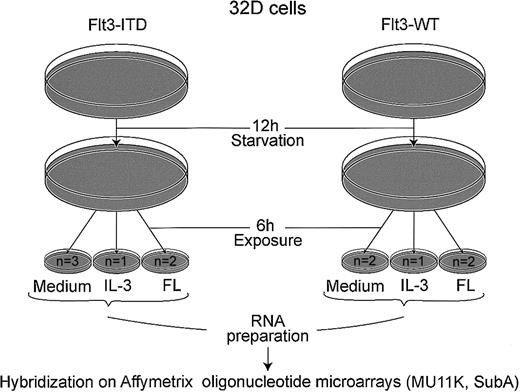
As shown in Figure 1, cells were starved from IL-3 for 12 hours and stimulated with either recombinant murine IL-3 (5 ng/mL) or recombinant human FL (20 ng/mL) for 6 hours. The mRNA was isolated by RNeasy mini kit (Qiagen GmbH, Hilden, Germany). A total of 20 μg mRNA was used to generate the first-strand cDNA with a T7-linked oligo(dT) primer. After second-strand synthesis, in vitro transcription was performed with Enzo BioArray HighYield RNA Transcript Labeling Kit (Enzo Diagnostics, Farmingdale, NY). Biotinylated cRNA (14 μg) was fragmented and hybridized to Affymetrix MU11K SubA arrays (Affymetrix, Santa Clara, CA). The arrays contain probe sets for 6586 murine genes. After washing, the arrays were stained with streptavidin-phycoerythrin (Molecular Probes, Eugene, OR) and biotinylated antistreptavidin antibody (Vector Laboratories, Burlingame, CA) and scanned on a Hewlett Packard scanner. Intensity for each feature of the array was captured with the Genechip software (Affymetrix).
Schematic representation of microarray experiment.
Cells (107) were used for each hybridization; n = number of microarrays hybridized. Following starvation from IL-3 for 12 hours, cells were incubated for 6 hours with either medium without cytokines, 5 ng/mL rmIL-3, or 20 ng/mL FL, as indicated.
Schematic representation of microarray experiment.
Cells (107) were used for each hybridization; n = number of microarrays hybridized. Following starvation from IL-3 for 12 hours, cells were incubated for 6 hours with either medium without cytokines, 5 ng/mL rmIL-3, or 20 ng/mL FL, as indicated.
Statistical methods
If not otherwise indicated, all statistical analyses were performed with the SPSS 10 software package (Munich, Germany). Initial analysis of microarray intensity data was performed by the Affymetrix Software Microarray Suite 4.0. Statistical analyses of the intensity data to identify differentially regulated genes were performed with a 2-class algorithm of the significance analysis of microarrays (SAM) software from Stanford University.21Average linkage clustering was performed with the use of centered correlations to calculate the distances of genes and samples, after all gene expression data had been log transformed, normalized, and median centered in relation to the variation of expression over all samples and genes. The software from Eisen et al22 was used for this purpose. Only genes were included in the cluster analyses that had been judged by the SAM algorithm to be significantly regulated by the ITD mutations, that were significantly expressed (average difference > 200) in either the wild-type or the ITD group, and that showed a relative expression (ratio of average difference) in one versus the other group of at least 2.
Real time RT-PCR
Total RNA was isolated from 32D cells expressing Flt3-WT or Flt3-ITD as described in “cDNA construction and gene expression.” The cDNA was diluted to 200 μL with ddH20, and 2.5 μL was used for each PCR reaction. The quantification of mRNA levels was carried out with the use of a real time fluorescence detection method as described before.23Relative gene expression levels were calculated using standard curves generated by the serial dilutions of cDNA from IL-3–stimulated 32D cells or U937 cells. All samples were independently analyzed at least twice for each gene. The housekeeping gene GAPDHserved as an additional control for the cDNA quality. For patient samples, blasts were enriched from bone marrow samples by density centrifugation of heparinized aspirates at the time of diagnosis and frozen at −80°C until the experiments were performed.
Western blot analysis
32D cells transfected with Flt3 constructs were starved from IL-3 and stimulated with cytokines for the indicated times. Subsequently, cells were washed once with ice-cold phosphate-buffered saline (PBS) and lysed with buffer containing 50 mM HEPES (N-2-hydroxyethylpiperazine-N′-2-ethanesulfonic acid) (pH 7.4), 10% glycerol, 150 mM NaCl, 1% Triton X-100, 1 mM EDTA (ethylenediaminetetraacetic acid), 1 mM EGTA (ethyleneglycoltetraacetic acid), 50 μM ZnCl2, 25 mM NaF, proteinase inhibitors (Complete, Boehringer Mannheim, Germany), 1 μM pepstatin, and 1 mM sodium orthovanadate. Cell lysates were clarified at 20 000g for 20 minutes. Total cell lysates were resuspended in sodium dodecyl sulfate (SDS) sample buffer, heated, and separated by SDS polyacrylamide gel electrophoresis (PAGE). Gels were blotted on Immobilon P membrane (Millipore, Bedford, MA) and stained with the indicated antibody. Antibody binding was detected by incubation with a horseradish peroxidase (HRP)–labeled secondary antibody followed by chemiluminescence detection (ECL-Plus; Amersham Pharmacia Biotech, United Kingdom).
Clonal growth in methylcellulose
Flt3-ITD–expressing 32Dcl3 cells were electroporated with 15 μg of the kinase-defective Pim-2 expression vector or control vector (not containing mammalian selection markers) with 2 μg pcDNA3.1, which confers G418 resistance. One day after electroporation, cells were seeded at a concentration of 1 × 105 cells per dish in 1 mL culture mix containing Iscoves modified Dulbecco medium (IMDM; Life Technologies, Grand Island, NY), 1% methylcellulose, 20% fetal calf serum (FCS), and 0.6 mg/mL G418. The colonies were counted on day 8.
Luciferase assay
To determine Pu.1 activity, we used the luciferase reporter construct for Pu.1, 3 × WT-MHC-luc, which had 3 tandem repeats of a sequence 5′-AAAGAGGAACTTGG-3′ just upstream of a minimal JunB promoter in JunB-MP-luc.24 32D/Flt3-WT or 32D/Flt3-ITD cells were transfected with 3 × WT-MHC-luc together with pRLnull, an expression vector of renilla luciferase, by electroporation. Luciferase assay was performed with the Dual Luciferase Reporter System (Promega, Madison, WI). The values of firefly luciferase were normalized to the respective values ofrenilla luciferase.
