Antigens associated with persistent viral infections can mimic epitopes presented on hematopoietic progenitor cells.
Respective epitopes can drive aplastic anemia by T cell–mediated elimination of hematopoietic progenitor cells.
Visual Abstract
Acquired aplastic anemia is a bone marrow failure syndrome characterized by hypocellular bone marrow and peripheral blood pancytopenia. Frequent clinical responses to calcineurin inhibition and antithymocyte globulin strongly suggest critical roles for hematopoietic stem/progenitor cell–reactive T-cell clones in disease pathophysiology; however, their exact contribution and antigen specificities remain unclear. We determined differentiation states and targets of dominant T-cell clones along with their potential to eliminate hematopoietic progenitor cells in the bone marrow of 15 patients with acquired aplastic anemia. Single-cell sequencing and immunophenotyping revealed oligoclonal expansion and effector differentiation of CD8+ T-cell compartments. We reexpressed 28 dominant T-cell receptors (TCRs) of 9 patients in reporter cell lines to determine reactivity with (1) in vitro–expanded CD34+ bone marrow, (2) CD34− bone marrow, or (3) peptide pools covering immunodominant epitopes of highly prevalent viruses. Besides 5 cytomegalovirus-reactive TCRs, we identified 3 TCRs that recognized antigen presented on hematopoietic progenitor cells. T cells transduced with these TCRs eliminated hematopoietic progenitor cells of the respective patients in vitro. One progenitor cell–reactive TCR (11A5) also recognized an epitope of the Epstein-Barr virus–derived latent membrane protein 1 (LMP1) presented on HLA-A∗02:01. We identified 2 LMP1-related mimotopes within the human proteome as activating targets of TCR 11A5, providing proof of concept that molecular mimicry of viral and self-epitopes can drive T cell–mediated elimination of hematopoietic progenitor cells in aplastic anemia.
Introduction
Aplastic anemia is defined by multilineage cytopenia and hypocellular bone marrow. With an incidence of 2 to 3 cases per 1 million inhabitants per year in western countries, the disease is considered rare. This study is focused on acquired/idiopathic aplastic anemia (hereafter referred to as aplastic anemia), which accounts for ∼70% of cases1 and should be distinguished from hereditary bone marrow failure syndromes such as telomere biology disorders that do typically not respond well to immunosuppressive treatment.2,3 The pathophysiology of aplastic anemia is not entirely understood; however, recovery of blood counts upon immunosuppressive therapy with antithymocyte globulin and cyclosporine A with/without eltrombopag, along with a substantial body of data suggest a critical involvement of the immune system, especially of T cells.4-8
Increased frequencies of T memory stem cells at diagnosis are associated with responsiveness to immunosuppressive therapy, and persistently increased frequencies after immunosuppressive therapy correlate with treatment failure or disease relapse.9 Furthermore, T helper type 1 differentiation, T-box expressed in T cells expression, and interferon gamma production by bone marrow and peripheral blood T cells have been shown to be associated with aplastic anemia development.10 Effector memory T cells have been described to be clonally expanded,11 and an association of aplastic anemia with selected human HLA haplotypes and polymorphisms12-14 supports the critical involvement of T cells in immune-mediated bone marrow aplasia.
Yet, clonal T-cell expansion and effector differentiation do not allow conclusions on their potential specificity for hematopoietic progenitor cells. Moreover, clonally expanded effector T cells can be specific for a variety of especially chronically persistent (viral) antigens.15-17 A link between Epstein-Barr virus (EBV) infection and autoimmune multiple sclerosis has recently been established18,19; however, in aplastic anemia, the mechanisms driving progenitor cell–directed T-cell immune responses remain unknown.
In summary, T cells have an undebatable role in aplastic anemia pathophysiology whereas drivers of clonal expansion and particular molecular targets remain incompletely understood.
In this study, we determined clonal bone marrow T-cell expansion and associated immune phenotypes at single-cell resolution, asked whether expanded clones recognized antigens presented on hematopoietic progenitor cells, and determined whether target epitopes were related to (persistent) viral infections.
Methods
Study approval and participants
The study was approved by the local ethics committee (Ethikkommission der Charité Universitätsmedizin Berlin, protocol no. EA2/096/15), all participants gave written informed consent, and all research was conducted in accordance with the Declaration of Helsinki. Bone marrow mononuclear cells were isolated using Ficoll-Paque PLUS (GE Healthcare) and cryopreserved in RPMI-1640 containing 50% fetal bovine serum (FBS, both Thermo Fisher Scientific) and 10% dimethyl sulfoxide (Carl Roth GmbH).
Cell lines
58α−β− cells were previously modified by stable transfection with human CD8 and nuclear factor of activated T cells–inducible green fluorescent protein (GFP) and will hereafter be referred to as 58α−β−.20 U266B1 (RRID: CVCL_0566), L-363 (RRID: CVCL_1357), and HEK293T (RRID: CVCL_0063) cells were cultured as previously described.21
For generation and expansion of stroma cells, human bone marrow mononuclear cells of an independent donor were cultured in stroma cell medium (10 mL StemMACS MSC Expansion Media [Miltenyi Biotec] supplemented with 1 μM hydrocortisone [StemCell Technologies]). Stroma cells were passaged at confluency of almost 100% for 5 passages before cells were used for expansion of CD34+ progenitor cells.
Identity of cell lines was determined by microscopy and flow cytometry. All cell lines were thawed and cultured for at least 5 days in RPMI-1640 (suspension cells) or Dulbecco modified Eagle medium (adherent cells), each supplemented with 10% FBS, 100 U/mL penicillin, and 100 μg/mL streptomycin (both Gibco) before use in functional assays.
Fluorescence-activated cell sorting
Index-sorting for single-cell T-cell receptor αβ (TCRαβ) and phenotype sequencing was performed as previously described.21-24 In brief, bone marrow cells were stained with a 13-parameter panel (supplemental Table 1, available on the Blood website), and 500 to 1000 single αβ T cells were index-sorted into 96-well plates prefilled with One Step reverse transcription polymerase chain reaction (PCR) buffer (Qiagen). Reverse transcription, PCR amplification, and molecular barcoding were performed as described previously. DNA was pooled and sequenced on a MiSeq instrument (Illumina, San Diego, CA). Computational scripts for data processing are available at https://github.com/HansmannLab/TRECA and the primary output can be found in the supplemental Appendix. Cytokines and transcription factors were considered expressed in single cells if we detected >10 reads per cell for the respective cytokine or transcription factor.25 T-cell clones were defined expanded if we detected at least 2 individual cells with identical TCRαβ complementarity determining region 3 (CDR3) amino acid sequences within the same sample.
Flow cytometry cell analysis was carried out using LSRFortessa (BD Biosciences) and Aurora (Cytek) instruments, and data were analyzed using FlowJo software version 10.8.1 (BD Biosciences).
