SH2–containing inositol 5′-phosphatase (SHIP) modulates the activation of immune cells after recruitment to the membrane by Shc and the cytoplasmic tails of receptors. A novel SHIP isoform of approximately 104 kd expressed in primitive stem cell populations (s-SHIP) is described. It was found that s-SHIP is expressed in totipotent embryonic stem cells to the exclusion of the 145-kd SHIP isoform expressed in differentiated hematopoietic cells. s-SHIP is also expressed in primitive hematopoietic stem cells, but not in lineage-committed hematopoietic cells. In embryonic stem cells, s-SHIP partners with the adapter protein Grb2 without tyrosine phosphorylation and is present constitutively at the cell membrane. It is postulated that s-SHIP modulates the activation threshold of primitive stem cell populations.
Introduction
Engagement of receptors on the surface of mammalian cells results in the activation of phosphatidylinositol 3-kinase (PI3K) and phosphorylation of phosphatidylinositols on the cytoplasmic side of the cell membrane.8,15 The generation of phosphatidylinositol 3,4,5-trisphosphate (PI3,4,5P3) by PI3K contributes to the activation of signaling pathways that drive cell proliferation.4-7 Removal of the phosphate group from the D5 position of phosphatidylinositols by the SH2–containing inositol 5′-phosphatase (SHIP) is now recognized as an important negative feedback mechanism on cell activation in the mammalian hematopoietic compartment.8,9 SHIP was identified based on its ability to bind Grb2,10 Shc,11,12 and the FcγRIIB receptor13 and by gene trapping.14 In vitro assays show that SHIP can remove the 5′-phosphate of PI3,4,5P3 or inositol 1,3,4,5-tetrakisphosphate,10,11 suggesting that SHIP may counteract the activity of PI3K8,9,15 or prevent the sustained influx of Ca++ into the cell.9,16,17SHIP can also form a complex with Shc in stimulated cells.17-21 Formation of a SHIP-Shc complex is proposed to prevent the recruitment of the nucleotide exchange factor mSos1 to the membrane, thus preventing the activation of the Ras/mitogen-activated protein kinase (MAPK) pathway.21,22 The ability of SHIP to convert PI3,4,5P3 to PI3,4P2 suggests that SHIP might also influence the activation of protein kinase B/Akt and thus influence the induction of programmed cell death.7,23 24
There is increasing genetic evidence that SHIP plays an important role as a negative regulator of cell activation in B-lymphoid cells,1-3,6,16 myeloid cells,1,3 and mast cells.25 These genetic studies and biochemical analysis of SHIP phosphorylation status indicate that SHIP responds to a wide variety of signals in the hematopoietic compartment, including cytokines,18,20,26 antigen,17,19,20 immune complexes,13,16,27,28 the Fc portion of immunoglobulin G (IgG) antibodies,29 and thrombin.30 These results demonstrate that SHIP is an important regulator of cellular responses in the mature cells of several hematopoietic lineages.
Here we describe the cloning and characterization of a novel SHIP isoform, s-SHIP. We show that transcription from an internal site within the SHIP gene promotes the expression of s-SHIP in totipotent embryonic stem (ES) cells and hematopoietic stem cells (HSCs), but not in mature hematopoietic cells. The s-SHIP isoform lacks the SH2 domain found in the previously described SHIP isoform whose expression is restricted to the hematopoietic system. Consistent with this structural difference, s-SHIP does not associate in vivo with the Shc adapter protein, but it does associate with Grb2. We propose that s-SHIP plays a unique signaling role in primitive stem cell populations.
Materials and methods
Cell culture
The ES cell line, TL1, was a kind gift of Dr Patricia A. Labosky31 (University of Pennsylvania, Philadelphia) and was cultured in Dulbecco modified Eagle medium (Gibco-BRL, Grand Island, NY) supplemented with 15% fetal bovine serum (FBS) (Summit Labs, Fort Collins, CO), 0.1 mM MEM nonessential amino acids, 2 mM L-glutamine, 50 μg/mL gentamicin, 50 μM β-mercaptoethanol (β-ΜΕ), and 1000 U/mL leukemia inhibitory factor (LIF) (Gibco-BRL). The ES cell line, E10, was derived from TL1. WEHI-231, 70Z/3, A20, and BAL17 (B-cell lines) were maintained in RPMI 1640 medium (Mediatech, Herndon, VA) with 10% FBS, 2 mM L-glutamine, 50 μM β-ME (Gibco-BRL), and antibiotics. Hepa 1-6 and 293T cell lines were maintained in DMEM medium with the same supplements as described for the RPMI medium.
Reverse transcription–polymerase chain reaction
Cells were washed once with ice-cold phosphate-buffered saline (PBS) and pelleted. Then cells were lysed, and total RNA was prepared using the RNeasy Mini Kit (Qiagen, Valencia, CA) according to the manufacturer's instructions. Reverse transcription (RT) of total cellular RNA and amplification of cDNA products for specific gene sequences was carried out using Ready-to-Go RT-PCR Beads (Amersham Pharmacia, Piscataway, NJ) following the manufacturer's instructions. In this series of RT–polymerase chain reaction (PCR), 1-μg total RNA and 1-μL each gene-specific primer (10 mM) were placed in a 50 μL reaction volume. In general, PCR conditions were as follows: denaturation for 30 seconds at 94°C, annealing for 30 seconds at a temperature optimal for the primer pair (see below), and a 1-minute extension at 72°C. Primer sequences used in RT-PCR reactions were as follows (numbers refer to the murine SHIP cDNA sequence, GenBank accession number U52044): SHIP155 (sense), 5′-CTCCAAGGCAGAGGAGCTAC-3′ (annealing temperature, 62°C); ASHIP675 (antisense), 5′-CCTGGATGGCTTTCAGGTGC-3′ (annealing temperature, 62°C); SHIP1970 (sense), 5′-CGACCAACTGCTCCTGGAGAGGAAGGAC-3′ (annealing temperature, 64°C); ASHIP3164 (antisense), 5′-GTTCTCAAACATCTCGGGCTTCGTCAGC-3′ (annealing temperature, 64°C); ASHIP2640 (antisense), 5′-CATGGTGGGTGAGAGGCGTGTAGATAGG-3′ (annealing temperature, 64°C); and SHIP989 (sense), 5′-CAACAGGCGTTCCCTTATCCCTCCG-3′ (annealing temperature, 64°C). Nested RT-PCR assays to detect mRNAs expressed in sorted cells were carried out as described below.
Cell sorts and nested RT-PCR assays
Femurs and tibias from adult (4- to 6-month-old) male C57BL/6 mice were isolated. Adult bone marrow (ABM) was flushed into chilled RPMI 1640 medium. Fetal livers (FLs) (day 14.5 of gestation) were isolated from pregnant female C57BL/6 mice. Cells were teased from the FLs into chilled RPMI. Cell clusters from the ABM and FL preparations were dispersed into single-cell suspensions with transfer pipettes and were passed over 70 μm nylon cell strainers. Cell suspensions were pelleted, and red blood cells (RBCs) were lysed for 5 minutes at 4°C in RBC lysis buffer (163 mM NH4Cl, 10 mM KHCO3, 0.13 mM EDTA). After RBC lysis, the cells were washed in PBS, passed over 70 μm strainers, counted, and resuspended in staining media (PBS with 3% FBS and 10 mM HEPES) at 107 cells/50 μL. Cells were incubated in Fc Block (Pharmingen, San Diego, CA) at 2 μg/107 cells/50 μL staining media and stained with the appropriate panel of fluorochrome-conjugated anti–murine antibodies at 1 to 2 μg/107 cells/50 μL staining media. Antibodies used were a lineage-fluorescein isothiocyanate panel of Mac-1 (Caltag Laboratories, Burlingame, CA), Gr-1, CD3ε, CD4 (GK1.5), CD8a (53-6.7) B220 (Pharmingen); B220-phycoerythrin (PE), Ter119-PE, c-kit-PE, and biotinylated Sca-1 (Pharmingen). Biotinylated Sca-1 was revealed with streptavidin-allophycocyanin (APC) (Pharmingen). After staining, the cells were washed and resuspended in staining media (50 μL/107 cells) plus propidium iodide (PI) at 1.0 μg/mL for dead cell exclusion. HSCs (Sca-1+c-kit+Lin−), B-lymphoid cells (B220+Gr-1−Mac-1−), myeloid cells (Gr-1+Mac-1+B220−), and erythroid cells (Ter119+Lin−) were sorted from ABM and FL using a FACStarPLUS flow cytometer (Becton Dickinson, San Jose, CA) equipped with an automatic cell deposition unit. ES cells (s-SHIP–positive control), WEHI-231 cells (SHIP-positive control), and Hepa 1-6 cells (negative control) were also prepared in staining media and PI (106 cells/50 μL). Fifty viable cells of each population were deposited into separate wells of a 96-well PCR plate containing lysis buffer (0.4% Nonidet P-40, 60 μM dNTP, 25 μM dithiothreitol, 0.5 U/μL RNasin [Promega, Madison, WI]) and were lysed for 15 minutes on ice (adapted from Hu et al32). RNA was reverse transcribed using multiple primer pairs for β-actin, s-SHIP, and SHIP and 48 U M-MLV reverse transcriptase per reaction in the buffer provided (Gibco-BRL). First-round PCR of 35 cycles was performed with addition of 40 μL buffered RedTaq DNA polymerase (Sigma, St Louis, MO). One-microliter aliquots of the first-round PCR product were further amplified for 35 cycles using fully nested primers for each gene. Aliquots of second-round products were subjected to gel electrophoresis and visualized by ethidium bromide staining. Primers used in the first-round PCRs were as follows: β-actin, 5′-TGGGTCAGAAGGACTCCTATG-3′ and 5′-ACCAGACAGCACTGTGTTGGC-3′; s-SHIP, 5′-GTTCCCACTAGTTGTTGAACT-3′ and 5′-CAGAACCATCCTTGGACTTCTTAA-3′; SHIP, 5′-TGCCTCCCAGAAACATTCCTATGT-3′ and 5′-CAGAACCATCCTTGGACTTCTTAA-3′. Primers for the second-round PCRs were as follows: β-actin, 5′-TGCTGTCCCTGTATGCCTCTG-3′ and 5′-GGAACCGCTCGTTGCCAATAG-3′; s-SHIP, 5′-TTACCTTGAACCTCTGCTCCCAG-3′ and 5′-CGATCAGTTTCCCAGACTCAACGTCCAC-3; SHIP, 5′-TGCAACAGAGAACCCCCGAGCCC-3′ and 5′-CGATCAGTTTCCCAGACTCAACGTCCAC-3′.