Results
To identify target genes of Flt3 mutations in myeloid cells, we analyzed the global gene expression of 32D cells transfected with different Flt3 isoforms by oligonucleotide microarrays. Following 12 hours of starvation, the cells were incubated for 6 hours in the presence of growth factors as shown in Figure1. We hybridized 11 microarrays (Mu 11k Sub A), 6 with cRNA from cells stably expressing Flt3-ITD and 5 with wild-type Flt3.
Significance analyses of microarray data
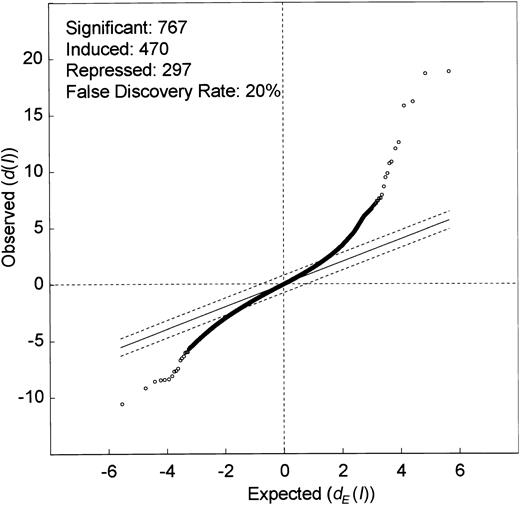
Genes, whose expression levels were altered by ITD mutations, were identified using the significance analysis of microarrays (SAM) algorithm reported by Tusher et al.21 We were interested in identifying genes that were differentially expressed in ITD-positive versus ITD-negative cases, irrespective of the presence of Flt3 ligand. Thus, we included into this first analysis all samples that had been stimulated with Flt3 ligand but excluded samples that had been stimulated with IL-3. The 9 microarrays were separated into 2 groups to compare: one group contained 4 arrays of wild-type Flt3 constructs with or without FL and another group contained 5 arrays of ITD1, ITD2, and ITD3 without FL as well as ITD1 and ITD2 with FL. The calculation of SAM led to a scatter plot of the observed relative difference d(I) versus the expected relative differencedE(I) (Figure2). At a Δ value of 0.77, illustrated by the broken lines, 767 genes were considered to be differentially expressed (470 induced and 297 repressed). These differentially expressed genes contained 471 expressed sequence tags (ESTs) and 296 known cDNAs.
Microarray analysis of ITD transfectants reveals multiple differentially regulated genes.
Significance analysis of the microarray data (SAM analysis) between one group containing 4 microarrays of Flt3-WT with or without Flt3 ligand and another group containing 5 microarrays of Flt3-ITD with or without Flt3 ligand. The scatter plot of the observed difference(d(I)) versus the expected relative difference(dE(I)) is shown. The broken lines are drawn at a distance of Δ = 0.77 from the solid line that indicatesd(I) = dE(I). Genes outside the broken lines are regarded as genes with significant changes in expression, yielding 767 genes. The median estimated false discovery rate is 20%. Dots above the broken line indicate genes induced by Flt3-ITD (470), and dots below the broken line indicate genes that are suppressed by the ITD mutations (297).
Microarray analysis of ITD transfectants reveals multiple differentially regulated genes.
Significance analysis of the microarray data (SAM analysis) between one group containing 4 microarrays of Flt3-WT with or without Flt3 ligand and another group containing 5 microarrays of Flt3-ITD with or without Flt3 ligand. The scatter plot of the observed difference(d(I)) versus the expected relative difference(dE(I)) is shown. The broken lines are drawn at a distance of Δ = 0.77 from the solid line that indicatesd(I) = dE(I). Genes outside the broken lines are regarded as genes with significant changes in expression, yielding 767 genes. The median estimated false discovery rate is 20%. Dots above the broken line indicate genes induced by Flt3-ITD (470), and dots below the broken line indicate genes that are suppressed by the ITD mutations (297).
Cluster analyses of ITD-regulated genes
We used cluster analyses to identify transcriptional programs induced by ITD mutations versus wild-type Flt3. To restrict the analysis to relevant genes, we chose only genes that were significant in the SAM analysis and that were regulated at least 2-fold, when the mean expression data of ITD versus wild-type Flt3 were compared. Also, the genes had to be significantly expressed in one of the 2 groups to enter the analyses.
Genes fitting into these categories were analyzed by hierarchical cluster analysis of self-organizing maps (SOMs; Figure3). Two large groups of genes could be distinguished, one repressed by ITD mutations, the other induced. Also, the main clusters formed by the cell lines were separated by the transfected Flt3 isoforms, which indicated that the mutations induce a different expression profile than the wild-type receptor. Here, the analysis was performed using all 11 microarrays. This analysis revealed that IL-3 incubation induced a transcriptional response that differed from ligand-stimulated Flt3. In contrast, the transcriptional profile of IL-3–stimulated ITD-transfected cells did not change in comparison to unstimulated ITD-transfected cells. These results indicated that many genes were coregulated by the mutations and IL-3 but not by the mutations and FL.
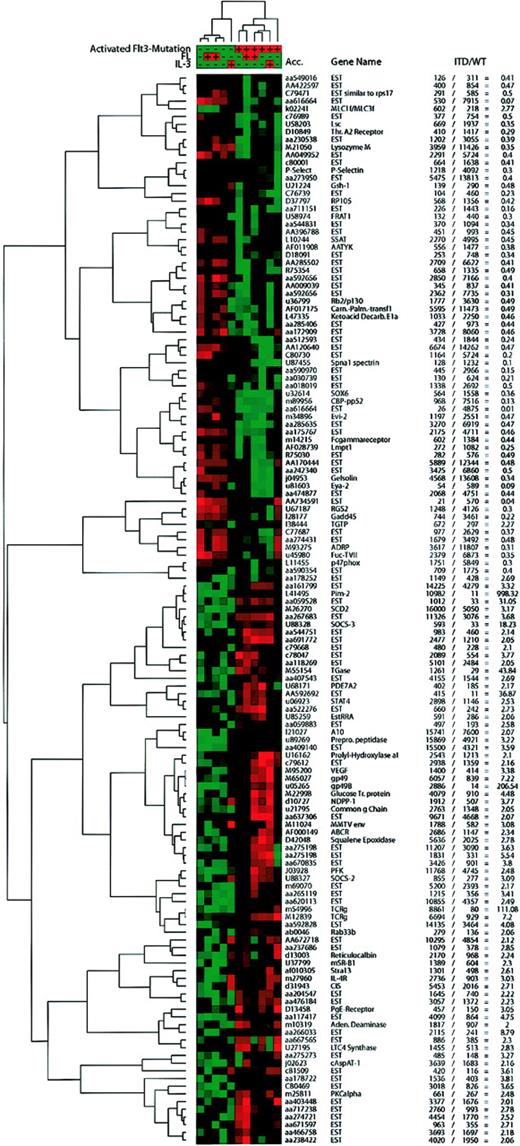
Hierarchical cluster analysis of self-organizing maps.