TCR expression in 58α−β− reporter cells
TCR expression in human T cells
Recombinant TCR expression in human lymphocytes was done as previously described.21,26-28 In brief, TCRβ and TCRα sequences modified with murine constant regions were codon optimized and synthesized (ThermoFisher) separated by a P2A cleavage site and cloned into the MP71 vector. Virus was produced after FuGeneHD (Promega) transfection of 293Vec-RD114 producer cells. Mononuclear cells of a healthy donor were stimulated with anti-CD3 and anti-CD28 (both BioLegend, see supplemental Appendix) before transduction. After expansion, transduced cells were rested for 2 days, analyzed by flow cytometry, and cryopreserved. Of TCR 10E7–transduced T cells, 85% were CD8+, and 27% expressed the transgene TCR determined by mouse TCRβ staining. Of TCR 11A5–transduced T cells, 84% were CD8+, and 36% expressed the transgene TCR determined by mouse TCRβ staining.
Magnetic enrichment and in vitro expansion of hematopoietic progenitor cells
CD34+ hematopoietic progenitor cells were enriched using the CD34 MicroBead Kit UltraPure, human (Miltenyi Biotec) according to the manufacturer’s instructions. CD34-enriched cells (10 000, or all CD34-enriched cells available) were expanded on 15 000 human mesenchymal stroma cells seeded in 24-well plates 1 day before the beginning of the expansion in 2 mL StemSpan serum-free medium II (Stemcell Technologies) supplemented with 50 ng/mL stem cell factor, 50 ng/mL thrombopoietin, 50 ng/mL Fms-related tyrosine kinase 3 ligand (all Peprotech), 1 μM StemRegenin1 (Hölzel Diagnostica), and 1 μM UM729 (Stemcell Technologies; called progenitor cell expansion medium), and were expanded at 37°C and 5% CO2 for 8 days without further medium changes. Expanded progenitor cells were harvested, washed in RPMI-1640 containing 10% FBS, and immediately analyzed and/or cultured with T cells.
For patients of whom not enough sample material of the initial bone marrow was available (patients 191, 198, 219, 228, and 230), follow-up samples (Table 1) were used for in vitro expansion of CD34+ cells.
Patient characteristics
| Patient . | Sex . | Initial sampling . | Follow-up . | HLA-class I . | EBV IgG . | CMV IgG . | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Age (y) . | Diagnosis . | Disease state . | Current treatment . | Time after initial sample (mo) . | Disease state . | Current treatment . | |||||
| 105 | M | 19 | mAA | PR after ATG + CsA | CsA + Epag | A∗24:02 B∗07:05; B∗51:01 C∗03:04; C∗15:05 | Pos. | Neg. | |||
| 144 | M | 57 | sAA | Relapse after ATG | CsA + Epag | A∗02:01; A∗03:01 B∗07:02; B∗51:01 C∗02:02; C∗07:02 | Pos. | Pos. | |||
| 191 | F | 69 | sAA | Relapse after ATG + Epag | CsA + Epag | 18 | Relapse | CsA + Epag | A∗24:02 B∗07:02 C∗07:02 | Pos. | Pos. |
| 192 | M | 56 | mAA | Relapse after CsA + Epag | Epag | A∗02:01; A∗32:01 B∗07:02; B∗40:02 C∗02:02; C∗07:02 | Pos. | Neg. | |||
| 198 | F | 48 | vsAA | Initial diagnosis | None | 20 | CR | CsA | A∗02:01; A∗33:01 B∗14:02; B∗40:01 C∗03:04; C∗08:02 | Pos. | Pos. |
| 199 | M | 23 | vsAA | Initial diagnosis | None | A∗01:01; A∗68:02 B∗08:01; B∗14:02 C∗07:01; C∗08:02 | Pos. | Pos. | |||
| 200 | F | 58 | sAA | Initial diagnosis | None | A∗26:01; A∗31:01 B∗40:02; B∗55:01 C∗01:02; C∗03:04 | Pos. | Pos. | |||
| 213 | M | 40 | vsAA | Relapse after ATG + CsA | None | A∗02:05; A∗31:01 B∗18:01; B∗50:01 C∗06:02; C∗07:01 | Pos. | Pos. | |||
| 219 | M | 62 | mAA | Relapse after CsA + Epag | CsA | 10 | Relapse after CsA | CsA | A∗26:01; A∗66:01 B∗07:02; B∗40:02 C∗07:02; C∗15:02 | Pos. | Pos. |
| 222 | F | 56 | vsAA | Initial diagnosis | None | A∗03:01; A∗24:02 B∗07:02; B∗55:01 C∗03:03; C∗07:02 | Pos. | Pos. | |||
| 227 | M | 76 | vsAA | Initial diagnosis | None | A∗02:01; A∗03:01 B∗07:02; B∗35:01 C∗04:01; C∗07:02 | Pos. | Pos. | |||
| 228 | M | 27 | vsAA | Initial diagnosis | None | 17 | CR after ATG + CsA + Epag | CsA | A∗02:01; A∗03:01 B∗07:02; B∗51:01 C∗07:02; C∗14:02 | Pos. | Pos. |
| 230 | M | 66 | mAA | Initial diagnosis | None | 5 17 | Refractory to CsA PR after Danazol | CsA + Epag Danazol + CsA | A∗02:01; A∗03:01 B∗07:02 C∗07:02 | Pos. | Pos. |
| 232 | M | 53 | mAA | Refractory to CsA + Epag | None | A∗02:01; A∗11:01 B∗07:02; B∗08:01 C∗07:01; C∗07:02 | Pos. | Neg. | |||
| 257 | F | 48 | mAA | PR after ATG + CsA | CsA + Epag | A∗02:01; A∗33:01 B∗14:02; B∗51:01 C∗02:02; C∗08:02 | Pos. | Pos. | |||
| Patient . | Sex . | Initial sampling . | Follow-up . | HLA-class I . | EBV IgG . | CMV IgG . | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Age (y) . | Diagnosis . | Disease state . | Current treatment . | Time after initial sample (mo) . | Disease state . | Current treatment . | |||||
| 105 | M | 19 | mAA | PR after ATG + CsA | CsA + Epag | A∗24:02 B∗07:05; B∗51:01 C∗03:04; C∗15:05 | Pos. | Neg. | |||
| 144 | M | 57 | sAA | Relapse after ATG | CsA + Epag | A∗02:01; A∗03:01 B∗07:02; B∗51:01 C∗02:02; C∗07:02 | Pos. | Pos. | |||
| 191 | F | 69 | sAA | Relapse after ATG + Epag | CsA + Epag | 18 | Relapse | CsA + Epag | A∗24:02 B∗07:02 C∗07:02 | Pos. | Pos. |
| 192 | M | 56 | mAA | Relapse after CsA + Epag | Epag | A∗02:01; A∗32:01 B∗07:02; B∗40:02 C∗02:02; C∗07:02 | Pos. | Neg. | |||
| 198 | F | 48 | vsAA | Initial diagnosis | None | 20 | CR | CsA | A∗02:01; A∗33:01 B∗14:02; B∗40:01 C∗03:04; C∗08:02 | Pos. | Pos. |
| 199 | M | 23 | vsAA | Initial diagnosis | None | A∗01:01; A∗68:02 B∗08:01; B∗14:02 C∗07:01; C∗08:02 | Pos. | Pos. | |||
| 200 | F | 58 | sAA | Initial diagnosis | None | A∗26:01; A∗31:01 B∗40:02; B∗55:01 C∗01:02; C∗03:04 | Pos. | Pos. | |||
| 213 | M | 40 | vsAA | Relapse after ATG + CsA | None | A∗02:05; A∗31:01 B∗18:01; B∗50:01 C∗06:02; C∗07:01 | Pos. | Pos. | |||
| 219 | M | 62 | mAA | Relapse after CsA + Epag | CsA | 10 | Relapse after CsA | CsA | A∗26:01; A∗66:01 B∗07:02; B∗40:02 C∗07:02; C∗15:02 | Pos. | Pos. |
| 222 | F | 56 | vsAA | Initial diagnosis | None | A∗03:01; A∗24:02 B∗07:02; B∗55:01 C∗03:03; C∗07:02 | Pos. | Pos. | |||
| 227 | M | 76 | vsAA | Initial diagnosis | None | A∗02:01; A∗03:01 B∗07:02; B∗35:01 C∗04:01; C∗07:02 | Pos. | Pos. | |||
| 228 | M | 27 | vsAA | Initial diagnosis | None | 17 | CR after ATG + CsA + Epag | CsA | A∗02:01; A∗03:01 B∗07:02; B∗51:01 C∗07:02; C∗14:02 | Pos. | Pos. |
| 230 | M | 66 | mAA | Initial diagnosis | None | 5 17 | Refractory to CsA PR after Danazol | CsA + Epag Danazol + CsA | A∗02:01; A∗03:01 B∗07:02 C∗07:02 | Pos. | Pos. |
| 232 | M | 53 | mAA | Refractory to CsA + Epag | None | A∗02:01; A∗11:01 B∗07:02; B∗08:01 C∗07:01; C∗07:02 | Pos. | Neg. | |||
| 257 | F | 48 | mAA | PR after ATG + CsA | CsA + Epag | A∗02:01; A∗33:01 B∗14:02; B∗51:01 C∗02:02; C∗08:02 | Pos. | Pos. | |||
On treatment indicates whether the patient was on treatment at the time of sample acquisition.