cDNA and genomic cloning
To determine the 5′ and 3′ termini of the s-SHIP cDNA, we performed rapid amplification of cDNA ends (RACE) cloning using the Smart Race cDNA Amplification Kit (Clontech, Palo Alto, CA). Primers for 5′ RACE reactions were ASHIP1098 (anti-sense) 5′-CGATCAGTTTCCCAGACTCAACGTCCAC-3′ and ASHIP1998 (anti-sense) 5′-GGTCCTTCCTCTCCAGGAGCAGTTGGTC-3′. Primers for 3′ RACE reactions were SHIP1970 (sense) 5′-CGACCAACTGCTCCTGGAGAGGAAGGAC-3′ and SHIP2766 (sense) 5′-AACCTCACCAGCCATGACCCTATG-3′. Touchdown PCR amplification conditions were used for 5′ and 3′ RACE reactions with the following cycling conditions: 30 seconds at 94°C and 4 minutes at 72°C for cycles 1 to 5; 30 seconds at 94°C, 30 seconds at 70°C, 4 minutes at 72°C for cycles 6 to 10; 30 seconds at 94°C, 30 seconds at 68°C, 4 minutes at 72°C for cycles 11 to 35. Smart Race products were cloned using the TOPO TA cloning kit (Invitrogen, Carlsbad, CA). Sequences derived from the termini of the 5′ and 3′ RACE clones of s-SHIP were used to design primers that amplify the entire s-SHIP cDNA by RT-PCR. Full-length s-SHIP cDNAs were obtained by 2 separate RT-PCR amplifications of overlapping fragments that contained either 5′ or 3′ termini. These 2 fragments were cloned into the pcDNA3 vector (Invitrogen). Genomic sequences were isolated using a mouse GenomeWalker Kit (Clontech). Primers for genomic cloning were as follows: ESHIP23 (sense), 5′-TTACCTTGAACCTCTGCTCCCAG-3′; ASHIP1098 (anti-sense), 5′-CGATCAGTTTCCCAGACTCAACGTCCAC-3′; SHIP741 (sense), 5′-CACCTGAAGAAGCTGATGTCACTGC-3′; AESHIP43 (anti-sense), 5′-TGGGAGCAGAGGTTCAAGGTAAAGTTCAAC-3′. Conditions for touchdown PCR amplification of genomic sequences were as described above. Sequencing of s-SHIP cDNAs and associated genomic clones were carried out at the DNA Sequencing Facility of the University of Pennsylvania Comprehensive Cancer Center.
Sequence analysis
DNA and protein sequences were analyzed with MacVector 7.0 (Oxford Molecular, Madison, WI). Sequence alignments and comparisons were performed with the ClustalW algorithm.33 The Celera Human Genome-Unassembled Fragments database (http://publication.celera.com)34 (Celera Genomics, Rockville, MD) was accessed to perform BlastN queries of the first 120 nucleotides of human 110-kD signaling inositol polyphosphate 5′-phosphatase (SIP-110) cDNA (GenBank accession numberU50040).
Northern blot analysis and ribonuclease protection assay
PolyA+ RNA was prepared as described above. Northern blot analysis of SHIP mRNA expression was as described previously.14 For ribonuclease protection assay (RPA), the target sequence contained the 44-nucleotide SSR of s-SHIP mRNA and nucleotides 45-247 of s-SHIP (nucleotides 785-987 of SHIP). It was amplified using PCR with primers XHOESHIP1 (sense) 5′-CCGCTCGAGGGTTCCCACTAGTTGTTGAAC-3′ and ASHIP987 (anti-sense) 5′-TAGATTCTGAGCCCTCGTGCAGC-3′. This PCR product was TA-cloned into pCR2.1-TOPO (Invitrogen) and then subcloned in a reverse orientation downstream of the T3 promoter of the pRRI-amp-18 vector (Ambion, Austin, TX). The vector was linearized for s-SHIP/SHIP probe synthesis. Anti-sense RNA probe was synthesized and labeled with32P-UTP using the Maxiscript T7/T3 kit (Ambion) following the manufacturer's instructions. The probe for mouse GAPDH had 25 times lower specific activity than the s-SHIP/SHIP probe. The 100 nucleotide RNA markers were generated using templates provided by Ambion. RPA assay was performed on 10 μg total RNA for each sample and 20 000 cpm for each probe using the RPAIII kit (Ambion) according to the manufacturer's instructions. RPA products were visualized by running Quick Point precast gels (6% polyacrylamide denaturing gel; Novex, San Diego, CA) and then by autoradiography.
Transfections
SHIP cDNA was kindly provided by Dr Larry Rohrschneider.11 SHIP and s-SHIP cDNAs were cloned separately into the pcDNA4/HisMaxC expression vector (Invitrogen). These pcDNA4/SHIP and pcDNA4/s-SHIP plasmids were transfected into 293T kidney cells using the FuGENE 6 transfection reagent (Roche Molecular Biochemicals, Indianapolis, IN) according to the manufacturer's instructions.
Antibodies
Purified mouse anti–SHIP monoclonal antibody, P2C6, was a kind gift of Drs David Lucas and Larry Rohrschneider.35 It was used for immunoprecipitation at a concentration of 4 μL/1 mg total protein and for immunoblotting at a dilution of 1:1000. Rabbit anti–Grb2 polyclonal antibody (sc-255; Santa Cruz Biotechnology, Santa Cruz, CA) was used for immunoprecipitation at a concentration of 2 μg/1 mg total protein and for immunoblotting at a dilution of 1:500. Rabbit anti–Shc polyclonal antibody (06-203; Upstate Biotechnology, Lake Placid, NY) was used for immunoblotting at a concentration of 1 μg/mL. Rabbit anti–mSos1 polyclonal antibody (06-246; Upstate Biotechnology) was used for immunoblotting at a concentration of 1 μg/mL. Mouse anti–phosphotyrosine monoclonal antibody (4G10; 05-321; Upstate Biotechnology) was used for immunoblotting at a concentration of 1 μg/mL.
Immunoprecipitations and immunoblots
Approximately 107 cells were washed once with ice-cold PBS and pelleted. Cell pellets were then lysed in 0.5 to 1.0 mL modified RIPA buffer (1% Nonidet P-40, 50 mM Tris-HCl [pH 7.4], 0.25% Na-deoxycholate, 150 mM NaCl, 1 mM EDTA, 1 mM Na3VO4, 1 mM phenylmethylsulfonyl fluoride, 10 μg/mL leupeptin, 10 μg/mL pepstatin, 10 μg/mL aprotinin). Lysates were rocked for 30 minutes at 4°C, and cellular debris was removed by centrifugation at 14 000g for 15 minutes. Total protein concentration of each lysate was determined by spectrophotometry using the BCA Protein Assay Kit (Pierce Chemical, Rockford, IL). Lysates were stored at −80°C. Whole cell lysates were subsequently mixed with NuPAGE 4× lithium dodecyl sulfate (LDS) sample buffer and 10× sample reducing agent, heated for 10 minutes at 70°C, and loaded onto NuPAGE 10% or 4% to 12% Bis-Tris 1.0 mm gels (Invitrogen). For immunoprecipitations, equal concentrations of total protein (either 500 μg in 500 μL or 1 mg in 1 mL lysis buffer) were used for each individual experiment. Samples were precleared with 0.25 μg of the appropriate control IgG (normal mouse or rabbit IgG; Santa Cruz Biotechnology) together with 50 μL appropriate agarose bead conjugate (Protein G-Agarose, Fast Flow [SHIP immunoprecipitation (I.P.)]; Protein A-Agarose, Fast Flow [Grb2 I.P.]; Upstate Biotechnology) for 1 hour at 4°C. Precleared samples were transferred to fresh tubes. Four microliters P2C6 anti–SHIP monoclonal antibody or 2 μg anti–Grb2 polyclonal antibody were added per milligram total protein and were incubated overnight with rocking at 4°C. Fifty microliters appropriate agarose bead conjugate was then added and rocked for 2 hours at 4°C. Agarose bead conjugates were pelleted (14 000g, 5 seconds), washed 3 times with cold PBS, and boiled in NuPAGE 4× LDS sample buffer with 10× sample reducing agent for 5 minutes at 100°C. Beads were pelleted, and equal volumes of the immunoprecipitate supernatant were loaded onto NuPAGE 10% or 4% to 12% Bis-Tris gels. Proteins were separated for 50 minutes at 200 V using MOPS sodium dodecyl sulfate running buffer in an XCell II Mini-Cell unit (Invitrogen). Protein molecular weights were compared using Novex MultiMark protein standards. Separated proteins were transferred to polyvinylidene difluoride (Bio-Rad Laboratories, Hercules, CA) or nitrocellulose [4G10 blots] (Millipore, Bedford, MA) membranes for 75 minutes at 30 V using the XCell II Blot Module. Immunoblotting was carried out using the enhanced chemiluminescence Western blotting analysis system following the manufacturer's instructions (Amersham Pharmacia).