Using the cluster software,22 we performed hierarchical clustering of a self-organizing map with the genes judged to be significantly regulated by SAM, which were expressed in either group at a significant level (average difference, > 200), and for which the “fold change” ( = mean of ITD/mean of WT average difference) was more than 2 or less than 0.5. IL-3–stimulated samples that had been excluded from SAM analyses were included in this analysis. Values from the Affymetrix software were normalized and median centered before the self-organizing map (SOM) algorithm was applied followed by hierarchical clustering. Red areas depict high expression and green areas depict low expression of a given gene in a given sample. The average difference and fold change values for each gene are presented in the table.
Hierarchical cluster analysis of self-organizing maps.
Using the cluster software,22 we performed hierarchical clustering of a self-organizing map with the genes judged to be significantly regulated by SAM, which were expressed in either group at a significant level (average difference, > 200), and for which the “fold change” ( = mean of ITD/mean of WT average difference) was more than 2 or less than 0.5. IL-3–stimulated samples that had been excluded from SAM analyses were included in this analysis. Values from the Affymetrix software were normalized and median centered before the self-organizing map (SOM) algorithm was applied followed by hierarchical clustering. Red areas depict high expression and green areas depict low expression of a given gene in a given sample. The average difference and fold change values for each gene are presented in the table.
Functional gene categorization
The cDNAs from the SAM analysis were further classified into several categories according to their published function (data not shown). Expectedly, significant numbers (n = 78) of differentially regulated genes are involved in signal transduction and regulation of proliferation and survival. Of note, we also found a significant number (n = 59) of genes that regulate hematopoietic differentiation or that encode proteins involved in the function of differentiated hematopoietic cells. Among the SAM-selected genes regulated by the ITD mutations and categorized as being important for cellular proliferation and survival were Pim-2 and several genes of the suppressor of cytokine signaling (SOCS) family. These genes have been reported to be STAT target genes,25 which is consistent with our earlier report that ITD mutations activate STAT5.6 Also, we saw the repression of several p53 target genes, like GADD45 and BTG-2, and of known inducers of apoptosis in 32D cells like AATYK. Finally, several transcription factors involved in myeloid gene regulation were repressed by Flt3-ITD (see “Specificity of ITD-induced gene regulation”).
Specificity of ITD-induced gene regulation
Because the cluster analyses had revealed that ITD mutations mimic the IL-3 transcriptional response, we were interested in identifying genes specifically regulated by ITD mutations but not by FL or IL-3. First, we plotted the relative gene expression (ITD versus WT) of all 767 genes against the relative gene expression induced by FL or IL-3 in the Flt3-WT cell lines (Figure 4). Again, we observed that the genes were significantly coregulated by the ITD mutations and IL-3 (R = 0.73), whereas a correlation between the ITD mutations and FL was much less pronounced (R = 0.49).
ITD mutations profoundly change gene expression profiles of myeloid progenitor cells.
(A) Scatter plot of the gene regulation values of IL-3 (ratio of IL-3 and medium [MED]) versus Flt3-ITD. (B) Scatter plot of the gene regulation values of ligand-activated Flt3-WT (ratio of FL and medium [MED]) versus Flt3-ITD. R depicts the correlation coefficient of logarithmically transformed expression ratios (Spearman nonparametric correlation).
ITD mutations profoundly change gene expression profiles of myeloid progenitor cells.
(A) Scatter plot of the gene regulation values of IL-3 (ratio of IL-3 and medium [MED]) versus Flt3-ITD. (B) Scatter plot of the gene regulation values of ligand-activated Flt3-WT (ratio of FL and medium [MED]) versus Flt3-ITD. R depicts the correlation coefficient of logarithmically transformed expression ratios (Spearman nonparametric correlation).
We then asked whether the correlation between growth factor incubation and ITD mutations differs between functional groups of genes (Figure5). ITD mutations have similar effects as IL-3 on 32D cell proliferation and survival. Thus, it was not surprising that we found a very strong correlation of the regulation of growth-related genes by the ITD mutations and IL-3 (Figure 5A). Indeed, most genes that had been reported in the literature to be promoting growth were induced by both IL-3 and the ITD mutations, whereas growth-inhibitory genes were corepressed. In contrast, these genes were not coregulated by the mutations and ligand-activated wild-type Flt3.
ITD mutations coregulate growth-related genes but not transcription factors with IL-3.
Scatter plot of IL-3– or FL-induced gene regulation versus ITD-induced gene regulation of functionally defined genes. R depicts the correlation coefficient of logarithmically transformed expression ratios (Spearman nonparametric correlation). Areas that contain differentially regulated genes are circled with continuous lines. Areas that contain coregulated genes are circled with dashed lines. (A) Growth-related genes. (B) Transcription factors.
ITD mutations coregulate growth-related genes but not transcription factors with IL-3.
Scatter plot of IL-3– or FL-induced gene regulation versus ITD-induced gene regulation of functionally defined genes. R depicts the correlation coefficient of logarithmically transformed expression ratios (Spearman nonparametric correlation). Areas that contain differentially regulated genes are circled with continuous lines. Areas that contain coregulated genes are circled with dashed lines. (A) Growth-related genes. (B) Transcription factors.
In contrast to growth-regulating genes, regulation of transcription factor expression differed between ITD mutations and IL-3 (Figure 5B). We concluded that these could be specific target genes of the ITD mutations.
Confirmation of microarray data by real time RT-PCR
The expression of 26 genes, selected based on their regulation by ITD mutations but not by FL, was verified by real time RT-PCR (Table1). After starvation, Flt3 wild-type– and ITD-transfected 32D cells were stimulated with either IL-3 or FL. RNA was isolated at 0, 3, 6, 12, 24, and 36 hours. As shown in Table 1, 18 of 26 analyzed genes showed significant regulation by Flt3-ITD and, thus, positively confirmed the microarray results.