ATG, antithymocyte globulin; CR, complete response; CsA, cyclosporin A; Epag, eltrombopag; F, female; IgG, immunoglobulin G; M, male; mAA, moderate aplastic anemia; Neg., negative; PR, partial response; Pos., positive; sAA, severe aplastic anemia; vsAA, very severe aplastic anemia.
Coculture assay
TCR-transgenic 58α−β− were labeled with specific combinations of CellTrace Far Red and CellTrace Violet (both Invitrogen) or left without labeling. Up to 4 CellTrace-labeled cell lines were pooled at 20 000 cells per label and cocultured with 60 000 to 80 000 CD34+exp or CD34− in 96-well U-bottom plates (Corning Incorporated). After 16 hours, cells were stained with APC/Fire750-labeled anti–mouse CD45 and Zombie Yellow (supplemental Table 1).
For HLA-blocking, CD34+exp were incubated with anti-HLA-ABC or isotype control (both at 20 μg/mL), anti–HLA-BC (40 μg/mL) or anti–HLA-A2 (40 μg/mL) for 30 minutes at 37°C before coculture (antibody details in supplemental Table 1).
To determine killing of hematopoietic progenitor cells, 50 000 CD34+exp were seeded in 100 μL progenitor cell expansion medium. TCR-transduced human T cells were thawed and rested for 4 hours in T-cell medium (RPMI-1640 supplemented with 10% FBS, 100 U/mL penicillin, 100 μg/mL streptomycin, 1 mM sodium pyruvate, 1× nonessential amino acids, and 50 μM 2-mercaptoethanol [all Gibco]). T cells were labeled with CellTrace Violet (Invitrogen), and 400 000 cells in 150 μL T-cell medium were added to CD34+exp, resulting in an effector-to-target ratio of 8:1, and cocultured at 37°C and 5% CO2, and harvested after 48 hours. Cells were subsequently stained with ZombieNIR (BioLegend) and analyzed by flow cytometry.
Peptides and peptide pools
The following peptide pools were used according to the manufacturer’s instructions: PepPool CMV (CD4 and CD8) human (Mabtech, order number 3619-1, 42 peptides from cytomegalovirus [CMV]); CERI-MHC Class I Control Peptide Pool (Immunospot, order number CTL-CERI-300, 124 peptides from CMV, EBV, influenza A, and respiratory syncytial virus); and PepTivator pools for pp65, IE1 (Miltenyi Biotec, order numbers 130-093-438, 130-093-493, both CMV), BMLF1, BRLF1, BZLF1, EBNA-1, and latent membrane protein 1 (LMP1; Miltenyi Biotec, order numbers 130-097-283, 130-097-284, 130-093-611, 130-093-613, and 130-095-930, respectively, all EBV). The 2 CMV PepTivator pools and the 5 EBV PepTivator pools were combined at equimolar ratios into 1 CMV and 1 EBV pool for coculture experiments.
For all peptide pools, CD34− bone marrow or peripheral blood mononuclear cells of the individuals of whom the TCRs in question were isolated, were used as antigen-presenting cells.
Individual peptides of the LMP1 PepTivator pool were provided by Miltenyi Biotec.
All single peptides were synthesized by Miltenyi Biotec or Genscript with purity >75% and used at a final concentration of 5 μM if not stated otherwise.
HLA, LMP1, and LMP2A constructs and transfection/nucleofection
Coding sequences of HLA-A∗02:01, A∗03:01, B∗07:02, and C∗07:02 were downloaded from IPD-IMGT/HLA (accession numbers: HLA00005, HLA00037, HLA00132, and HLA00434), cloned into the pHSE3′ expression vector as previously described,20 and transfected using FuGene HD (Promega).
LMP1 (UniProt: P03230) and LMP2A (UniProt: P13285) protein sequences were reverse transcribed, codon optimized, and synthesized (ThermoFisher). Constructs were cloned into the pcDNA6 expression vector with a T2A-linked mCherry. L-363 (91 × 106) were nucleofected using the Nucleofector 4D (Lonza), the SF kit (Lonza), 1 μg plasmid, and program DN-100.
Transfected HEK293T were cultured in Dulbecco modified Eagle medium for 32 hours, and nucleofected L-363 were cultured in RPMI-1640 for 24 hours at 37°C and 5% CO2 before coculture assays.
TCRβ repertoire sequencing
DNA was extracted from bone marrow mononuclear cells or flow cytometry–sorted bone marrow T cells using the AllPrep DNA/RNA Kit (Qiagen). Libraries for amplicon TCRβ next-generation sequencing were prepared according to the EuroClonality-NGS Working Group protocols.29 The total DNA input in the first PCR step was normalized to the equivalent of ∼75 000 nucleated cells. Sequencing was performed on the MiSeq (Illumina) sequencer. Raw sequencing data were demultiplexed using the bcl2fastq conversion software (Illumina) with 0 mismatches in barcode sequences allowed. The resulting fastq files were analyzed using the bioinformatics tool ARResT/Interrogate,30 and functionality (productivity) of each TCRβ rearrangement was assessed using IMGT/V-QUEST.31
Statistics and data visualization
Statistics and data visualization were done using R version 4.2.2.32 Applied tests are stated in the individual figure descriptions.
For uniform manifold approximation and projection and boxplot immune phenotype visualization of expanded T-cell clones, a clone was determined positive for a given parameter if the majority of cells belonging to the clone were positive.