Cell stimulation
ES-TL1 cells were split into 4 different 100-mm TC plates and grown overnight at 37°C in ES media supplemented with 1000 U/mL LIF to approximate 80% confluence. LIF-supplemented media was removed, cells were washed twice with PBS, media without LIF was added, and cells were incubated for 5 hours. Cells were washed once with PBS and twice with Hanks balanced salt solution (HBSS; Mediatech). Then they were pre-incubated in HBSS at 37°C for 10 minutes. Cells were stimulated by the addition of HBSS containing 2000 U/mL LIF and by incubation at 37°C for either 2, 5, or 10 minutes. A control ES plate (0 minute) containing HBSS without LIF was incubated at 37°C in parallel for 10 minutes. Stimulations were stopped by the removal of buffer and the addition of 10 mL ice-cold PBS/1 mM Na3VO4. PBS/Na3VO4 was removed, and 1 mL ice-cold modified RIPA buffer was added immediately. Cells were scraped, and cell lysates were processed as described above. A20 cells were grown in RPMI and were washed once with PBS and twice with HBSS. Then 2 × 107 cells were placed in separate 15-mL conicals and were pre-incubated in HBSS at 37°C for 10 minutes. Cells were spun down, and HBSS was removed. After that, cells were stimulated with 1 mL HBSS containing 20 μg/mL goat anti–mouse IgG (Southern Biotechnology Associates, Birmingham, AL) and were incubated at 37°C for 5 minutes. Control A20 cells (0 minute) containing HBSS without anti–mouse IgG were incubated in parallel. Stimulations were stopped by placing the conicals on ice and adding 2 mL ice-cold PBS/Na3VO4. Cells were pelleted, supernatant was removed, and 1 mL ice-cold modified RIPA buffer was added. Total protein (1.0 mg in 1 mL) of each preparation was used for subsequent immunoprecipitation experiments.
Membrane and cytosol fractionation
Two confluent 150 mm × 25 mm plates of ES-TL1 cells cultured in LIF were washed twice with chilled PBS, then collected with a chilled cell scraper. Cells were pelleted by centrifugation at 300g and resuspended in 2 mL chilled sonication buffer (20 mM Tris-HCl, pH 8, 137 mM NaCl, 10% glycerol, 2 mM EDTA, 1 mM Na3VO4, 1 mM phenylmethylsulfonyl fluoride, 10 μg/mL leupeptin, 10 μg/mL pepstatin, and 10 μg/mL aprotinin). This suspension was sonicated on ice for 20 seconds at 20% full output using a Fisher Model 60 Sonic Dismembrator equipped with a one-eighth–inch diameter tip (Fisher Scientific, Pittsburgh, PA). Trypan blue dye exclusion and examination by phase-contrast microscopy confirmed that no intact cells remained. The suspension was centrifuged at 500g for 5 minutes at 4°C to remove the nuclear fraction. Postnuclear supernatant was centrifuged in Beckman 15 × 31 mm polycarbonate tubes at 100 000g for 20 minutes at 4°C in a Beckman TL-100 centrifuge equipped with a TLA-100.3 rotor. The supernatant cytosol fraction was removed, and Triton X-100 was added to a final concentration of 1%. The membrane pellet was rinsed once with chilled sonication buffer and then resuspended in chilled sonication buffer plus 1% Triton X-100. Resuspension was achieved by needle probe sonication on ice for 30 seconds at 20% full output, and the resuspended membrane fraction was allowed to solubilize at 4°C for 45 minutes with frequent gentle vortexing. Insoluble material was removed by centrifugation at 10 000g for 15 minutes at 4°C. ES-TL1 cells from a third 150 mm plate were collected in parallel with the cells collected for fractionation in order to prepare an unfractionated whole cell lysate for comparison purposes. These cells were lysed in 1 mL of chilled sonication buffer/1% Triton X-100, rocked for 45 minutes at 4°C, and centrifuged at 10 000gfor 15 minutes at 4°C to remove cellular debris. Total protein concentrations of the whole cell lysate, cytosol, and membrane fractions were determined by spectrophotometry using the BCA Protein Assay Kit. Whole cell lysate, cytosol, and membrane fractions were then diluted in sonication buffer/1% Triton X-100 to a final concentration of 1 mg/mL. Fractions were stored at −80°C and 1 mg total protein (in 1 mL) of each preparation was used for subsequent immunoprecipitation experiments.
Results
Identification and characterization of s-SHIP mRNA
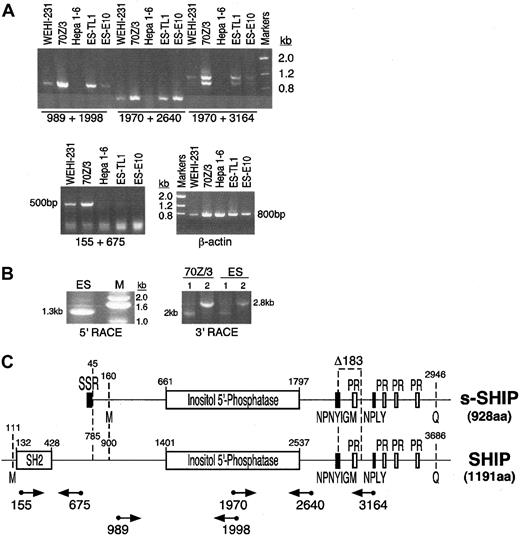
Previous analysis of SHIP expression indicated that its expression is restricted to cells of the hematopoietic system.10,11,14 36 However, while evaluating other cell types for expression of the SHIP gene, we found that ES cells express SHIP mRNA. We detected the presence of SHIP mRNA in polyA+ RNA isolated from ES cells (TL1) using several different RT-PCR assays that amplify distinct regions of SHIP mRNA (Figure 1A,C). However, primers that amplify the 5′ portion of SHIP mRNA that encodes its SH2 domain failed to amplify the SHIP mRNA species expressed in ES cells, but they yielded the expected product when amplifying mRNA from B-lymphocyte cell lines (70Z/3, WEHI-231). Furthermore, we found that an ES cell line (E10) with both SHIP alleles mutated by the insertion of a GFP transgene into exon 1 (J Wang et al, unpublished data, September 1997) retains expression of SHIP mRNA as detected by several independent RT-PCR assays for SHIP expression (Figure 1A,C). These results indicate that ES cells express a SHIP transcript, which we designate s-SHIP, that differs at its 5′ end from the SHIP mRNA previously described in differentiated hematopoietic cells.
ES cells express a SHIP mRNA species different from that expressed in mature hematopoietic cells.
(A) RT-PCR analysis of SHIP expression in ES cells and lineage-committed hematopoietic cell lines. All primers used in the RT-PCR assays were designed to amplify regions of SHIP mRNA encoded by separate exons; hence, the expected amplification products must arise from amplification of cDNA, not from contaminating genomic DNA. The following cell types were analyzed: B-lymphocyte cell lines 70Z/3 and WEHI-231, hepatoma cell line Hepa 1-6 (negative control), and ES cell lines TL1 and E10. Amplification of β-actin was included as a control for the RNA isolation and cDNA synthesis steps. PCR products were resolved on a 1% agarose gel and stained with ethidium bromide. (B) RACE cloning of s-SHIP mRNA. PCR products representing the 5′ or 3′ termini of the s-SHIP mRNA expressed in ES cells were generated by 5′ and 3′ RACE, respectively. For 3′ RACE cloning of SHIP mRNA species in 70Z/3 and ES cells, the primers SHIP2766 (sense; lane 1) or SHIP1970 (sense; lane 2) were used. (C) Schematic depiction of the structure of the s-SHIP and SHIP cDNAs. Nucleotide numbering refers to GenBank accession numbers AF184912 and U52044, respectively. Functional domains and sequence motifs encoded in the s-SHIP and SHIP mRNAs ares-SHIP Region (SSR), Src-homology 2 domain (SH2), proline-rich motifs (PR), and NPXY and YIGM motifs. The internal Δ183 nucleotide deletion is illustrated. Initial methionine (M) and terminal glutamine (Q) amino acids for each mRNA are indicated, yielding proteins of 928 aa (s-SHIP) and 1191 aa (SHIP). Relative location and orientation of the primers used in the RT-PCR analysis or RACE cloning are indicated below the SHIP cDNA. Numbers below the primers (arrows) represent the 5′ nucleotide in the primer based on its position in the SHIP cDNA sequence.
ES cells express a SHIP mRNA species different from that expressed in mature hematopoietic cells.