Confirmation of microarray results by real time RT-PCR analysis
| Gene . | Microarray . | Real time RT-PCR . | |||||||
|---|---|---|---|---|---|---|---|---|---|
| 32D . | Ba/F3 . | 32D . | |||||||
| ITD/WT . | IL-3/Med . | FL/Med . | ITD/WT . | P . | IL-3/Med . | FL/Med . | ITD/WT . | D-65476/Med . | |
| Up-regulated | |||||||||
| PIM2 | 998.30 | 352.80 | 1.00 | 42.9 | .001 | 7.7 | 0.51 | 9.73 | 0.052 |
| GP49B | 206.50 | 43.60 | 1.54 | 226.4 | .006 | 61.5 | 12-0.2 | 0.28 | 0.047 |
| SOCS-2 | 3.10 | 3.30 | 0.70 | 8.4 | .009 | 5.4 | 2.1 | 2.32 | 0.72-1.35 |
| SOCS-3 | 18.23 | 31.05 | 4.92 | 6.19 | .009 | 3.3 | 1.7 | 5.83 | 0.042 |
| PGE-R | 3.10 | 2.70 | 0.60 | 1.68 | .019 | 2.47 | 0.71 | 0.4-27.55 | 0.4-1.72 |
| Down-regulated | |||||||||
| maf-B | 0.10 | 0.50 | 1.60 | 0.6 | .004 | 1.8 | 1.4 | 2.07 | 0.7-1.74 |
| PU.1 | 0.60 | 1.10 | 1.30 | 0.31 | .006 | 2.05 | 3.56 | 0.33 | 2.00 |
| RARγ | 0.60 | 1.20 | 1.10 | 0.378 | .011 | 0.71 | 3.46 | 0.04 | 6.3 |
| MAID | 0.66 | 0.53 | .014 | 0.61 | 1.44 | 0.5 | 1.84 | ||
| GADD45 | 0.22 | 0.16 | 0.69 | 0.35 | .028 | 0.18 | 1.61 | 0.12 | 4.2 |
| RB2/p130 | 0.49 | 0.15 | 0.58 | 0.55 | .002 | 0.52 | 1.98 | 0.61 | 3.9 |
| p16 ink 4A | 0.75 | 0.90 | 1.00 | 0.19 | .011 | 6.47 | 5.35 | 77.17 | 2.00 |
| Frat1 | 0.30 | 0.02 | 0.68 | 0.143 | .028 | 0.25 | 1 | 0.34 | 4.7 |
| LSC | 0.40 | 0.90 | 1.00 | 0.37 | .027 | 3.16 | 4.31 | 0.4 | 1.85 |
| RGS2 | 0.30 | 0.30 | 1.00 | 0.33 | .002 | 0.26 | 1.8 | 0.01 | 1.92 |
| Gα15 | 0.60 | 1.10 | 1.20 | 0.265 | .022 | 2.27 | 3.04 | 0.79-1.28 | 1.22 |
| TISS11 | 0.57 | 0.50 | 0.90 | 0.28 | .016 | 1.1 | 3.25 | 0.35 | 1.78 |
| Evi-2 | 0.47 | 1.01 | 1.37 | 0.37 | .005 | 0.47 | 1.31 | 0.66 | 2.54 |
| Gene . | Microarray . | Real time RT-PCR . | |||||||
|---|---|---|---|---|---|---|---|---|---|
| 32D . | Ba/F3 . | 32D . | |||||||
| ITD/WT . | IL-3/Med . | FL/Med . | ITD/WT . | P . | IL-3/Med . | FL/Med . | ITD/WT . | D-65476/Med . | |
| Up-regulated | |||||||||
| PIM2 | 998.30 | 352.80 | 1.00 | 42.9 | .001 | 7.7 | 0.51 | 9.73 | 0.052 |
| GP49B | 206.50 | 43.60 | 1.54 | 226.4 | .006 | 61.5 | 12-0.2 | 0.28 | 0.047 |
| SOCS-2 | 3.10 | 3.30 | 0.70 | 8.4 | .009 | 5.4 | 2.1 | 2.32 | 0.72-1.35 |
| SOCS-3 | 18.23 | 31.05 | 4.92 | 6.19 | .009 | 3.3 | 1.7 | 5.83 | 0.042 |
| PGE-R | 3.10 | 2.70 | 0.60 | 1.68 | .019 | 2.47 | 0.71 | 0.4-27.55 | 0.4-1.72 |
| Down-regulated | |||||||||
| maf-B | 0.10 | 0.50 | 1.60 | 0.6 | .004 | 1.8 | 1.4 | 2.07 | 0.7-1.74 |
| PU.1 | 0.60 | 1.10 | 1.30 | 0.31 | .006 | 2.05 | 3.56 | 0.33 | 2.00 |
| RARγ | 0.60 | 1.20 | 1.10 | 0.378 | .011 | 0.71 | 3.46 | 0.04 | 6.3 |
| MAID | 0.66 | 0.53 | .014 | 0.61 | 1.44 | 0.5 | 1.84 | ||
| GADD45 | 0.22 | 0.16 | 0.69 | 0.35 | .028 | 0.18 | 1.61 | 0.12 | 4.2 |
| RB2/p130 | 0.49 | 0.15 | 0.58 | 0.55 | .002 | 0.52 | 1.98 | 0.61 | 3.9 |
| p16 ink 4A | 0.75 | 0.90 | 1.00 | 0.19 | .011 | 6.47 | 5.35 | 77.17 | 2.00 |
| Frat1 | 0.30 | 0.02 | 0.68 | 0.143 | .028 | 0.25 | 1 | 0.34 | 4.7 |
| LSC | 0.40 | 0.90 | 1.00 | 0.37 | .027 | 3.16 | 4.31 | 0.4 | 1.85 |
| RGS2 | 0.30 | 0.30 | 1.00 | 0.33 | .002 | 0.26 | 1.8 | 0.01 | 1.92 |
| Gα15 | 0.60 | 1.10 | 1.20 | 0.265 | .022 | 2.27 | 3.04 | 0.79-1.28 | 1.22 |
| TISS11 | 0.57 | 0.50 | 0.90 | 0.28 | .016 | 1.1 | 3.25 | 0.35 | 1.78 |
| Evi-2 | 0.47 | 1.01 | 1.37 | 0.37 | .005 | 0.47 | 1.31 | 0.66 | 2.54 |
The cell lines 32D and Ba/F3 stably expressing Flt3-WT or Flt3-ITD were cultured under the same conditions as in the microarray experiments depicted in Figure 1. Subsequently the expression of 23 genes showing significant regulation in the microarray experiments was analyzed by real time RT-PCR analyses. The numbers given represent the ratios of glyceraldehyde phosphate dehydrogenase (GAPDH)–normalized expression values (PCR) or of “average difference” values (microarrays). For significance analyses, a Student t test for paired samples on multiple experiments at different time points of FL stimulation was performed. The last column shows the ratio of expression in 32D/Flt3-ITD cells under the influence of 0.5 μM D-65476 compared with controls treated with solvent (0.01% dimethyl sulfoxide [DMSO]). Each number is representative of at least 3 independent experiments.