Results
T cells in aplastic anemia bone marrow show oligoclonal expansion and CD8+ cytotoxic effector differentiation
We determined clonal T-cell expansion and associated immune phenotypes in bone marrow of 15 patients with aplastic anemia. The study cohort (10 male and 5 female patients) had a median age of 56 years. Seven patients were newly diagnosed, 8 had received treatment at some point before, and all patients had active disease at the time of initial sampling (detailed characteristics in Table 1).
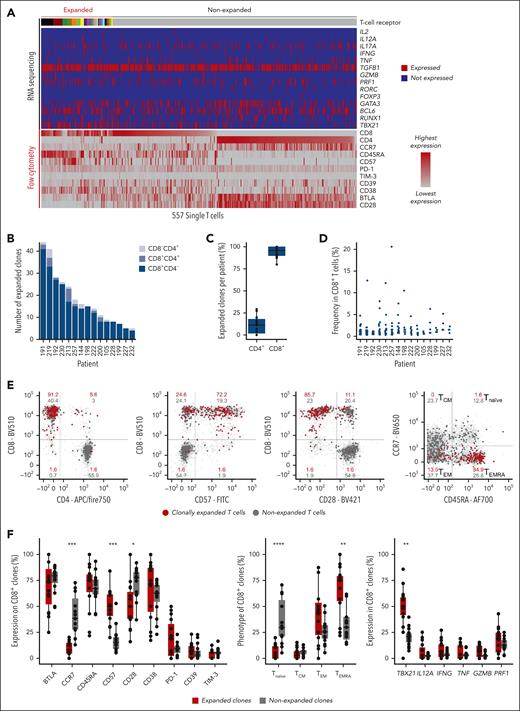
We index sorted (antibody panel in supplemental Table 1) 8144 single bone marrow αβ T cells (median, 569 T cells per patient; range, 344-731 T cells) for subsequent TCRαβ, cytokine, and transcription factor sequencing. Clones with identical CDR3 amino acid sequences in at least 2 cells of the same patient were termed “expanded,” and data of patient 230 are shown in Figure 1A as an example (individual data of all patients in supplemental Figure 1). We detected at median 15 expanded T-cell clones per patient (range, 5-44 expanded clones; Figure 1B). Expanded T-cell clones were almost exclusively CD8+ (92% of expanded clones; Figure 1C) and the most dominant clone within each patient occupied at median 4.7% of CD8+ T cells (range, 1.1%-20.6%; Figure 1D). Of 282 expanded T-cell clones, 2 were detectable in >1 patient, all other expanded clones were unique (supplemental Table 2; supplemental Appendix).
Clonal expansion and immune phenotypes of bone marrow T cells in aplastic anemia. (A) Bone marrow single-cell TCRαβ sequencing and immune phenotyping of patient 230 as an example. Single cells are arranged in columns with each column representing 1 single cell. The top bar indicates TCR sequences; adjacent cells with the same color in the top bar share identical TCRαβ CDR3 amino acid sequences. Clonal expansion was defined as the detection of at least 2 cells with identical TCRαβ CDR3 sequences. The upper part of the heat map indicates single-cell gene expression determined by targeted panel sequencing. The lower part of the heat map visualizes corresponding flow cytometry index-sort data. (B) Numbers of expanded bone marrow T-cell clones per patient. (C) Expression of CD4 and CD8 on expanded T-cell clones. Data points indicate mean frequencies of CD4+ or CD8+ expanded T-cell clones per patient. (D) Frequencies of expanded T-cell clones within total CD8+ T cells. (E) Immune phenotype of index-sorted T cells from patient 230 as an example. Numbers indicate percentages within quadrants. (F) Phenotype determined by flow cytometry (left and middle panels) and RNA sequencing (right panel) for all n = 15 patients. Each data point represents the frequency of positive clones within all expanded (red) or nonexpanded (gray) CD8+ clones of each patient. Boxes reach from the first to third quartiles, lines within boxes indicate medians. The upper whisker extends from the hinge to the largest value, no further than 1.5× interquartile range (IQR). The lower whisker extends from the hinge to the smallest value, at most 1.5× IQR. Statistics were calculated using Wilcoxon signed-rank test and adjusted for multiple testing by Bonferroni correction. ∗P < .05, ∗∗P < .01, ∗∗∗P < .001, and ∗∗∗∗P < .0001. CM, central memory; EM, effector memory; EMRA, effector memory with CD45RA expression.
Clonal expansion and immune phenotypes of bone marrow T cells in aplastic anemia. (A) Bone marrow single-cell TCRαβ sequencing and immune phenotyping of patient 230 as an example. Single cells are arranged in columns with each column representing 1 single cell. The top bar indicates TCR sequences; adjacent cells with the same color in the top bar share identical TCRαβ CDR3 amino acid sequences. Clonal expansion was defined as the detection of at least 2 cells with identical TCRαβ CDR3 sequences. The upper part of the heat map indicates single-cell gene expression determined by targeted panel sequencing. The lower part of the heat map visualizes corresponding flow cytometry index-sort data. (B) Numbers of expanded bone marrow T-cell clones per patient. (C) Expression of CD4 and CD8 on expanded T-cell clones. Data points indicate mean frequencies of CD4+ or CD8+ expanded T-cell clones per patient. (D) Frequencies of expanded T-cell clones within total CD8+ T cells. (E) Immune phenotype of index-sorted T cells from patient 230 as an example. Numbers indicate percentages within quadrants. (F) Phenotype determined by flow cytometry (left and middle panels) and RNA sequencing (right panel) for all n = 15 patients. Each data point represents the frequency of positive clones within all expanded (red) or nonexpanded (gray) CD8+ clones of each patient. Boxes reach from the first to third quartiles, lines within boxes indicate medians. The upper whisker extends from the hinge to the largest value, no further than 1.5× interquartile range (IQR). The lower whisker extends from the hinge to the smallest value, at most 1.5× IQR. Statistics were calculated using Wilcoxon signed-rank test and adjusted for multiple testing by Bonferroni correction. ∗P < .05, ∗∗P < .01, ∗∗∗P < .001, and ∗∗∗∗P < .0001. CM, central memory; EM, effector memory; EMRA, effector memory with CD45RA expression.
Immune phenotypes were heterogeneous, but expanded T-cell clones were more frequently CD57+CD28− compared with nonexpanded T cells. Furthermore, expanded T-cell clones were CD45RA+CCR7−, which is characteristic of effector memory RA (TEMRA) differentiation, and showed a cytotoxic cytokine profile (expression of perforin, granzyme B, and interferon gamma) associated with significantly higher levels of TBX21 than nonexpanded T cells. Data of patient 230 are shown as an example in Figure 1E, combined data of all patients and a selected set of parameters are presented in Figure 1F (for all parameters, see supplemental Figure 2).