(A) RT-PCR analysis of SHIP expression in ES cells and lineage-committed hematopoietic cell lines. All primers used in the RT-PCR assays were designed to amplify regions of SHIP mRNA encoded by separate exons; hence, the expected amplification products must arise from amplification of cDNA, not from contaminating genomic DNA. The following cell types were analyzed: B-lymphocyte cell lines 70Z/3 and WEHI-231, hepatoma cell line Hepa 1-6 (negative control), and ES cell lines TL1 and E10. Amplification of β-actin was included as a control for the RNA isolation and cDNA synthesis steps. PCR products were resolved on a 1% agarose gel and stained with ethidium bromide. (B) RACE cloning of s-SHIP mRNA. PCR products representing the 5′ or 3′ termini of the s-SHIP mRNA expressed in ES cells were generated by 5′ and 3′ RACE, respectively. For 3′ RACE cloning of SHIP mRNA species in 70Z/3 and ES cells, the primers SHIP2766 (sense; lane 1) or SHIP1970 (sense; lane 2) were used. (C) Schematic depiction of the structure of the s-SHIP and SHIP cDNAs. Nucleotide numbering refers to GenBank accession numbers AF184912 and U52044, respectively. Functional domains and sequence motifs encoded in the s-SHIP and SHIP mRNAs ares-SHIP Region (SSR), Src-homology 2 domain (SH2), proline-rich motifs (PR), and NPXY and YIGM motifs. The internal Δ183 nucleotide deletion is illustrated. Initial methionine (M) and terminal glutamine (Q) amino acids for each mRNA are indicated, yielding proteins of 928 aa (s-SHIP) and 1191 aa (SHIP). Relative location and orientation of the primers used in the RT-PCR analysis or RACE cloning are indicated below the SHIP cDNA. Numbers below the primers (arrows) represent the 5′ nucleotide in the primer based on its position in the SHIP cDNA sequence.
To confirm the presence of a SHIP mRNA species with a different 5′ end in ES cells and to determine its sequence, we performed 5′ Smart Race cDNA cloning using anti–sense primers complementary to the SHIP cDNA sequence. We used primers from regions of SHIP that our analysis in Figure 1A indicated were also present in s-SHIP mRNA. Two separate 5′ RACE reactions with primers ASHIP1998 and ASHIP1098 yielded a single product of either 1.3 kb (Figure 1B) or 0.4 kb (data not shown), respectively. Sequence analysis of these 5′ RACE products showed that the SHIP mRNA present in ES cells begins with a 44-nucleotide region not previously found in any murine SHIP cDNA identified to date (Figure2). We designate this novel 44-nucleotide region the stem-SHIP region (SSR). The complete sequence of these 5′ RACE products showed that the SSR forms the 5′ end of the s-SHIP mRNA and is fused with the previously identified murine SHIP cDNA sequence beginning at nucleotide 785 and is identical up to nucleotide 1998 (GenBank accession number U52044).
Nucleotide sequence of s-SHIP cDNA and the predicted amino acid sequence of its major ORF.
The first 784 nucleotides of the SHIP cDNA are provided for comparison above the bold arrow. s-SHIP cDNA sequence is indicated below the bold arrow, starting with the numeral 1 and continuing through nucleotide 2946. Remaining s-SHIP cDNA sequence (nucleotides 2947-4125) is not shown because of size constraints, but it is identical to nucleotides 3687 to 4865 of the SHIP cDNA (data not shown). The number in parentheses (785) at nucleotide 45 of the s-SHIP cDNA indicates the nucleotide in the SHIP cDNA, where identity between the s-SHIP and SHIP cDNAs begins. ATG initiator codons for the SHIP and s-SHIP major ORFs are indicated in bold and underlined, and the termination codon shared by both isoforms is indicated by an asterisk. SH2 (dotted box), SSR (solid box), and inositol 5′-phosphatase (dashed box) sequences are shown. The nucleotide sequence that encodes the Δ183 nucleotide deletion is underlined. NPXY and proline-rich motifs are shaded black and gray, respectively.
Nucleotide sequence of s-SHIP cDNA and the predicted amino acid sequence of its major ORF.
The first 784 nucleotides of the SHIP cDNA are provided for comparison above the bold arrow. s-SHIP cDNA sequence is indicated below the bold arrow, starting with the numeral 1 and continuing through nucleotide 2946. Remaining s-SHIP cDNA sequence (nucleotides 2947-4125) is not shown because of size constraints, but it is identical to nucleotides 3687 to 4865 of the SHIP cDNA (data not shown). The number in parentheses (785) at nucleotide 45 of the s-SHIP cDNA indicates the nucleotide in the SHIP cDNA, where identity between the s-SHIP and SHIP cDNAs begins. ATG initiator codons for the SHIP and s-SHIP major ORFs are indicated in bold and underlined, and the termination codon shared by both isoforms is indicated by an asterisk. SH2 (dotted box), SSR (solid box), and inositol 5′-phosphatase (dashed box) sequences are shown. The nucleotide sequence that encodes the Δ183 nucleotide deletion is underlined. NPXY and proline-rich motifs are shaded black and gray, respectively.
To determine the sequence of the remainder of the s-SHIP transcript present in ES cells, we performed 2 separate 3′ RACE reactions (Figure1B) with sense strand specific primers (SHIP1970 and SHIP2766). Cloning and sequencing of these 3′ RACE products indicated that s-SHIP mRNA is identical to SHIP mRNA from nucleotide 1970 through the end of the SHIP cDNA. To further confirm the sequence of the s-SHIP cDNA, we obtained full-length s-SHIP cDNA by RT-PCR. Sequence analysis (Figure 2) of the full-length s-SHIP cDNA confirmed that s-SHIP mRNA lacks the SH2 domain but has the 44-nucleotide SSR fused to nucleotide 785 of SHIP. However, like its hematopoietic counterpart,35,37,38 s-SHIP mRNA encodes an inositol 5′-phosphatase domain and several protein interaction motifs, including 2 NPXY motifs, a YIGM motif, and 4 proline-rich motifs (Figures 1C, 2). As Lucas and Rohrschneider35 found for SHIP mRNA in hematopoietic cells, s-SHIP mRNA is alternatively spliced to generate a form that lacks a 183-nucleotide region (nucleotides 2129-2311 of s-SHIP [GenBank accession number AF184912] corresponding to nucleotides 2869-3051 of SHIP). Forms of s-SHIP mRNA that contain or lack this 183-nucleotide region are present in ES cells, as indicated by the RT-PCR reaction, with primers that span this region (1970 + 3164 in Figure 1A). Sequence analysis of these products confirmed that the alternative splice event that results in the Δ183 s-SHIP mRNA (accession number AF184913) is identical to that described by Lucas and Rohrschneider35 for SHIP (Figure 2).
Genomic location of the s-SHIP first exon indicates its expression arises from an internal promoter in the SHIP gene
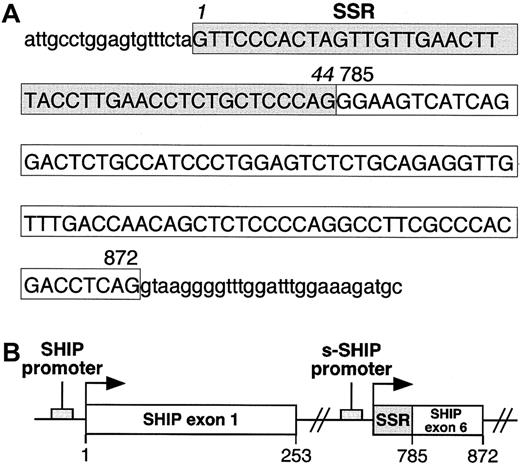
To determine whether the 44-nucleotide SSR at the 5′ end of s-SHIP mRNA represents an independent exon or is part of an existing exon in SHIP, we isolated and sequenced the genomic DNA that spans this region. Sequence analysis of genomic DNA revealed that the 44-nucleotide SSR is directly adjacent to a sequence that serves as exon 6 (nucleotides 785-872) in the SHIP isoform expressed in lineage-committed hematopoietic cells39 (Figure3). Thus, the 44-nucleotide SSR is not a discrete exon but is part of a larger 132-nucleotide region that serves as exon 1 in s-SHIP. Furthermore, because the 132-nucleotide s-SHIP exon 1 is located at an internal site in the SHIP gene, s-SHIP expression must arise from an internal promoter that is 3′ to the SH2-encoding exons and 5′ to the exons that encode the inositol 5′-phosphatase domain (Figure 3B). The location of s-SHIP exon 1 requires that its transcription initiate internally in the SHIP genomic locus in a region that serves as an intron for the SHIP mRNA expressed in differentiated hematopoietic cells.
Organization of the first exon of s-SHIP.
(A) Genomic sequence of the first exon of s-SHIP and adjacent intronic sequence. Uppercase letters represent the nucleotides in the first exon of s-SHIP, and lowercase letters represent the intronic sequence immediately flanking this exon. The 44 nucleotides enclosed in the shaded box indicate the SSR, and the 88 nucleotides in the clear box represent those found in both the s-SHIP and the 145-kd SHIP cDNAs. Numbers 785 and 872 indicate the position of each corresponding nucleotide in the murine 145-kd SHIP cDNA sequence (GenBank accession number U52044). (B) Schematic representation of the orientation of the predicted promoter regions and first exons of SHIP and s-SHIP relative to each other in the SHIP locus. Arrows indicate the transcriptional start points. The s-SHIP first exon consists of the 44-nucleotide SSR and the adjacent 88-nucleotide SHIP exon 6.39 Nucleotide numbering refers to the 145-kd SHIP cDNA sequence (accession number U52044).