To exclude the possibility that the observed effects are restricted to 32D cells, we analyzed the expression of 18 regulated genes in another IL-3–dependent murine hematopoietic progenitor cell line, Ba/F3, that we stably transfected with either Flt3-WT or Flt3-ITD. Expression of 13 of the 18 (72%) genes tested was altered by Flt3-ITD in these cells in the same way as in 32D cells.
To confirm that the changes in gene expression profiles were based on direct effects of Flt3 and did not occur because of a commonly selected phenotype, we analyzed the effects of Flt3 inhibition on the expression of the regulated genes. To inhibit Flt3 activity, we used a tyrosine kinase inhibitor (D-65476) that we previously described as having inhibitory activity toward a number of tyrosine kinases with selectivity toward Flt3. Specifically, the substance inhibited Flt3-mediated growth and survival of 32D cells, without influencing effects mediated by IL-3.18 Here, we treated 32D cells expressing Flt3-ITD with D-65476 for 6 hours and subsequently analyzed its effect on the expression of ITD-regulated genes. Fifteen of the 18 genes confirmed to be regulated by Flt3-ITD were regulated in the opposite way by the inhibitor. These findings provide evidence that the regulation of target gene expression required ongoing activity of the Flt3 kinase.
Pim-2 is a functionally important target gene of Flt3-ITD
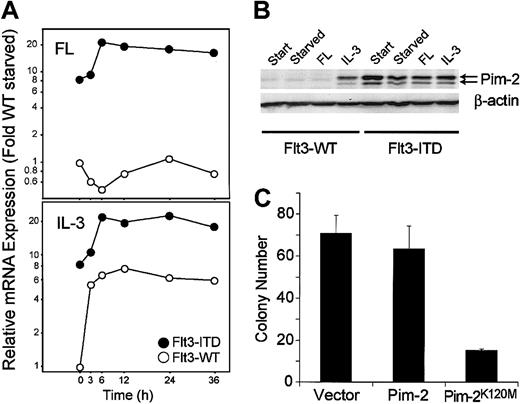
One of the genes up-regulated by the ITD mutations was the serine/threonine kinase Pim-2 (Figure6). Stimulation of 32D cells with IL-3 but not with FL also induced enhanced expression of this mRNA. By Western blot, we confirmed that induction of the Pim-2 mRNA was accompanied by increased protein expression as well (Figure 6B). We then were interested in whether Pim-2 activity was necessary for ITD-mediated cellular transformation. In earlier results, we had shown that Flt3-ITD but not ligand-activated Flt3-wild-type can support 32D cell colony growth.6 Therefore, we were interested in the effect of dominant-negative Pim-2 on 32D colony growth. We constructed a Pim-2 molecule with a mutation at residue Lys120Met. The homologue of this mutation in Pim-1 has been reported to abolish Pim-1 catalytic activity and to inhibit endogenous Pim-1 kinase activity in a dominant-negative manner.26 Recently, the same mutation in Pim-2 has been described to abolish Pim-2 kinase activity.27 When cotransfected with the ITD mutations, kinase-defective Pim-2 inhibited ITD-mediated colony growth by approximately 50% (Figure 6C).
Pim-2 is an ITD target gene involved in ITD-mediated transformation.
(A-B) Flt3-ITD induces Pim-2 mRNA and protein selectively. (A) mRNA was obtained from 32Dcl3/flt3-ITD (●) and 32Dcl3/Flt3-WT (○) cells, which were stimulated with FL or IL-3 for the time indicated. The amount of Pim-2 mRNA was analyzed in each sample by real time RT-PCR. The ratio of expression compared with flt3-WT after normalization to GAPDH levels is shown. (B) Total lysates of 32D/flt3-ITD and 32D/flt3-WT stimulated with or without cytokines for 24 hours were electrophoresed and blotted with anti–Pim-2 antibody. The time point “start” indicates the starting point of the starvation period. The concentrations of FL and IL-3 were 50 ng/mL and 1 ng/mL, respectively. (C) Pim-2K120M inhibits colony formation by Flt3-ITD. 32D/Flt3-ITD was electroporated with 15 μg of the expression vectors for kinase-defective Pim-2K120M or control vectors and 2 μg pcDNA3. One day after electroporation, cells were seeded on 1% methylcellulose. The colonies were counted on day 8. The mean colony numbers and standard deviation of 3 experiments is shown.
Pim-2 is an ITD target gene involved in ITD-mediated transformation.
(A-B) Flt3-ITD induces Pim-2 mRNA and protein selectively. (A) mRNA was obtained from 32Dcl3/flt3-ITD (●) and 32Dcl3/Flt3-WT (○) cells, which were stimulated with FL or IL-3 for the time indicated. The amount of Pim-2 mRNA was analyzed in each sample by real time RT-PCR. The ratio of expression compared with flt3-WT after normalization to GAPDH levels is shown. (B) Total lysates of 32D/flt3-ITD and 32D/flt3-WT stimulated with or without cytokines for 24 hours were electrophoresed and blotted with anti–Pim-2 antibody. The time point “start” indicates the starting point of the starvation period. The concentrations of FL and IL-3 were 50 ng/mL and 1 ng/mL, respectively. (C) Pim-2K120M inhibits colony formation by Flt3-ITD. 32D/Flt3-ITD was electroporated with 15 μg of the expression vectors for kinase-defective Pim-2K120M or control vectors and 2 μg pcDNA3. One day after electroporation, cells were seeded on 1% methylcellulose. The colonies were counted on day 8. The mean colony numbers and standard deviation of 3 experiments is shown.
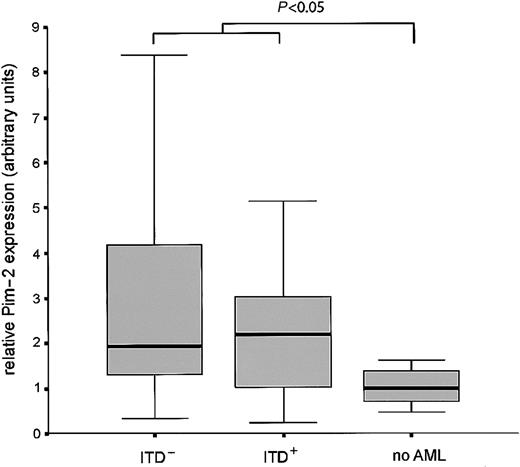
Finally, we analyzed the expression of Pim-2 mRNA in bone marrow samples from 84 patients newly diagnosed with AML in comparison to healthy bone marrow. Pim-2 expression was significantly induced in the AML samples. No difference in the expression level was observed between ITD-positive versus -negative cases (Figure7).
Pim-2 is overexpressed in leukemic bone marrow.