Dominant CD8+ T-cell clones eliminate hematopoietic progenitor cells, persist over time, and can be reactive against virus-related epitopes
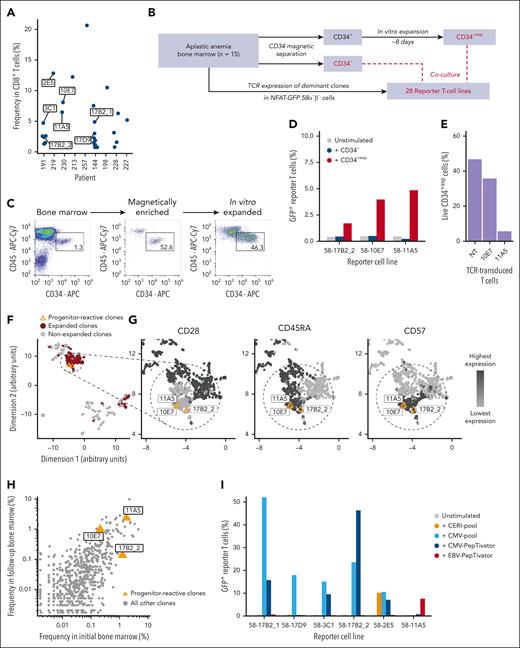
To determine their specificities, we selected 27 dominant CD8+ T-cell clones of 9 patients based on their frequencies (Figure 2A; supplemental Table 3) for TCR reexpression in 58α−β− reporter T cells. One T-cell clone (14G5) expressed 2 productive TCRα chains, which were individually expressed in combination with the corresponding TCRβ chain (named 14G5a1 and 14G5a2) resulting in a total of 28 cell lines each expressing 1 single TCR in question. TCR-transgenic cell lines were named 58–“identifier of the TCR” (eg, 58-11A5) and expressed GFP under control of the nuclear factor of activated T cells promoter resulting in GFP expression upon activation. Each TCR-transgenic cell line was tested for antigen recognition in coculture with (1) CD34-depleted bone marrow (denoted CD34−), and (2) CD34-enriched in vitro–expanded hematopoietic progenitor cells (denoted CD34+exp; schematic in Figure 2B). In vitro expansion of CD34-enriched bone marrow cells was necessary to obtain enough cells for coculture with all TCR-recombinant cell lines. We observed a median 10-fold (range, 7.3-21.3-fold) in vitro expansion while CD34+CD45dim expression was maintained (example shown for patient 198 in Figure 2C; detailed expansion rates and phenotypes of all patients in supplemental Table 4).
Recognition of progenitor cells and viral epitopes by dominant T-cell clones. (A) Frequencies of the 27 dominant CD8+ T-cell clones that were selected for reexpression in reporter cell lines. Clones of further mention in the manuscript were labeled individually. (B) Schematic overview of cell and TCR sources for determination of reactivity with hematopoietic precursor cells. (C) Immune phenotypes of hematopoietic progenitor cells (CD34+CD45dim) upon magnetic enrichment and after in vitro expansion. The plots show data of patient 198 as an example. Flow cytometry plots are pregated on live, single cells. (D) Percentage of GFP+ reporter T cells after coculture with CD34− and CD34+exp for all 3 progenitor cell-reactive TCRs. (E) CD34+exp of patient 230 were cocultured with TCR-transduced or no-transduced (NT) primary human T cells. Bars show percentages of live CD34+exp after coculture determined by flow cytometry. Coculture details and gating strategy are outlined in supplemental Figure 5. (F-G) Uniform manifold approximation and projection (UMAP) visualization of all index-sorted T-cell clones of the 15 patients. Single data points indicate individual clones. The progenitor cell-reactive T-cell clones are highlighted in orange. (H) Relative frequencies of T-cell clones in the initial and follow-up bone marrow samples of patients 191 and 230 determined by TCRβ repertoire sequencing. Orange triangles indicate progenitor cell-reactive clones. (I) Reactivity of 6 TCRs against viral antigens of CMV (orange, light blue, and blue) and EBV (red). CD34− bone marrow or peripheral blood mononuclear cells of the patients in whom the respective TCRs were identified were used as antigen-presenting cells. The progenitor cell-reactive TCR 11A5 also recognized an epitope within the EBV-PepTivator peptide pools.
Recognition of progenitor cells and viral epitopes by dominant T-cell clones. (A) Frequencies of the 27 dominant CD8+ T-cell clones that were selected for reexpression in reporter cell lines. Clones of further mention in the manuscript were labeled individually. (B) Schematic overview of cell and TCR sources for determination of reactivity with hematopoietic precursor cells. (C) Immune phenotypes of hematopoietic progenitor cells (CD34+CD45dim) upon magnetic enrichment and after in vitro expansion. The plots show data of patient 198 as an example. Flow cytometry plots are pregated on live, single cells. (D) Percentage of GFP+ reporter T cells after coculture with CD34− and CD34+exp for all 3 progenitor cell-reactive TCRs. (E) CD34+exp of patient 230 were cocultured with TCR-transduced or no-transduced (NT) primary human T cells. Bars show percentages of live CD34+exp after coculture determined by flow cytometry. Coculture details and gating strategy are outlined in supplemental Figure 5. (F-G) Uniform manifold approximation and projection (UMAP) visualization of all index-sorted T-cell clones of the 15 patients. Single data points indicate individual clones. The progenitor cell-reactive T-cell clones are highlighted in orange. (H) Relative frequencies of T-cell clones in the initial and follow-up bone marrow samples of patients 191 and 230 determined by TCRβ repertoire sequencing. Orange triangles indicate progenitor cell-reactive clones. (I) Reactivity of 6 TCRs against viral antigens of CMV (orange, light blue, and blue) and EBV (red). CD34− bone marrow or peripheral blood mononuclear cells of the patients in whom the respective TCRs were identified were used as antigen-presenting cells. The progenitor cell-reactive TCR 11A5 also recognized an epitope within the EBV-PepTivator peptide pools.
In coculture experiments, 3 TCRs of 2 patients (17B2_2 from patient 191; 10E7 and 11A5 from patient 230) were activated by CD34+exp (Figure 2D; gating strategy in supplemental Figure 3). TCR 17B2_2 was also detectable in patient 144 (17B2_1) but did not recognize CD34+exp of this patient. TCRs 10E7 and 11A5 were unique to patient 230 and activation by CD34+exp was confirmed with (1) follow-up bone marrow of the same patient (clinically in partial response after immunosuppressive therapy at the time of follow-up), and (2) bone marrow of 1 additional patient in complete remission (patient 228), who shared all HLA-class I alleles with patient 230 (supplemental Figure 4; Table 1). Furthermore, when expressed in primary human T cells of a healthy donor, TCRs 10E7 and 11A5 eliminated hematopoietic progenitor cells of patient 230 in vitro (Figure 2E; gating strategy in supplemental Figure 5). None of the 28 TCRs recognized CD34− bone marrow (Figure 2D; supplemental Figure 6).
All 3 T-cell clones that recognized CD34+exp showed CD45RA and CD57 expression (Figure 2F-G) and persisted over time without major changes in relative clone size within the lymphocyte compartment as determined by TCRβ repertoire sequencing at initial and follow-up time points (Figure 2H).