Organization of the first exon of s-SHIP.
(A) Genomic sequence of the first exon of s-SHIP and adjacent intronic sequence. Uppercase letters represent the nucleotides in the first exon of s-SHIP, and lowercase letters represent the intronic sequence immediately flanking this exon. The 44 nucleotides enclosed in the shaded box indicate the SSR, and the 88 nucleotides in the clear box represent those found in both the s-SHIP and the 145-kd SHIP cDNAs. Numbers 785 and 872 indicate the position of each corresponding nucleotide in the murine 145-kd SHIP cDNA sequence (GenBank accession number U52044). (B) Schematic representation of the orientation of the predicted promoter regions and first exons of SHIP and s-SHIP relative to each other in the SHIP locus. Arrows indicate the transcriptional start points. The s-SHIP first exon consists of the 44-nucleotide SSR and the adjacent 88-nucleotide SHIP exon 6.39 Nucleotide numbering refers to the 145-kd SHIP cDNA sequence (accession number U52044).
Embryonic and hematopoietic stem cells express s-SHIP mRNA
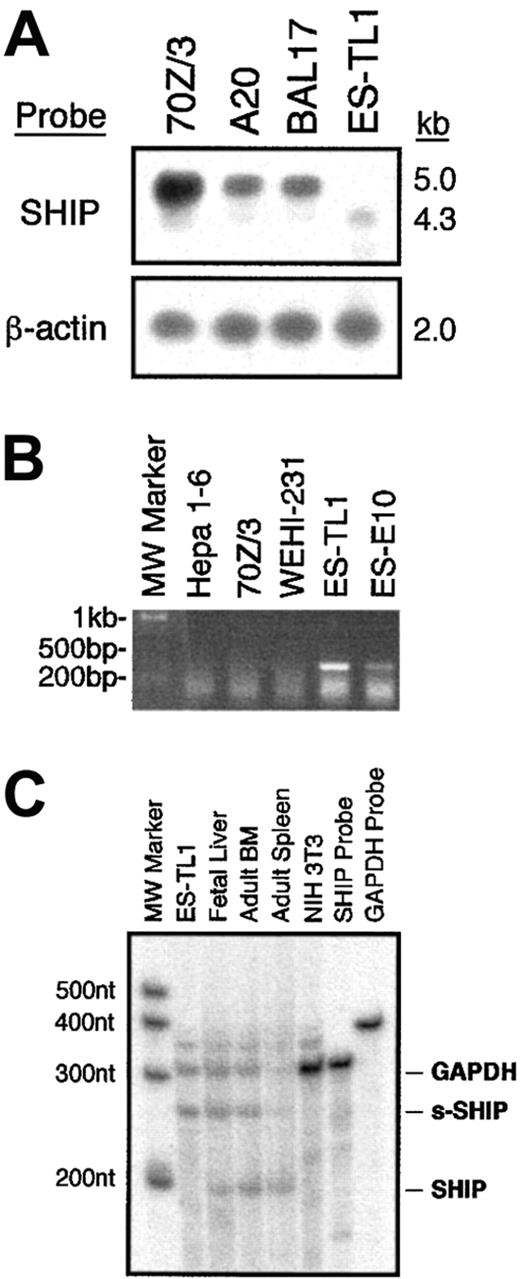
Our RT-PCR analysis indicated that s-SHIP is the only SHIP transcript expressed in ES cells (Figure 1). However, to determine whether s-SHIP is also expressed in the hematopoietic system along with its SH2-encoding counterpart, we analyzed total polyA+ RNA isolated from ES cells and lineage-committed hematopoietic cell lines by Northern blot. This analysis showed that B-lymphocyte cell lines (70Z/3, A20, BAL17) express only the approximately 5 kb SHIP mRNA, whereas the ES cell line TL1 expresses only the smaller, approximately 4.3 kb s-SHIP mRNA (Figure4A). In addition, an RT-PCR assay that is specific for s-SHIP expression showed that ES cells (TL1, E10) express s-SHIP, whereas B-lymphocyte cell lines (70Z/3, WEHI-231) do not (Figure 4B). These results demonstrate that s-SHIP expression is silenced in differentiated hematopoietic cells.
Analysis of s-SHIP and SHIP mRNA expression in ES cells and hematopoiesis.
(A) Northern Blot analysis of SHIP mRNA species expressed by ES cells (TL1) and lineage-committed hematopoietic cell lines (70Z/3, A20, BAL17). A full-length SHIP cDNA clone was used to probe the filter. After initial hybridization with the SHIP probe, the filter was stripped and reprobed for β-actin to confirm that comparable amounts of RNA were loaded in each lane. (B) RT-PCR analysis of s-SHIP expression in ES cells (TL1, E10) and lineage-committed hematopoietic cell lines (70Z/3, WEHI-231). The primer pair ESHIP23 and ASHIP1098 selectively amplifies s-SHIP mRNA and not SHIP mRNA. Analysis of the hepatoma cell line, Hepa 1-6, serves as a negative control for SHIP. (C) RPA analysis of s-SHIP and SHIP expression in ES cells and hematopoietic development. After annealing and RNase digestion, the s-SHIP/SHIP probe is protected for 247 nucleotides by s-SHIP mRNA and for 203 nucleotides by SHIP mRNA. Thus, a single probe is used to detect the relative abundance of the s-SHIP and SHIP mRNAs. Each RNA sample was incubated with a GAPDH probe as an internal control. Analysis of NIH3T3 cell RNA is provided as a negative control for both SHIP and s-SHIP mRNA expression.
Analysis of s-SHIP and SHIP mRNA expression in ES cells and hematopoiesis.
(A) Northern Blot analysis of SHIP mRNA species expressed by ES cells (TL1) and lineage-committed hematopoietic cell lines (70Z/3, A20, BAL17). A full-length SHIP cDNA clone was used to probe the filter. After initial hybridization with the SHIP probe, the filter was stripped and reprobed for β-actin to confirm that comparable amounts of RNA were loaded in each lane. (B) RT-PCR analysis of s-SHIP expression in ES cells (TL1, E10) and lineage-committed hematopoietic cell lines (70Z/3, WEHI-231). The primer pair ESHIP23 and ASHIP1098 selectively amplifies s-SHIP mRNA and not SHIP mRNA. Analysis of the hepatoma cell line, Hepa 1-6, serves as a negative control for SHIP. (C) RPA analysis of s-SHIP and SHIP expression in ES cells and hematopoietic development. After annealing and RNase digestion, the s-SHIP/SHIP probe is protected for 247 nucleotides by s-SHIP mRNA and for 203 nucleotides by SHIP mRNA. Thus, a single probe is used to detect the relative abundance of the s-SHIP and SHIP mRNAs. Each RNA sample was incubated with a GAPDH probe as an internal control. Analysis of NIH3T3 cell RNA is provided as a negative control for both SHIP and s-SHIP mRNA expression.
To further assess the relative ratio of s-SHIP to SHIP mRNA expressed in different cell types and tissues, we developed an RPA that simultaneously detects both s-SHIP and SHIP mRNAs (Figure 4C). RPA analysis indicates that s-SHIP is expressed to the exclusion of SHIP in ES cells and that SHIP mRNA is the predominant form in adult spleen. However, in FL and ABM—tissues that contain significant numbers of pluripotent HSC—we find coexpression of s-SHIP and SHIP mRNA. Interestingly, FL has significantly more s-SHIP than SHIP mRNA, whereas ABM has slightly less s-SHIP than SHIP mRNA. These results suggest that the relative abundance of s-SHIP mRNA in a tissue may correlate to the relative abundance of HSC activity in the tissue. For instance, FL is substantially enriched for primitive HSC activity relative to ABM,40 whereas the spleen is a poor source of HSC activity.
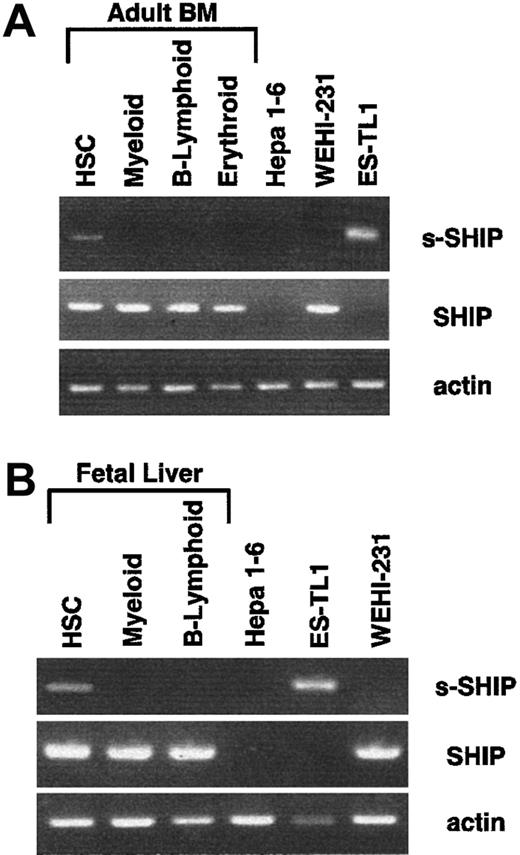
To directly assess the possibility that s-SHIP is expressed in HSC, we developed a nested RT-PCR procedure to detect the expression of s-SHIP and SHIP mRNAs. To define the s-SHIP expression pattern in vivo, we sorted HSC (Sca-1+c-Kit+Lin−) as well as cells of the B-lymphoid (B220+), myeloid (Gr-1+Mac-1+), and erythroid (Ter119+) lineages from day 14.5 FL or ABM. Fifty cells of each population were deposited into single wells of a 96-well PCR plate containing lysis buffer and dNTPs.32 RNA was reverse transcribed by adding multiple primer pairs for s-SHIP, SHIP, and β-actin together with M-MLV reverse transcriptase. Nested RT-PCR reactions were performed (see “Materials and methods”). Representative RT-PCR results from multiple independent sorts of ABM and FL are shown in Figure 5. We detected s-SHIP mRNA in Sca-1+c-kit+Lin− HSC from both ABM and FL, but not in cells of the B-lymphoid, myeloid, or erythroid lineages. RT-PCR analysis of 50 sorted ES-TL1, WEHI-231, and Hepa 1-6 cells served as s-SHIP, SHIP, and negative controls. These results provide further support for our hypothesis that s-SHIP expression is restricted to primitive stem cell populations. We also detected SHIP expression in HSC; thus, it remains to be determined whether the 2 isoforms are coexpressed in all HSC or whether they are discordantly expressed in distinct subsets of HSC.