Bone marrow from 84 patients with AML was analyzed for Pim-2 expression by real time RT-PCR and compared with healthy bone marrow. Box plots are shown for the relative expression levels of Pim-2 normalized for GAPDH in Flt3-ITD positive, Flt3-ITD negative AML samples, and healthy bone marrow. The difference of expression between leukemic samples and healthy bone is statistically significant (P < .05, Mann-Whitney U test).
Pim-2 is overexpressed in leukemic bone marrow.
Bone marrow from 84 patients with AML was analyzed for Pim-2 expression by real time RT-PCR and compared with healthy bone marrow. Box plots are shown for the relative expression levels of Pim-2 normalized for GAPDH in Flt3-ITD positive, Flt3-ITD negative AML samples, and healthy bone marrow. The difference of expression between leukemic samples and healthy bone is statistically significant (P < .05, Mann-Whitney U test).
Taken together, Pim-2 is induced by the ITD mutations on the mRNA and the protein levels, and Pim-2 function is important for ITD-mediated cellular transformation in 32D cells. Expression of this kinase is significantly up-regulated in AML.
Flt3-ITD represses expression and function of Pu.1 and c/EBPα
The strong differences observed in the regulation of transcription factors by the ITD mutations versus growth factor stimulation prompted us to analyze these effects in more detail. First, we confirmed the microarray results for several transcription factors by real time RT-PCR (Table 1). Pu.1 has been reported to play a crucial role in monocytic and B-cell lineage differentiation.28,29 Similar functions have been ascribed to Flt3.1,30 Also, although not represented on the microarray, we were interested in the effects of the ITD mutations on the expression and function of C/EBPα that has been shown to be important for myeloid differentiation and that is frequently inactivated in myeloid leukemias.31
Therefore, we analyzed ligand-stimulated expression of Pu.1 in a time-course experiment (Figure 8A). In Flt3-WT–expressing cells, Flt3 ligand induced expression of Pu.1 at 3 hours, with Pu.1 mRNA levels steadily increasing over 36 hours. However, in ITD-expressing cells, neither Flt3 ligand nor IL-3 significantly induced Pu.1 expression. Thus, the transcriptional regulation of Pu.1 clearly differed between Flt3-WT and -ITD.
ITD mutations suppress PU.1 expression and function.
(A) Real time RT-PCR analysis of PU.1 mRNA from 32D/Flt3-WT (○) and 32D/Flt3-ITD (●) stimulated with IL-3 or FL as indicated for 36 hours or more. The expression level is shown in the same way as in Figure 6. (B) Western blot analysis of Pu.1 protein from 32D/Flt3-WT and 32D/Flt3-ITD stimulated with cytokines for 48 hours. (C) Analysis of Pu.1-transactivating activity on a Pu.1-responsive promoter construct, 3 × WT-MHC-luc. 32D/Flt3-WT and 32D/Flt3-ITD were starved from IL-3 for 2 hours and transfected with 3 × WT-MHC-Luc and pRLnull. After stimulation with the indicated cytokines for 16 hours, cells were lysed, and the luciferase activity of the cell lysates was determined. The relative luciferase activity was normalized to renilla luciferase activity (activity of cotransfected pRLnull vector). med indicates medium, no cytokines.
ITD mutations suppress PU.1 expression and function.
(A) Real time RT-PCR analysis of PU.1 mRNA from 32D/Flt3-WT (○) and 32D/Flt3-ITD (●) stimulated with IL-3 or FL as indicated for 36 hours or more. The expression level is shown in the same way as in Figure 6. (B) Western blot analysis of Pu.1 protein from 32D/Flt3-WT and 32D/Flt3-ITD stimulated with cytokines for 48 hours. (C) Analysis of Pu.1-transactivating activity on a Pu.1-responsive promoter construct, 3 × WT-MHC-luc. 32D/Flt3-WT and 32D/Flt3-ITD were starved from IL-3 for 2 hours and transfected with 3 × WT-MHC-Luc and pRLnull. After stimulation with the indicated cytokines for 16 hours, cells were lysed, and the luciferase activity of the cell lysates was determined. The relative luciferase activity was normalized to renilla luciferase activity (activity of cotransfected pRLnull vector). med indicates medium, no cytokines.
The specific repression of Pu.1 by mutationally activated Flt3 in comparison to ligand-activated Flt3 was also confirmed by Western blot analyses (Figure 8B). Finally, we assessed the transcriptional activity of Pu.1, using a Pu.1 reporter luciferase construct. At baseline, Flt3-ITD displayed slightly enhanced Pu.1 activity compared with Flt3-WT. However, on FL stimulation, Pu.1 activity in 32D cells transfected with Flt3-WT increased more than 5-fold, whereas Flt3 ligand did not have any effect on Pu.1 activity in cells expressing Flt3-ITD (Figure 8C). These results suggested that ITD mutations inhibit Flt3-mediated induction of Pu.1 expression and function.
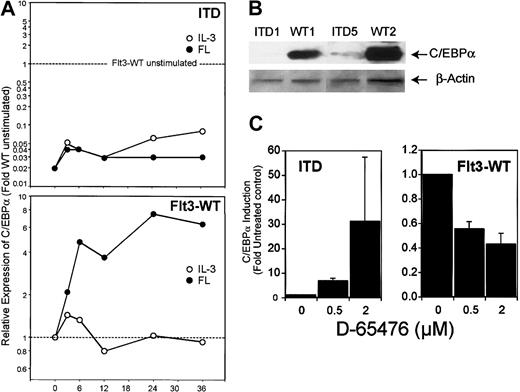
Analysis of the expression of C/EBPα by real time RT-PCR and by Western blot revealed similar results (Figure9): incubation with IL-3 did not change C/EBPα expression, whereas Flt3 ligand induced it up to 7.5-fold. In contrast, C/EBPα levels in Flt3-ITD–transfected cells were significantly lower. Also, in Ba/F3 cells expressing Flt3-ITD, C/EBPα expression was 5-fold lower compared with Ba/F3-cells expressing Flt3-WT (data not shown). We were further interested in how inhibition of Flt3 kinase activity would influence C/EBPα transcription under ligand stimulation of Flt3-WT compared with Flt3-ITD. FL-stimulated 32D/Flt3-WT and 32D/Flt3-ITD were treated with different amounts of a kinase inhibitor that we described to be an inhibitor of Flt3, D-65476.18 As shown in Figure 9C, inhibition of ligand-activated wild-type Flt3 caused a decrease of C/EBPα expression of 50% or more in a dose-dependent manner. In contrast, inhibition of mutation-activated Flt3-ITD led to up-regulation of C/EBPα up to 30-fold.
ITD mutations decrease C/EBPα expression.