Immune phenotypes of expanded T-cell clones (TEMRA), including those with reactivity against hematopoietic progenitor cells, suggested reactivity with chronically persistent (eg, viral) antigens33; therefore, we tested all 28 reexpressed TCRs for recognition of peptide pools derived from CMV (CMV-pool and CMV-PepTivator), EBV (EBV-PepTivator), and a selection of immunodominant epitopes (CERI-pool) of highly prevalent viruses (influenza A, respiratory syncytial virus, CMV, and EBV). We determined 5 of 28 TCRs to be CMV reactive (Figure 2I), and immune phenotypes of CMV epitope-specific TCRs did not differ from the progenitor cell–reactive TCRs (supplemental Figure 7). However, TCR 11A5 of patient 230, which was reproducibly activated by CD34+exp, also recognized an epitope within the EBV peptide pool (Figure 2I) suggesting reactivity with an EBV-related epitope that was presented on hematopoietic progenitor cells.
TCR 11A5 recognized the EBV LMP1–derived epitope WQLLAFFLA presented on HLA-A∗02:01
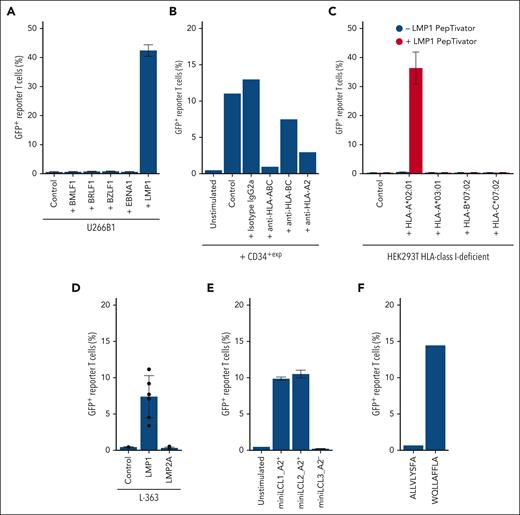
TCR 11A5 recognized an EBV-related epitope presented on CD34+exp, raising the question of its exact target and HLA restriction. To narrow down the target epitope of TCR 11A5, we tested commercially available peptide pools (PepTivator) that cover selected EBV proteins as oligopeptides. The multiple myeloma cell line U266B1 that shared all HLA-class I alleles with patient 230 (HLA type of U266B1; alleles of patient 230 underlined: A∗02:01, A∗03:01, B∗07:02, B∗40:01, C∗03:04, and C∗07:02) was used for peptide presentation. Only the LMP1 peptide pool activated TCR 11A5 (Figure 3A).
58-11A5 recognizes an LMP1-derived peptide in an HLA-A∗02:01–restricted manner. (A) Coculture of 58-11A5 with U266B1 loaded with EBV protein–spanning peptide pools (PepTivator). Frequencies of GFP+ among all 58α−β− were measured to detect T-cell activation. The plot shows median ± standard deviation from n = 3 independent experiments. (B) CD34+exp of an independent individual were incubated with 58-11A5 in presence of blocking antibodies against HLA-ABC, HLA-BC, or HLA-A2. (C) HLA-class I–deficient HEK293T were transfected with all HLA-class I alleles, 1 at a time, of patient 230 (median recombinant HLA-expression: 54%; range, 37.3%-66.2%), loaded with LMP1 PepTivator, and incubated with 58-11A5. The plot shows median ± standard deviation from n = 3 replicates. (D) LMP1 or LMP2A expressing L-363 were cocultured with 58-11A5 (median LMP1 and LMP2A expression: 7.5%; range, 3.5%-15.3%). The plot shows median ± standard deviation from 6 replicates of n = 2 independent experiments. (E) Coculture of HLA-A∗02:01+ (A2+) or A2− miniLCLs with 58-11A5. The plot shows median ± standard deviation from n = 3 replicates. All plots show frequencies of GFP+ as percentage of 58α−β− cells. “Control” indicates coculture with nontreated/nontransfected cells as negative control. “Unstimulated” indicates 58-11A5 alone. (F) Only WQLLAFFLA of the LMP1 PepTivator pool activated 58-11A5. All other LMP1 PepTivator pool peptides did not activate 58-11A5 as shown for ALLVLYSFA as a representative example.
58-11A5 recognizes an LMP1-derived peptide in an HLA-A∗02:01–restricted manner. (A) Coculture of 58-11A5 with U266B1 loaded with EBV protein–spanning peptide pools (PepTivator). Frequencies of GFP+ among all 58α−β− were measured to detect T-cell activation. The plot shows median ± standard deviation from n = 3 independent experiments. (B) CD34+exp of an independent individual were incubated with 58-11A5 in presence of blocking antibodies against HLA-ABC, HLA-BC, or HLA-A2. (C) HLA-class I–deficient HEK293T were transfected with all HLA-class I alleles, 1 at a time, of patient 230 (median recombinant HLA-expression: 54%; range, 37.3%-66.2%), loaded with LMP1 PepTivator, and incubated with 58-11A5. The plot shows median ± standard deviation from n = 3 replicates. (D) LMP1 or LMP2A expressing L-363 were cocultured with 58-11A5 (median LMP1 and LMP2A expression: 7.5%; range, 3.5%-15.3%). The plot shows median ± standard deviation from 6 replicates of n = 2 independent experiments. (E) Coculture of HLA-A∗02:01+ (A2+) or A2− miniLCLs with 58-11A5. The plot shows median ± standard deviation from n = 3 replicates. All plots show frequencies of GFP+ as percentage of 58α−β− cells. “Control” indicates coculture with nontreated/nontransfected cells as negative control. “Unstimulated” indicates 58-11A5 alone. (F) Only WQLLAFFLA of the LMP1 PepTivator pool activated 58-11A5. All other LMP1 PepTivator pool peptides did not activate 58-11A5 as shown for ALLVLYSFA as a representative example.
To narrow down the HLA restriction, we used CD34+exp of an independent bone marrow donor that activated 58-11A5 and expressed 3 of 4 HLA-class I alleles of patient 230 (HLA type of the independent donor, shared alleles underlined: A∗02:01, A∗29:02, B∗07:02, B∗39:01, C∗07:01, and C∗07:02). CD34+exp of the independent donor were cocultured with 58-11A5 in presence of blocking antibodies against pan HLA-class I (HLA-ABC), HLA-BC, or HLA-A2. Blocking pan HLA-class I and HLA-A2 substantially reduced 58-11A5 activation by CD34+exp suggesting HLA-A∗02:01 restriction (Figure 3B). To confirm restriction of TCR 11A5 for HLA-A∗02:01, we transfected HLA class I–deficient HEK293T with all HLA class I alleles of patient 230, 1 at a time (transfection efficiency in supplemental Table 5), loaded them with LMP1 PepTivator, and tested activation of 58-11A5. Activation of 58-11A5 by LMP1 PepTivator specifically required HLA-A∗02:01 expression (Figure 3C).
To determine the role of LMP1 as a potentially relevant target antigen for TCR 11A5, we investigated whether the target epitope could be efficiently processed and presented. We expressed full length LMP1, or LMP2A as a negative control, in the multiple myeloma cell line L-363, which endogenously expresses HLA-A∗02:01 (called L-363LMP1 or L-363LMP2A, nucleofection efficiency in supplemental Table 6). Only L-363LMP1 but not L-363LMP2A activated 58-11A5 (Figure 3D).