HSCs express s-SHIP mRNA.
HSCs and lineage-committed cells were sorted from ABM or day 14.5 FL and analyzed by nested RT-PCR assay for the expression of s-SHIP, SHIP, and β-actin mRNA. In both ABM and FL, s-SHIP mRNA was detected in HSCs (Sca-1+c-kit+Lin−) but not in the myeloid (Gr-1+/Mac-1+), B-lymphoid (B220+), or erythroid (Ter119+) lineages. Hepa 1-6 cells were sorted as the negative control for both s-SHIP and SHIP mRNA expression, and ES-TL1 and WEHI-231 cells were sorted as positive controls for s-SHIP and SHIP mRNA expression, respectively.
HSCs express s-SHIP mRNA.
HSCs and lineage-committed cells were sorted from ABM or day 14.5 FL and analyzed by nested RT-PCR assay for the expression of s-SHIP, SHIP, and β-actin mRNA. In both ABM and FL, s-SHIP mRNA was detected in HSCs (Sca-1+c-kit+Lin−) but not in the myeloid (Gr-1+/Mac-1+), B-lymphoid (B220+), or erythroid (Ter119+) lineages. Hepa 1-6 cells were sorted as the negative control for both s-SHIP and SHIP mRNA expression, and ES-TL1 and WEHI-231 cells were sorted as positive controls for s-SHIP and SHIP mRNA expression, respectively.
Embryonic stem cells express a lower molecular weight isoform of SHIP, consistent with the predicted open-reading frame in the s-SHIP cDNA
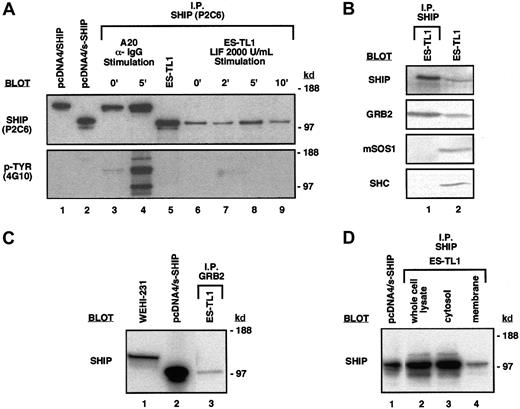
The major predicted open-reading frame (ORF) in s-SHIP mRNA begins with a Kozak41 consensus ATG that is 3′ to the 44-nucleotide SSR (Figure 2). This ATG and its associated ORF of 928 amino acids (aa) is predicted to yield a translated product of 104 kd. Immunoprecipitation and blotting with the anti–SHIP monoclonal antibody P2C6, a kind gift of Drs David Lucas and Larry Rohrschneider,35 revealed a SHIP protein in ES cell lysates with an apparent molecular weight (MW) of approximately 104 kd, consistent with the predicted MW (Figure6A). The P2C6 antibody reacts with a region of SHIP from aa-866 to aa-1020 that, based on our cDNA sequence, should also be present in the s-SHIP isoform. This smaller s-SHIP isoform is not detected in the B-lymphoid cell lines A20 and WEHI-231 (Figure 6A-C). In addition, the endogenous s-SHIP protein immunoprecipitated from ES cells comigrated with the translated product of the s-SHIP cDNA when expressed after transfection of a non-ES cell line, 293T (Figure 6A). Thus, this approximately 104-kd species represents the s-SHIP protein translated from the s-SHIP mRNA in ES cells, and not the COOH-terminal truncation product of SHIP with a similar MW identified by Damen et al.42 Furthermore, we found that s-SHIP appeared as a doublet of approximately 104/97 kd in ES cells and in transfected 293T cells (Figure 6A-D). This doublet presumably represents the translation products of 2 different s-SHIP mRNAs (928-aa and 867-aa [predicted MW, 97.7 kd], respectively) generated by the alternative 183 nucleotide splice reaction identified in hematopoietic cells35 that also takes place in ES cells (Figure 1A).
ES cells express the s-SHIP protein isoform that associates with the Grb2 adapter protein.
(A) Immunoprecipitation and immunoblot detection of s-SHIP in ES cell lysates. Lysate from ES-TL1 cells cultured in LIF was immunoprecipitated with the P2C6 anti–SHIP monoclonal antibody, separated on gels, transferred to membranes, and probed with P2C6, revealing 104-kd and 97-kd proteins (lane 5). No tyrosine phosphorylation of these proteins was detected when they were probed with the 4G10 anti–phosphotyrosine antibody. For comparison, lysates from 293T cells transfected with SHIP cDNA (lane 1) and s-SHIP cDNA (lane 2) were included in the blots. To further assess the tyrosine phosphorylation status of s-SHIP, timed LIF stimulation studies were performed. ES-TL1 cells incubated for 5 hours without LIF were stimulated with 2000 U/mL LIF for 0, 2, 5, or 10 minutes and were rapidly lysed. Equal amounts of total protein were immunoprecipitated with the P2C6 antibody and probed separately with the P2C6 and 4G10 antibodies (lanes 6-9). No tyrosine phosphorylation of s-SHIP was detected at any time point with the 4G10 antibody. For comparison, A20 B-lymphoid cells were stimulated with anti–IgG antibody for 0 or 5 minutes, lysed, immunoprecipitated with P2C6, and probed with P2C6 and 4G10 (lanes 3, 4), showing prominent tyrosine phosphorylation of SHIP at 5 minutes. Molecular mass standards are indicated on the right. (B) s-SHIP associates with Grb2 but not Shc in ES cells. ES-TL1 cell lysates were prepared and immunoprecipitated with the P2C6 anti–SHIP monoclonal antibody. Resolved immunoprecipitates were then blotted with antibodies specific for SHIP, Grb2, mSos1, or Shc (lane 1). Whole cell lysates from ES-TL1 cells were also included to confirm the expression of these proteins in ES cells (lane 2). (C) Grb2 associates with s-SHIP in ES cells. ES-TL1 cell lysate was prepared and immunoprecipitated with an anti–Grb2 polyclonal antibody. Resolved immunoprecipitate was then blotted with the P2C6 anti–SHIP monoclonal antibody (lane 3). For comparison, whole cell lysates from WEHI-231 cells (lane 1) and 293T cells transfected with s-SHIP cDNA (lane 2) were included. (D) Subcellular localization of s-SHIP protein in ES cells. Whole cell lysate, cytosol, and membrane fractions from ES-TL1 cells were prepared as described. One milligram total protein from each fraction was immunoprecipitated with the P2C6 anti–SHIP monoclonal antibody, and equal volumes of immunoprecipitate from each preparation were separated on gels, transferred, and blotted with the P2C6 anti–SHIP monoclonal antibody (lanes 2-4). For comparison, whole cell lysate from 293T cells transfected with s-SHIP cDNA was included (lane 1).
ES cells express the s-SHIP protein isoform that associates with the Grb2 adapter protein.