(A) Real time RT-PCR analysis of 32D/Flt3-WT and 32D/Flt3-ITD stimulated with FL (●) or IL-3 (○) for the indicated times (hours). The relative expression was calculated in the same way as described for Figure 6. (B) Western blot analysis for C/EBPα from 32D cells expressing wild-type or ITD isoforms of Flt3. All cells were cytokine and serum starved for 12 hours prior to preparing the lysates for Western blot analyses. (C) Real time RT-PCR analysis for C/EBPα of FL-stimulated 32D/Flt3-WT and 32D/Flt3-ITD treated with the Flt3 kinase inhibitor D-65476. Cells were exposed to the indicated concentration of inhibitor for 12 hours. Mean and standard deviation of 2 independent experiments are shown.
ITD mutations decrease C/EBPα expression.
(A) Real time RT-PCR analysis of 32D/Flt3-WT and 32D/Flt3-ITD stimulated with FL (●) or IL-3 (○) for the indicated times (hours). The relative expression was calculated in the same way as described for Figure 6. (B) Western blot analysis for C/EBPα from 32D cells expressing wild-type or ITD isoforms of Flt3. All cells were cytokine and serum starved for 12 hours prior to preparing the lysates for Western blot analyses. (C) Real time RT-PCR analysis for C/EBPα of FL-stimulated 32D/Flt3-WT and 32D/Flt3-ITD treated with the Flt3 kinase inhibitor D-65476. Cells were exposed to the indicated concentration of inhibitor for 12 hours. Mean and standard deviation of 2 independent experiments are shown.
Because Pu.1 and C/EBPα are both involved in myelomonocytic differentiation programs, we were interested in the effects of ITD mutations on granulocyte colony-stimulating factor (G-CSF)–induced differentiation of 32D cells. We analyzed the expression of the granulocytic surface antigen CD11b on the different transfectants after continuous incubation with G-CSF for 12 days. Whereas Flt3-WT cells expressed this antigen after differentiation-inducing treatment, no CD11b expression could be detected on the surface of 32D-Flt3-ITD cells (data not shown). These results indicate that the ITD mutations inhibited the expression and function of transcription factors involved in myeloid differentiation.
Discussion
Genome-wide analyses of target genes of Flt3-ITD revealed striking differences in signal transduction of mutationally versus ligand-activated Flt3. The ITD receptor mutations induce a transcriptional program that resembles activation of the IL-3 receptor. Other parts of the transcriptional response to these mutations are novel and counteract the transcriptional response to ligand-activated Flt3.
The targeted analysis of the transcriptional consequences of a relevant molecular aberration in a cell line has several key advantages over a direct analysis of AML samples: first, it is more likely to identify pathway-specific changes in expression, because the genetic background in a cell line is well defined; second, many different pathways may induce changes in expression levels of genes that are relevant in leukemic transformation. Therefore, it is difficult to trace the pathway back to its origin from expression data in primary AML blasts. The analysis of our data by SAM provided a statistically sound basis for evaluation of the single genes that were differentially regulated. The high rate of confirmation of differential expression by real time RT-PCR matched exactly the statistical probabilities from the analysis. Further confirmation of our data came from the high correlation of ITD-regulated growth-related genes with IL-3–regulated genes that were as expected, given the similar effect of the mutations and IL-3 on 32D cell proliferation and survival.
The time point of the microarray experiments was chosen at 6 hours after cytokine addition, to avoid comparison of the Flt3-WT immediate early response with the late response to constitutively, long-term activated Flt3-ITD. This approach led to confirmation of most observed expression differences in time course experiments of up to 36 hours.
Several genes in our analyses were specifically regulated by Flt3-ITD and not by Flt3-WT. Interestingly, we also observed genes that were unregulated by Flt3-WT and repressed by Flt3-ITD. This finding suggests that qualitatively different signals are emitted by the 2 receptor isotypes. Dose or time effects of Flt3-WT versus Flt3-ITD do not explain the observed antagonist effect of Flt3-ITD on some Flt3-WT target genes.
In the current study, we demonstrate that several known target genes of STAT proteins are induced by ITD mutations. This confirms our previous finding that a major difference of signal transduction between Flt3-WT and Flt3-ITD is constitutive STAT5 activation.6 The aberrant activation of STAT or Janus kinase (JAK)/STAT pathways is now recognized as a common characteristic of several hematopoietic malignancies. It is thought to be especially important in the transformation by activating tyrosine kinase receptor mutations such as KitD816V and the TEL-PDGFR.32-34Although wild-type Kit as well as Flt3-WT has been reported to activate STAT pathways, the activity appeared not to be strong enough to induce the transcription of relevant target genes.4,6 35 These results from several groups suggested that full activation of STAT pathways is not necessary for the normal function of wild-type receptor tyrosine kinases expressed on primitive hematopoietic cells. Rather, STAT activation by tyrosine kinases in hematopoietic cells seems to be associated with their aberrant, oncogenic signaling.
In our analyses, among the STAT3/5 target genes, Pim-2showed strong induction in ITD mutations. The colony assays also revealed an important role in Flt3-ITD–mediated transformation. Pim-1, a protein closely related to Pim-2, was not induced by Flt3-ITD, although it is also known to be a STAT3/5 target gene in other cells, such as BA/F3 cells.25 26 The expression of Pim-1 was not induced by IL-3 in our cells either, suggesting that Pim-1 expression may be already deregulated in 32D cells by other mechanisms.
Nevertheless, 32D cells expressing Flt3-WT showed a very low transforming efficiency in colony formation or in transplantation in mice.6 Because Pim-1 and Pim-2 cooperatively induce cellular transformation, high expression of Pim family proteins may be necessary for the transforming activity of Flt3-ITD proteins. Recently, Dhanasekaran et al36 reported that the expression level of Pim-1 was significantly correlated with the clinical outcome of patients with prostate cancer. We found significant overexpression of Pim-2 in primary AML samples, which was not related to the presence or absence of the ITD mutations. On one hand, this finding indicates that additional mechanisms contribute to Pim-2 up-regulation in AML blasts. On the other hand, this finding points to a possible role of Pim-2 as a “signal integrator” in leukemic transformation, with Flt3 mutations being one of several possible mechanisms to induce it. Together with the expression data in our cell line models, we conclude that Pim-2 is a functionally relevant target gene of Flt3-ITD but not of Flt3-WT.
We have also found SOCS2 and SOCS3 specifically induced in 32D/Flt3-ITD, both of which are STAT3/5 target genes and known negative regulators of receptor signaling.37Schultheis et al38 reported that SOCS2 was overexpressed in advanced stages of chronic myeloid leukemia, suggesting that the activation of STAT pathways is associated with disease transformation. These results further imply the possible role of STAT pathways as targets for therapeutic intervention.