In addition to transient LMP1 expression in L-363, we tested miniLCL cells that express a selected set of EBV proteins including LMP1.34 HLA-A∗02:01+ but not HLA-A2− miniLCL activated 58-11A5 (Figure 3E).
To determine the target epitope of TCR 11A5, we tested all LMP1 PepTivator peptides individually for activation of 58-11A5. Only WQLLAFFLA activated 58-11A5 when loaded onto L-363 as antigen–presenting cells (Figure 3F).
TCR 11A5 recognizes epitopes on hematopoietic progenitor cells via molecular mimicry
The LMP1-derived epitope WQLLAFFLA is not part of the human proteome and its identification as target of TCR 11A5 raised the question whether TCR activation by CD34+exp was because of potentially EBV-infected cells in the cocultures or mechanisms of molecular mimicry (a similar but not identical epitope presented on hematopoietic progenitor cells). EBV primarily infects B lineage and epithelial cells, and patient 230 (in whom TCR 11A5 was expanded) was EBV seropositive (Table 1). However, CD19+ B-lineage cells within the CD34− population of patient 230 accounted for 3.7% of nucleated cells, did not activate 58-11A5, and showed similar frequencies in in vitro–expanded CD34+exp cells (supplemental Figure 8), rendering potentially EBV-infected B cells within the cocultures an unlikely trigger for TCR activation. Furthermore, activation of 58-11A5 by CD34+exp from bone marrow of a healthy EBV-negative donor (HLA type: A∗02:01, B∗07:02, B∗13:02, C∗06:02, C∗07:02; supplemental Figure 9) and elimination of these CD34+exp by TCR 11A5–transduced healthy donor T cells (supplemental Figure 10) confirmed an LMP1-independent mimotope as target peptide presented on CD34+exp cells.
To identify potential mimotopes that activate TCR 11A5, we performed a National Center for Biotechnology Information (NCBI) protein BLAST search (https://blast.ncbi.nlm.nih.gov) using WQLLAFFLA as input sequence. From the top results of the BLAST output, we selected 5 peptides (Figure 4A) and tested them for activation of 58-11A5. WQLLSFFLL and LQLLAFILA derived from alternative protein NUP205 and Claudin-20, respectively, activated TCR 11A5 (Figure 4B) providing proof of concept that a virus-reactive TCR expanded in the bone marrow of a patient with aplastic anemia can be cross-reactive with human auto-epitopes.
LMP1-related mimotopes that activate 58-11A5. (A) Results of an NCBI protein BLAST search (https://blast.ncbi.nlm.nih.gov, accessed in October 2023) of the human proteome (taxid:9606) using “WQLLAFFLA” as input sequence. Sequences to be tested for 58-11A5 activation (highlighted in yellow) were selected by investigators’ choice considering numbers and positions as well as size, charge, and hydrophobicity of exchanged amino acids. Amino acids that differ from the LMP1 target peptide are colored in red. (B) Mimotopes that activated 58-11A5. The LMP1 epitope WQLLAFFLA was included as positive control, GFPT1-derived LQLLAFHLA was shown as negative control and representative example for peptides that did not activate 58-11A5. Amino acids that differ from WQLLAFFLA are colored in red. The plot shows frequencies of GFP+ as percentage of 58α−β− cells. Data indicate median ± standard deviation of n = 3 replicates.
LMP1-related mimotopes that activate 58-11A5. (A) Results of an NCBI protein BLAST search (https://blast.ncbi.nlm.nih.gov, accessed in October 2023) of the human proteome (taxid:9606) using “WQLLAFFLA” as input sequence. Sequences to be tested for 58-11A5 activation (highlighted in yellow) were selected by investigators’ choice considering numbers and positions as well as size, charge, and hydrophobicity of exchanged amino acids. Amino acids that differ from the LMP1 target peptide are colored in red. (B) Mimotopes that activated 58-11A5. The LMP1 epitope WQLLAFFLA was included as positive control, GFPT1-derived LQLLAFHLA was shown as negative control and representative example for peptides that did not activate 58-11A5. Amino acids that differ from WQLLAFFLA are colored in red. The plot shows frequencies of GFP+ as percentage of 58α−β− cells. Data indicate median ± standard deviation of n = 3 replicates.
In summary, expanded T-cell clones in the bone marrow of patients with aplastic anemia can eliminate hematopoietic progenitor cells in an antigen-dependent fashion. Molecular mimicry of epitopes derived from EBV and related human proteins presented on hematopoietic progenitor cells potentially contribute to immune-mediated bone marrow aplasia as presented for TCR 11A5 as an example.
Discussion
A critical role of T cells in aplastic anemia has already been suggested by indirect experimental and clinical evidence. We systematically studied clonal expansion, associated immune phenotypes, targeted cell populations, and target antigens of bone marrow T cells during the clinical course of aplastic anemia. Our data provide experimental proof that expanded bone marrow T-cell clones can eliminate hematopoietic progenitor cells and that epitopes derived from chronic viral infections can potentially drive hematopoiesis-directed T-cell responses by molecular mimicry.
T-cell expansion is an antigen-driven process and T cells can be grouped into functional units by their TCRαβ CDR3 sequences. We have developed a methodology that combines multidimensional single-cell flow cytometry index-sorting with TCRαβ and phenotype sequencing for determination of clone-associated T-cell differentiation at gene expression and protein levels.25,35 This methodology is highly accurate, efficient, and allows expression of T-cell clones of choice for downstream determination of potential target epitopes.23,24 Our experimental approach was optimized for immune phenotyping and TCR reexpression of dominant clones, which, in case of autoreactivity, can be expected to be enriched at sites of inflammation, in this case the bone marrow. Clonal expansion occurred predominantly within the CD8+ bone marrow T-cell compartment, and expanded clones showed TEMRA differentiation, which is in line with previous studies,9 and a frequent differentiation state of T cells associated with persistent viral infections.36,37
Despite advances in computational grouping of TCRs by similar specificities,38,39 reliable target identification requires TCR reexpression and experimental investigation in functional assays. Dominant TEMRA differentiation of expanded bone marrow T-cell clones encouraged us to determine reactivity with peptide pools derived from chronic and frequent viral infections. In total, we determined 6 of 28 dominant TCRs to be virus specific, which is not surprising given that bone marrow is a site of T- and B-cell memory,15 and major proportions of expanded bone marrow and circulating T-cell clones can be specific for persistent pathogens.16,17 Furthermore, reactivation of herpesviruses can frequently occur upon immunosuppressive therapy.40 Peptide pools cannot cover all potential viral epitopes, and nonreactivity of a TCR does not exclude specificity for the respective virus. Indeed, we suspect frequencies of virus-specific T cells among dominant clones to be substantially higher; nevertheless, to our knowledge, this study is the first to systematically address virus reactivity of dominant bone marrow T-cell clones in aplastic anemia bone marrow with a standardized and comprehensive set of immunodominant epitopes of highly prevalent viruses.