(A) Immunoprecipitation and immunoblot detection of s-SHIP in ES cell lysates. Lysate from ES-TL1 cells cultured in LIF was immunoprecipitated with the P2C6 anti–SHIP monoclonal antibody, separated on gels, transferred to membranes, and probed with P2C6, revealing 104-kd and 97-kd proteins (lane 5). No tyrosine phosphorylation of these proteins was detected when they were probed with the 4G10 anti–phosphotyrosine antibody. For comparison, lysates from 293T cells transfected with SHIP cDNA (lane 1) and s-SHIP cDNA (lane 2) were included in the blots. To further assess the tyrosine phosphorylation status of s-SHIP, timed LIF stimulation studies were performed. ES-TL1 cells incubated for 5 hours without LIF were stimulated with 2000 U/mL LIF for 0, 2, 5, or 10 minutes and were rapidly lysed. Equal amounts of total protein were immunoprecipitated with the P2C6 antibody and probed separately with the P2C6 and 4G10 antibodies (lanes 6-9). No tyrosine phosphorylation of s-SHIP was detected at any time point with the 4G10 antibody. For comparison, A20 B-lymphoid cells were stimulated with anti–IgG antibody for 0 or 5 minutes, lysed, immunoprecipitated with P2C6, and probed with P2C6 and 4G10 (lanes 3, 4), showing prominent tyrosine phosphorylation of SHIP at 5 minutes. Molecular mass standards are indicated on the right. (B) s-SHIP associates with Grb2 but not Shc in ES cells. ES-TL1 cell lysates were prepared and immunoprecipitated with the P2C6 anti–SHIP monoclonal antibody. Resolved immunoprecipitates were then blotted with antibodies specific for SHIP, Grb2, mSos1, or Shc (lane 1). Whole cell lysates from ES-TL1 cells were also included to confirm the expression of these proteins in ES cells (lane 2). (C) Grb2 associates with s-SHIP in ES cells. ES-TL1 cell lysate was prepared and immunoprecipitated with an anti–Grb2 polyclonal antibody. Resolved immunoprecipitate was then blotted with the P2C6 anti–SHIP monoclonal antibody (lane 3). For comparison, whole cell lysates from WEHI-231 cells (lane 1) and 293T cells transfected with s-SHIP cDNA (lane 2) were included. (D) Subcellular localization of s-SHIP protein in ES cells. Whole cell lysate, cytosol, and membrane fractions from ES-TL1 cells were prepared as described. One milligram total protein from each fraction was immunoprecipitated with the P2C6 anti–SHIP monoclonal antibody, and equal volumes of immunoprecipitate from each preparation were separated on gels, transferred, and blotted with the P2C6 anti–SHIP monoclonal antibody (lanes 2-4). For comparison, whole cell lysate from 293T cells transfected with s-SHIP cDNA was included (lane 1).
s-SHIP forms a complex with Grb2 and is present constitutively at the membrane in embryonic stem cells
The SHIP isoform expressed in mature hematopoietic cells is tyrosine phosphorylated on stimulation by growth factor,18,20,26 immune complexes,13,20,27,28or BCR engagement.17,19,20 Phosphorylated SHIP is found associated with the adapter protein Shc, which facilitates SHIP recruitment to the plasma membrane,19 where it can then act on phosphatidylinositol substrates such as PI3,4,5P3. However, we found that s-SHIP is not tyrosine phosphorylated in either ES cells grown in LIF at steady state or ES cells deprived of LIF for 5 hours and subsequently stimulated with 2× LIF (Figure 6A). Furthermore, s-SHIP in ES cells grown in LIF did not associate with Shc (Figure 6B). Instead, immunoprecipitation of s-SHIP by the P2C6 anti–SHIP antibody coimmunoprecipitated the adapter protein Grb2. Similarly, immunoprecipitation with a polyclonal antibody specific for Grb2 coimmunoprecipitated s-SHIP (Figure 6C). These results demonstrate an in vivo association between s-SHIP and the major adapter protein Grb2 in totipotent stem cells that does not require tyrosine phosphorylation of s-SHIP. s-SHIP does not appear to be associated with either Shc or mSos1 in ES cells, suggesting that it may have different preferences for partnering than the SHIP isoform (Figure6B). Immunoblotting of ES cell lysates confirmed that Shc and mSos1 are expressed in ES cells. To further assess the potential for s-SHIP to play a signaling role in primitive stem cells, we prepared cytosolic and membrane fractions of ES cells. Immunoprecipitation of s-SHIP from these subcellular fractions (Figure 6D) demonstrates the presence of s-SHIP protein in the membrane fraction of ES cells. This membrane localization of s-SHIP and its association with a major adapter protein indicate a role for s-SHIP in signaling pathways active in primitive stem cells.
s-SHIP demonstrates significant homology with the human 110-kd signaling inositol polyphosphate 5′-phosphatase
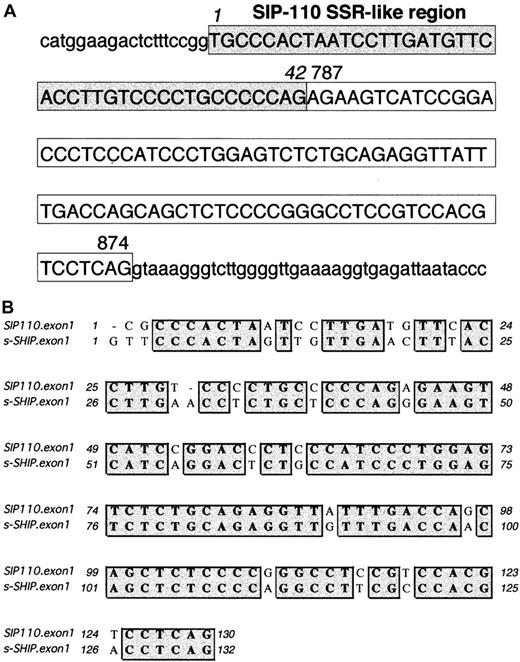
Clustal W Sequence Analysis33 revealed that s-SHIP and the human SHIP isoform SIP-110 (GenBank accession number U50040), described by Kavanaugh et al,12 share 78% nucleotide identity. Of note, Kavanaugh et al12 predicted a 976-aa protein (calculated MW, 109 kd) based on an ATG initiation codon at position 19 of GenBank accession number U50040. This ATG initiation site lacks strong Kozak41 consensus features. However, a second ATG initiation codon at position 160 displays a strong Kozak consensus sequence of CAnCATGG and would predict a translated protein of 929 aa (calculated MW, 104 kd). This shorter ORF of SIP-110 shares 88% amino acid identity with the s-SHIP ORF. Nucleotides 43-2786 of human SIP-110 cDNA are identical to nucleotides 787-3537 of human 145-kd SHIP cDNA (GenBank accession number U50041). Reminiscent of s-SHIP, nucleotides 1-42 of SIP-110 are not found in the 145-kd SHIP cDNA. Kavanaugh et al12 proposed that SIP-110 is a splice variant of the 145-kd SHIP. However, we hypothesized that SIP-110 may represent the human homologue of s-SHIP and may arise by alternative promoter use. To confirm this, we conducted a BlastN query of the first 120 nucleotides of SIP-110 in the Celera Human Genome-Unassembled Fragments database (http://publication.celera.com).34 We found 2 unassembled and uncharacterized genomic fragments (Celera accession numbers GA_x4N24A5J0RM:1..262 and GA_x4N24A6F9UD:1..508) that span the first exon of SIP-110. These genomic fragments revealed that the first 42 nucleotides of SIP-110 are found in the SHIP intronic sequence and are followed directly by an 88-nucleotide sequence representing an internal SHIP exon corresponding to nucleotides 787-874 of 145-kd SHIP (Figure 7A). Thus, analogous to s-SHIP, the first exon of SIP-110 is comprised of a 42-nucleotide SSR-like region unique to SIP-110 and an adjacent 88-nucleotide exon shared by both SIP-110 and 145-kd SHIP. The first exons of SIP-110 and s-SHIP show 82% nucleotide identity (Figure 7B). These results establish that SIP-110 is the human homologue of s-SHIP.
Organization of the first exon of SIP-110, the human homolog of s-SHIP.
(A) Genomic sequence of the SIP-110 first exon and adjacent intronic sequence as compiled from the Celera Human Genome-Unassembled Fragments database (Celera accession numbers GA_x4N24A5J0RM:1..262 and GA_x4N24A6F9UD:1..508). Uppercase letters represent the nucleotides in the SIP-110 first exon, and lowercase letters represent the intronic sequence immediately flanking this exon. The 42 nucleotides enclosed in the shaded box indicate the SSR-like region, and the 88 nucleotides in the clear box represent those nucleotides found in both the SIP-110 and the human 145-kd SHIP cDNAs. Based on the genomic fragment sequences, position 1 is listed as a “T” rather than as the “C” assigned in the SIP-110 cDNA (GenBank accession number U50040). Numbers 787 and 874 indicate the position of the corresponding nucleotide in the human 145-kd SHIP cDNA sequence (GenBank accession number U50041). (B) Clustal W alignment33 of the first exon sequences of human SIP-110 and murine s-SHIP. Matching nucleotides are shaded gray. Overall, the first exons of SIP-110 and s-SHIP show 82% nucleotide identity.
Organization of the first exon of SIP-110, the human homolog of s-SHIP.
(A) Genomic sequence of the SIP-110 first exon and adjacent intronic sequence as compiled from the Celera Human Genome-Unassembled Fragments database (Celera accession numbers GA_x4N24A5J0RM:1..262 and GA_x4N24A6F9UD:1..508). Uppercase letters represent the nucleotides in the SIP-110 first exon, and lowercase letters represent the intronic sequence immediately flanking this exon. The 42 nucleotides enclosed in the shaded box indicate the SSR-like region, and the 88 nucleotides in the clear box represent those nucleotides found in both the SIP-110 and the human 145-kd SHIP cDNAs. Based on the genomic fragment sequences, position 1 is listed as a “T” rather than as the “C” assigned in the SIP-110 cDNA (GenBank accession number U50040). Numbers 787 and 874 indicate the position of the corresponding nucleotide in the human 145-kd SHIP cDNA sequence (GenBank accession number U50041). (B) Clustal W alignment33 of the first exon sequences of human SIP-110 and murine s-SHIP. Matching nucleotides are shaded gray. Overall, the first exons of SIP-110 and s-SHIP show 82% nucleotide identity.