We provide evidence for the first time that the ITD mutations suppress the expression and function of transcription factors important for myeloid differentiation.
Although ligand-activated wild-type Flt3 induced expression of Pu.1 and C/EBPα, the ITD mutations did not induce expression of Pu.1, with only slight and inconsistent increase of its activity, and severely repressed the expression of C/EBPα. Thus, the effects of Flt3 on these transcription factors were mutation specific. Furthermore, no significant regulation by IL-3 was observed. What are the possible consequences of down-regulation of these 2 transcription factors by the ITD mutations?
Although Flt3 and kit are important receptors for the proliferation and survival of early hematopoietic progenitors, several recent reports suggest that stimulation of normal early hematopoietic progenitors by the Flt3 signal does not enhance but reduces their self-renewal capacity. When early mouse progenitor cells start to express Flt3, they rapidly lose their capacity to give rise to long-term myeloid reconstitution but gain potential to induce lymphocyte development.30 ITD mutations provide chronic stimulation of Flt3 signal transduction to AML blasts. Thus, some mechanisms have to be instrumental to inhibit Flt3-mediated differentiation and restriction of self-renewal in ITD-positive AML blasts.
DeKoter and Singh29 reported that Pu.1 induced monocytic or B-cell lineage commitment of early hematopoietic progenitors, depending on the duration and level of expression. Because Flt3 is important for monocytic and B-cell differentiation, Flt3-WT–mediated Pu.1 induction could be involved in this function, and failure of Flt3-ITD to induce Pu.1 could inhibit differentiation. CD11b (Mac-1), a marker of granulocytic differentiation, has been shown to be regulated by Pu.1.39 Also, G-CSF did not induce CD11b in 32D cells in the presence of Flt3-ITD (data not shown).
Inhibition of C/EBPα is an important feature in myeloid leukemias. The effects of translocation-associated fusion proteins in AML like AML1-ETO and PML-RARα are thought to be partially mediated by inhibition of the expression or function of C/EBPα.31,40,41 Point mutations of C/EBPα have been found in AML.42,43 C/EBPα is essential for granulocytic differentiation39 and has direct antiproliferative effects by inhibition of cdk2 and cdk4 function,44,45 repression of E2F transcription factors,46 and down-regulation of c-myc.47 The strong repression of C/EBPα expression by the ITD mutations is surprising, given that IL-3 does not regulate this factor and that Flt3-WT induced its expression by almost 10-fold.
The block of differentiation is a cardinal feature of AML. It was suggested that the pathogenesis of AML is accomplished cooperatively by 2 different kinds of mutations: the chromosomal translocations work as a differentiation blocker and activated tyrosine kinase receptors such as Flt3-ITD or Kit mutants act by augmenting proliferation and survival of the leukemic progenitors.48 Our results suggest that Flt3-ITD may work as both a differentiation blocker and an augmentator of proliferation and survival. However, AML1-ETO has been reported to augment G-CSF–dependent proliferation and expansion of human hematopoietic stem cells.49 50 Furthermore, in a screen for common target genes of translocation-associated fusion proteins, we found several induced genes that, when overexpressed, led to enhanced proliferation of myeloid progenitors, and not only to a block in cellular differentiation (C.M.-T. et al, unpublished observation, December 2001). Taken together, our results imply that the 2 classes of genetic abnormalities are not as functionally exclusive and cooperative as previously thought. Both the translocations involving transcription factors and the activating tyrosine kinase mutations may act in both differentiation block and proliferation enhancement.
In conclusion, Flt3-ITD mutations induce a transcriptional program that is fundamentally different from the program induced by Flt3-WT. The Flt3-ITD mutations mimic IL-3 activation and activate genes involved in cellular growth and proliferation. In addition, Flt3-ITD regulates transcription factors independent of the IL-3–mimicking effects and antagonizes the activity of wild-type Flt3 on myeloid differentiation programs. The combination of both mechanisms is the basis for the strong oncogenic function of ITD mutations.
We thank Beate Surmann, Marion Baas, and Silvia Klümpen for their excellent technical assistance; Anton Berns for the Pim-2 expression construct; and the AMLCG study group for providing the bone marrow samples. We also thank Sven Diederichs and Christian Brandts for critical review of the manuscript and for fruitful discussions.
Prepublished online as Blood First Edition Paper, December 5, 2002; DOI 10.1182/blood-2002-06-1677.
Supported by grants from the Deutsche Forschungsgemeinschaft (Se 600/2, Mu 1328/2-I), the IZKF (H4) and the IMF-Program (Mü429926, Mü529905, SE119908) at the University of Münster, the José-Carreras Leukemia Foundation, and the Deutsche Krebshilfe (10-1539-MÜ), as well as by grants from the Japanese Ministry of Education, Culture, Sports, Science and Technology and the Japan Society for the Promotion of Science (M.M.).
M.M. and J.S. contributed equally to this study.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 U.S.C. section 1734.
References
Author notes
Hubert Serve, Department of Medicine, Hematology and Oncology, University of Münster, Albert-Schweitzer-Strasse 33, 48129 Münster, Germany; e-mail:serve@uni-muenster.de.


![Fig. 4. ITD mutations profoundly change gene expression profiles of myeloid progenitor cells. / (A) Scatter plot of the gene regulation values of IL-3 (ratio of IL-3 and medium [MED]) versus Flt3-ITD. (B) Scatter plot of the gene regulation values of ligand-activated Flt3-WT (ratio of FL and medium [MED]) versus Flt3-ITD. R depicts the correlation coefficient of logarithmically transformed expression ratios (Spearman nonparametric correlation).](https://ash.silverchair-cdn.com/ash/content_public/journal/blood/101/8/10.1182_blood-2002-06-1677/4/m_h80834133004.jpeg?Expires=1769157804&Signature=sOiYG4ALcehmyQEAL6qWaUm4HmXJfw6SqojXkoFb8aeDi7j0Y~1BiU-VlrgMYWEDnkv2AUc-eeZ2KJMUWLtbc0iMAQdJ6ReV3togFW7ObfHd5Hs3jf9jz8X4cXLq55i7EE8ooAW-pm6EbUsU2evj3EGdH57YpScYHdr3uB4zEJmg7D05sthlVv42h~-V2R3p0OjDGoBom4N371NyhNJmoLjjJj8206PtMHQv8Jrl-0XRUNMZM7iPjPsno0pJiAoZxdpnqPCbwRMBYIlssyaaymr5sHqhKgttsFrfVY2TOvcQ4hQSXqQBVo4kMKoFd43Vwgod8agCXJq98Gyh55uyNA__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)


This feature is available to Subscribers Only
Sign In or Create an Account Close Modal