Of 28 dominant TCRs, we identified 3 to be progenitor cell reactive, and determination of reactivity required in vitro expansion of hematopoietic progenitor cells. Progenitor cells are rare; sample size is limited, especially in hypocellular conditions such as aplastic anemia; and in vitro expansion was necessary to obtain enough cells for cocultures with T-cell lines of choice. There are a variety of protocols for in vitro expansion of hematopoietic progenitor cells41,42; our protocol was modified from 4 different protocols,43-46 and maintained the CD34+CD45dim immune phenotype during expansion. We cannot exclude that antigen abundance, processing, or presentation were affected by in vitro expansion processes, and it is difficult to predict whether additional TCRs would have been reactive against CD34+exp under different expansion conditions. Furthermore, bone marrow of patients with active aplastic anemia could be depleted of some of the immunologically targeted progenitor cell populations and could recover heterogeneity during in vitro expansion. The exact target populations among CD34+exp cannot be concluded from our data; however, CD34− cells did not activate any of the dominant T-cell clones, and selective activation of all 3 progenitor cell–reactive TCRs by CD34+exp was confirmed with follow-up bone marrow of the same patients.
For tracking T-cell clones over time, we applied TCRβ repertoire sequencing as it exceeds single-cell approaches in sequencing depth while preserving relative quantitation between samples at an affordable cost. Persistence of hematopoietic progenitor cell–reactive T-cell clones over time has previously been suggested47; however, one could hypothesize that hematopoiesis-reactive T-cell clones should decrease in frequency or disappear upon successful immunosuppressive therapy. The hematopoiesis-reactive T-cell clones identified in 2 patients of our cohort persisted over time and both patients had active disease at the studied time points. Whether frequencies of hematopoiesis-reactive T-cell clones could potentially be used as predictive biomarkers, requires investigation in larger cohorts.
One of 3 hematopoietic progenitor cell–reactive TCRs recognized the LMP1-derived epitope WQLLAFFLA in an HLA-A∗02:01–restricted manner. We proved experimentally that the target epitope can be processed from full-length LMP1 in B-lineage cells presenting EBV as a relevant driver of hematopoietic progenitor cell–reactive T-cell expansion in aplastic anemia. HLA-A∗02:01 restriction is in line with a recent study determining this allele as a frequent risk allele in aplastic anemia.48 Cross-reactivity of the LMP1-reactive TCR 11A5 with epitopes presented on hematopoietic progenitor cells suggested molecular mimicry. We identified 2 additional target epitopes of TCR 11A5 that were part of the human proteome. Identification of these epitopes was based on an NCBI BLAST search followed by manual selection of promising candidates, taking into account numbers and positions as well as size, charge, and hydrophobicity of exchanged amino acids. We identified NUP205- and CLDN20-derived epitopes that activated TCR 11A5 and were closely related to the LMP1 target epitope. NUP205 and CLDN20 are expressed in a variety of tissues including the bone marrow49 and present only 2 of a potentially larger set of cross-reactive epitopes that could not be entirely identified by our approach. Furthermore, tissue-specific antigen processing is likely involved in activation of TCR 11A5 exclusively by CD34+exp but not CD34− bone marrow.
Clinical evidence for a link between EBV or other viral infections and aplastic anemia has so far been limited to case reports.50-55 However, the recent identification of HLA risk alleles48 and long-term incidence reduction associated with the COVID-19 pandemic56 suggest critical roles for T-cell responses to viral antigens in aplastic anemia pathophysiology. Whether herpesvirus reactivation, which can frequently occur under immunosuppressive therapy,40 is critical for the observed clonal T-cell expansion, cannot be concluded from our data. We would rather hypothesize that initial EBV infection associated with a state of infection-induced inflammation triggers expansion of EBV-specific T-cell clones that, later on, after currently unknown additional cues, eliminate hematopoietic progenitor cells via molecular mimicry. It is likely that other not only viral diseases can potentially trigger immune-mediated bone marrow aplasia. Furthermore, aplastic anemia follows a bimodal age distribution; because exposure to different viruses is associated with age, results of our cohort comprised of relatively older individuals require confirmation in younger patients.
In conclusion, we provide experimental evidence that antigens associated with persistent viral infections can mimic epitopes presented on hematopoietic progenitor cells driving hematopoiesis-directed effector T-cell reactions in aplastic anemia.
Acknowledgments
The authors thank Cornelia Eckert and Juan Lázaro-Navarro for help with stroma cell cultures. The authors are grateful to Kirstin Rautenberg and Hans-Peter Rahn at MDC Berlin for expert support with flow cytometry sorting, as well as to Linda Hammerich and Michaela Schwarz at Charité Universitätsmedizin Berlin for providing analytical flow cytometry infrastructure and clinical data.
This work was supported by Deutsche Krebshilfe e.V. (70113355) (L.H.), the German Cancer Consortium (L.H.), and the European Union (ERC Advanced Grant NeoT 882963, to T.B.). A.B.H. was supported by the scholarships BIA-MD-Promotionsstipendium of the Berlin Institute of Health and Deutschlandstipendium.
Authorship
Contribution: A.B.H. and L.H. conceived the project; A.B.H., C.W., M.B., S.S., J.S., H.H., K. Dornmair, F.B., and L.H. designed experiments; T.W. and A.M. provided critical material; T.H.B., F.B., and L.H. provided and interpreted clinical data; T.H.B. and F.B. contributed critical clinical expertise; A.B.H., C.W., S.S., and K. Dietze performed experiments; A.B.H., C.W., S.S., M.B., L.B., T.B., L.R., M.L., T.K., W.H., T.H.B, O.B, J.S., H.H., K. Dornmair, F.B., and L.H. analyzed and interpreted data; O.B. provided critical reagents; A.B.H. and L.H. wrote the manuscript with input from all authors; and L.H. coordinated and supervised the project.
Conflict-of-interest disclosure: A.M. reports collaboration with Biosyngen. F.B. received research support from Novartis and Pfizer; and reports research collaboration with RepeatDx. L.B. serves on advisory committees for AbbVie, Amgen, Astellas, Bristol Myers Squibb, Celgene, Daiichi Sankyo, Gilead, Hexal, Janssen, Jazz Pharmaceuticals, Menarini, Novartis, Pfizer, Sanofi, and Seattle Genetics; and reports research support from Bayer and Jazz Pharmaceuticals. T.H.B. is a consultant for Janssen, Merck, Novartis, and Pfizer; received research support from Novartis and Pfizer; and reports research collaboration with RepeatDx. L.H. serves on advisory committees for Janssen-Cilag, Pierre-Fabre, and Sanofi; and received travel support from Amgen, Gilead, and Janssen-Cilag. M.B. serves on advisory committees for Amgen and Incyte; serves on the speaker’s bureau of Amgen, Becton Dickinson, Janssen, and Pfizer; and reports research support from Amgen. M.L. reports grant/research support from T-knife GmbH. The remaining authors declare no competing financial interests.
Correspondence: Leo Hansmann, Department of Internal Medicine III, University Hospital Regensburg, Franz-Josef-Strauß-Allee 11, 93053 Regensburg, Germany; email: leo.hansmann@ukr.de.
References
Author notes
F.B. and L.H. are joint senior authors.
All data have been made publicly available along with this manuscript in a supplemental Appendix. Original data are available on request from the corresponding author, Leo Hansmann (leo.hansmann@ukr.de).
The online version of this article contains a data supplement.
There is a Blood Commentary on this article in this issue.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.



This feature is available to Subscribers Only
Sign In or Create an Account Close Modal