Discussion
The SHIP protein is a key signaling component that participates in controlling the responses of several different hematopoietic cell types in the adult mouse.1-3,13,16 25 Our results provide the first evidence that a SHIP isoform plays a specific role in the biologic composition of stem cell populations. Four lines of evidence support this hypothesis: (1) Totipotent ES cells express the s-SHIP isoform exclusively, (2) s-SHIP partners preferentially with Grb2 in ES cells, (3) s-SHIP is present constitutively in the membrane fraction of ES cells, and (4) s-SHIP expression within the hematopoietic compartment is restricted to fetal and adult hematopoietic stem cells. Our results also suggest the evolution of a transcription control mechanism that promotes the expression of s-SHIP in stem cell populations.
The striking discordance between stem cells and mature hematopoietic cells in the expression of s-SHIP mRNA suggests a distinct signaling role for this SHIP isoform in stem cells. The SH2 domain of SHIP allows it to participate in signal transduction pathways in mature hematopoietic cells that involve partnering with Shc18-20,26,43 or binding to ITIM motifs of receptors.13 The lack of an SH2 domain in s-SHIP may change its binding preferences and, therefore, the pathways with which it interacts. Consistent with this hypothesis, we detected no association of s-SHIP with Shc in ES cells at steady state or after LIF stimulation. We did find an association of s-SHIP with Grb2 in ES cells in the absence of tyrosine phosphorylation. Several groups have shown that SHIP can associate with Grb2 in stimulated hematopoietic cells through an interaction of the SH3 domain of Grb2 and the proline-rich motifs of SHIP.10,12,17-19,44 s-SHIP contains these same proline-rich motifs. In addition to SH3 domains, Grb2 contains an SH2 domain that enables its binding to other signal transduction components or receptors at the cell membrane.45 46 Grb2, through its SH2 domain, could recruit s-SHIP to tyrosine-phosphorylated receptors that are important in the biologic makeup of ES cells and HSCs. Thus, the absence of an SH2 domain may not preclude s-SHIP from acting at the cell membrane, but it may alter how it is recruited to the cell membrane and which signaling pathways it impacts.
Membrane recruitment of s-SHIP would enable its inositol 5′-phosphatase domain to access phosphatidylinositols such as PI3,4,5P3 and to influence proliferation, differentiation, or apoptosis of stem cell populations. Because the association of s-SHIP with Grb2 does not appear to require tyrosine phosphorylation, this complex is likely to be present in quiescent stem cell populations. This preformed complex could rapidly associate with receptors that become tyrosine phosphorylated after basal level growth factor stimulation, enabling s-SHIP to prevent the accumulation of PI3,4,5P3 to significant levels. In this way, s-SHIP could participate in the process of maintaining a stem cell population in a quiescent state. Furthermore, analogous to the proposal that SHIP interferes with the mSos1/Shc/Grb2 complex of the MAPK pathway in mature cells,47 one can envision a competition in stem cells between s-SHIP and mSos1 for Grb2. The outcome of this competition may influence the decision to remain quiescent or to activate the Ras/MAPK pathway that triggers proliferation and differentiation.
The identification of a distinct SHIP isoform expressed from an internal site within the SHIP gene suggests the possibility that the recently reported SHIP-null mice may not be absolute nulls for expression from the SHIP locus.1-3,25,48-50 The mutation strategy used by both groups involved the insertional mutation of the first exon that encodes a portion of the SH2 domain at the amino terminus of the SHIP isoform. This insertional mutation strategy clearly leads to the ablation of the SHIP isoform in mature hematopoietic cells. However, one would not predict that expression of the s-SHIP isoform in the embryo or in HSCs would be affected given that the s-SHIP first exon consists of the terminal 44 nucleotides of intron V, together with exon 6 of SHIP.39 This region is approximately 27 kb downstream of the 145-kd SHIP promoter. Nevertheless, ablation of SHIP expression by this strategy may lead to the dysregulation of s-SHIP expression by an unknown mechanism. In preliminary experiments, we were unable to detect s-SHIP protein expression from total bone marrow of adult C57BL/6 mice in which the first exon of SHIP was insertionally mutated (data not shown). If it can be determined that s-SHIP expression during embryogenesis and hematopoietic development is intact in these SHIP-null strains, one might predict a novel phenotype in mice designed to be null for both s-SHIP and SHIP.
Kavanaugh et al12 cloned and characterized the human SHIP isoform, SIP-110, that lacks an SH2 domain. They cloned SIP-110 cDNA (GenBank accession number U50040) from a human placenta cDNA expression library based on its affinity for a Grb2 SH3 domain in the absence of tyrosine phosphorylation. The overall structure of SIP-110 was similar to s-SHIP. However, SIP-110 was described as a 976-aa protein with an estimated molecular weight of 110 kd. Although the origin of the first 42 nucleotides of SIP-110 within the human SHIP gene was unknown, it was proposed that SIP-110 results from an alternative splice event.12,38,39 Our analysis of the structure and the genomic location of the first exon of SIP-110 confirms that SIP-110 is the human homologue of s-SHIP. Furthermore, the structure and internal location of the SIP-110 first exon within the SHIPgene indicates that SIP-110 arises through alternative promoter use rather than as a splice variant, as was previously proposed. Although there are 2 potential ORFs for SIP-110 that predict proteins of 976 aa and 929 aa, the distal ATG initiation codon has Kozak consensus features that may favor translation of the 929-aa, 104-kd protein, which more closely resembles s-SHIP.51-53 Our results demonstrate that s-SHIP is expressed in murine HSCs. Because cord blood within placenta is known to be enriched for HSCs,54-56 we hypothesize that the SIP-110 cDNA isolated by Kavanaugh et al12 may have originated from cord blood HSC cDNA present in the placenta library.
In vitro studies with recombinant SIP-110 confirmed that it can bind Grb2, but its inositol 5′-phosphatase activity is unaffected by Grb2 binding.12 44 Furthermore, SIP-110 did not associate with Shc. These results can be extrapolated to s-SHIP, and they suggest that the purpose of Grb2 binding is not to alter s-SHIP enzymatic activity but rather to direct the subcellular localization of this activity.
Geier et al57 similarly found a 100-kd isoform of SHIP in both human and murine bone marrow using a polyclonal antibody to an amino acid sequence of murine SHIP (aa 670 to aa 868) that is also present in s-SHIP. Interestingly, they also found that a 100-kd isoform was the most prominent species in human peripheral blood mononuclear cells. Examination of the human ML-1 myeloblastic leukemia cell line revealed the presence of a 100-kd SHIP isoform and a 105-kd isoform and the absence of higher MW isoforms. When these ML-1 myeloblasts were induced to differentiate into monocytes in the presence of 12-O-tetradecanoylphorbol-13-acetate, higher MW isoforms (175 kd, 145 kd, and 130 kd) appeared, whereas the lower MW isoforms (105 kd, 100 kd) were no longer detectable. These results with ML-1 cells indicate that as human myeloid cells differentiate, human SHIP expression changes from the lower MW 105-kd and 100-kd isoforms to the higher MW isoforms. This is analogous to what we found with s-SHIP and SHIP isoforms in murine hematopoietic cell development, and it suggests that the lower MW human isoforms characterized by Geier et al57 represent the SIP-110 identified by Kavanaugh et al.12 However, it is unclear why in their studies human peripheral blood displayed only the lower MW 100-kd isoform. Wolf et al39 describe an additional 110-kd SHIP isoform, SHIPδ, which is the product of an out-of-frame splice event with a deletion of 167 nucleotides in the C-terminal region. This clearly represents a different isoform than s-SHIP. It remains to be determined which SHIP-related proteins, among these several isoforms encoding 100 kd to 110 kd, predominate during different stages of hematopoietic development.
Recently, Krause et al58 elegantly demonstrated that a single murine adult bone marrow stem cell can repopulate hematopoietic cells and a diverse array of nonhematopoietic epithelial tissues, including lung, liver, gastrointestinal tract, and skin. Given that s-SHIP is expressed in totipotent ES cells and at least a subset of bone marrow HSCs, this study raises the possibility that s-SHIP expression may serve to further characterize and delineate the pluripotent bone marrow stem cell capable of multi-organ engraftment that Krause et al58 identified.
We thank Drs David Lucas and Larry Rohrschneider for providing the SHIP cDNA and the generous quantities of P2C6 anti–SHIP monoclonal antibody. We also thank Dr Patricia A. Labosky for providing the TL1 ES cell line and Dr Diane Krause for assisting us with the methodology for the cell sort–nested RT-PCR reactions. We thank Hank Pletcher, Andrew Morschauser, Nikki Brake, and Dr Jonni Moore of the University of Pennsylvania Cancer Center Flow Cytometry Facility for their excellent technical assistance with the cell sorts.
Supported by National Institutes of Health grant DK 54767; the Penn Research Foundation; the H. Lee Moffitt Cancer Center and Research Institute academic development funds; the National Heart, Lung and Blood Institute training grant T32 HL07775 (J.M.N.); and the National Cancer Institute training grant T32 CA09140 (J.M.N.).
Z.T. and J.M.N. contributed equally to this work.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 U.S.C. section 1734.
References
Author notes
William G. Kerr, Immunology Program, H. Lee Moffitt Cancer Center and Research Institute, 12902 Magnolia Dr, Tampa, FL 33612; e-mail: kerrw@moffitt.usf.edu.


This feature is available to Subscribers Only
Sign In or Create an Account Close Modal