Abstract
Introduction
The emergence of next generation RNA sequencing (RNA-Seq) technologies will likely advance diagnostic, prognostic and therapeutic strategies for patients (pts) with various cancers.Novel fusions have recently been described in AML and solid tumors using RNA-Seq, and many were out-of-frame.It is not known whether novel fusions are generated at diagnosis (Dx) of CML and if so, their impact on treatment outcome. We used RNA-Seq coupled with whole exome sequencing to identify and characterize novel fusions at Dx of CML and at blast crisis (BC). A highly complex pattern of genomic rearrangements of chromosome (chr) 9 and 22 was found in some pts at Dx that generated novel fusions associated with multiple genomic breaks, multiple non-contiguous deletionsand inversion of genomic sequences, including BCR and ABL.
Method
RNA-Seqwas performed on Dx samples of chronic phase pts treated with first line TKI representing 2 extreme response groups: 14 pts with BC at a median of 6 months (mos), range 3-25 (group A, poor response), and 16 pts with rapid major molecular response by 3mosof imatinib (group B, optimal response). RNA-Seqwas also performed for 9 of 14 pts at BC (group C).The TruSeq Stranded Total RNA-RiboZero Gold Sample Prep Kit (Illumina) was used. This method enables computation of transcription direction and detection of genomic breaks from precursor RNA. Fusions were identified usingthe STAR algorithm and those detected in 4 normal controls were filtered out. Fusions with a high unique read count, supporting genomic breaks or detection atDxand BC for individual pts were prioritized for validation and their somatic status confirmed by RT-PCR.Correspondingwhole exome sequencingwas conducted for 30 samples. Copy number variation was detected usingSequenzaand exon level resolution ofdeletionswas achieved using an in-house sequence read normalization method.
Results
BCR-ABL fusions were detected by RNA-Seq in 29/30 pts at Dx and all pts at BC. In addition, novel fusions were identified in eachptgroup.
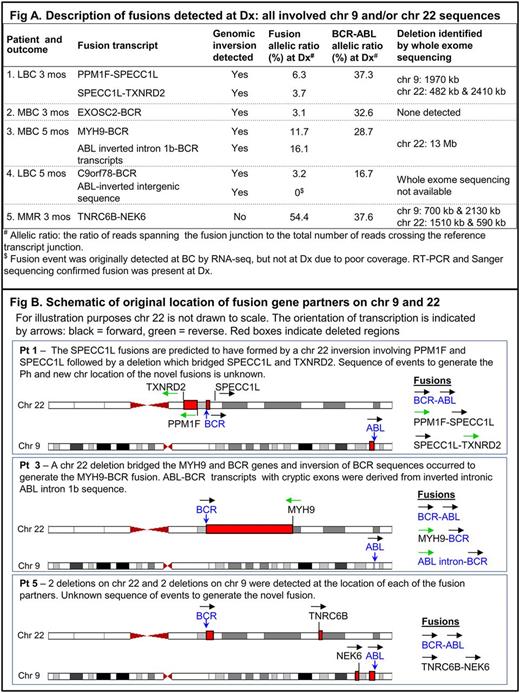
GroupA(poor response). AtDx, 8 cytogenetically cryptic novel fusion transcripts were detected in 4/14 pts, Fig A pts1-4. All fusions involved genes or sequences onchr9 and/or 22 and all 4 pts had concomitant genomic inversion events. Fusion partners included inverted ABL intronic sequences and an inverted intergenic region on chr 22, potentially derived from the generation and activation of cryptic splice sites. BCR was a frequent fusion partner (5/8 fusion transcripts). Genomicdeletionswere detected adjacent to some fusions (3deletionsin 1pt),indicatingdeletionsmay have contributed to fusion formation, Fig B. All 4 pts with novel fusions and inversions had very rapid BC (within 5mosofDx).
Group B (optimal response).AtDx, only 1/16 pts had a fusion detected in addition to BCR-ABL: TNRC6B (chr22)-NEK6 (chr9), Fig Apt5. Thisptalso had multiple non-contiguousdeletions: 2 each onchr9 and 22 associated with fusion formation, but no inversions, Fig B.
Group C (BC). At BC, 3/9 pts gained fusions. No inversions were detected. Two pts had MLL fusions; MLL-BCAT1 (novel) and MLL-MLLT6. The MLL gene is a known fusion partner in acute leukemia, associated with poor prognosis. Both pts had sudden onset BC after a complete cytogenetic response. These fusions were supported by translocation events detected by cytogenetic analysis;t(11;12)(q23;p12) and t(11;17)(q23;q21). The thirdptgained an out-of-frame ANKRD11-UBQLN1 fusion at BC. Indeed, ANKRD11 expression was reduced by 3-fold at BC. Interestingly, thispthad a germline gain of function TP53 mutation. ANKRD11 is a p53 coactivator and loss of expression defined poor prognosis in breast cancer pts that harbored gain of function p53 mutations (Noll, 2012). The ANKRD11 fusion detected at BC in CML may have been selected with disease progression in the context of mutant p53.
Conclusion
We identified a subset of pts with novel fusions and inversion events at Dx involvingchr9 and 22. These inversions were detected among the pts studied with very rapid BC. The biological effects of the novel fusions remain to be determined. Our data support the presence of novel fusions, additional to BCR-ABL in CML and add a further layer of genetic heterogeneity associated with the Philadelphia translocation. Whether genomic inversions identify a small subset of CML pts with very poor prognosis requires expanded analysis.
Yeung:BMS: Honoraria, Membership on an entity's Board of Directors or advisory committees, Research Funding; Novartis Pharmaceuticals: Honoraria, Membership on an entity's Board of Directors or advisory committees, Research Funding; Ariad: Research Funding. Mueller:Ariad: Honoraria; Institute for Hematology and Oncology GmbH: Employment; Bristol-Myers Squibb: Honoraria; Novartis: Honoraria; Pfizer: Honoraria. Dietz:Institute for Hematology and Oncology GmbH: Employment. Ross:Novartis Pharmaceuticals: Honoraria, Research Funding; BMS: Honoraria. Hughes:Ariad: Honoraria, Membership on an entity's Board of Directors or advisory committees, Research Funding; BMS: Honoraria, Membership on an entity's Board of Directors or advisory committees, Research Funding; Novartis: Honoraria, Membership on an entity's Board of Directors or advisory committees, Research Funding. Branford:Novartis: Honoraria, Membership on an entity's Board of Directors or advisory committees, Research Funding; Ariad: Research Funding; Bristol Myers Squibb: Research Funding; Qiagen: Honoraria, Membership on an entity's Board of Directors or advisory committees; Cepheid: Consultancy.
Author notes
Asterisk with author names denotes non-ASH members.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal