Abstract
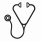
Introduction
New drugs have improved survival for multiple myeloma (MM) patients, however, patient outcome remains highly variable, unpredictable and often very poor. To identify novel treatments and potential biomarkers, we applied high throughput ex vivo drug sensitivity testing combined with exome and transcriptome sequencing to samples collected from newly diagnosed and relapsed MM patients. Integration of results from the different platforms indicated several oncogenic signaling pathways driving drug response and highlighted the importance of a multi-targeted approach for treatment.
Methods
Bone marrow (BM) aspirates (n=48) were collected from MM patients (newly diagnosed n=14; relapsed/refractory n=26) and healthy individuals (n=8). CD138+ plasma cells were enriched by Ficoll separation followed by immunomagnetic bead selection. Cells were screened against 306 oncology drugs with the drugs tested in a 10,000-fold concentration range. Drug sensitivity scores were calculated based on the normalized area under the dose response curve (Yadav et al, Sci Reports, 2014). Importantly, MM selective responses were determined by comparing data from MM patients with those of healthy BM cells. Clustering of drug sensitivity profiles was performed using unsupervised hierarchical ward-linkage clustering with Spearman and Manhattan distance measures of drug and sample profiles. Somatic mutations were identified by exome sequencing of DNA from CD138+ cells and skin biopies from each patient, while gene expression profiles were derived from RNA sequencing of CD138+ cells.
Results
Cluster analysis of drug response profiles segregated the samples into four MM specific groups (Figure). Group I patients (n=12) were highly sensitive to many drugs, including several signal transduction inhibitors such as those targeting PI3K-AKT, MAPK and IGF pathways, as well as HSP90 and BCL2 inhibitors plus epigenetic/chromatin modifiers such as BET and HDAC inhibitors. Group II (n=15) showed a more modest response profile and were moderately sensitive to signal transduction inhibitors and epigenetic modifiers. Group III (n=9) were largely insensitive to most drugs in the panel except for BCL2 and proteasome inhibitors, while group IV (n=3) were resistant to all drugs except BCL2 inhibitors. Many samples were selectively sensitive to navitoclax (55%), dual PI3K/mTOR inhibitors (45%) and aminopeptidase inhibitors (20%), which had little effect on healthy control or MM CD138- cells. Only 33% of the samples responded to glucocorticoids. The majority of samples including healthy BM controls were sensitive to proteasome and CDK inhibitors, suggesting low selective cytotoxicity. However, drug sensitivity profiles of healthy control and CD138- cell populations were distinct from MM CD138+ samples indicating that observed CD138+ drug responses were specific for malignant plasma cells. In addition, we observed that drugs with overlapping target profiles tended to cluster together, indicating sample responses were similar to related drugs.
Diagnostic and relapse samples were spread across the different response groups. Samples with mutations to genes involved in PI3K and NF-κB signaling tended to cluster in group I, while most samples with t(4;14) fell in Group II. Samples with RAS mutations were present in all response groups and no correlation with MEK inhibitor sensitivity was observed. 17p deletion samples were also found in all response groups, however, those with additional TP53 mutation tended to have increased drug sensitivity.
Summary
Our results indicate that PI3K/mTOR, MAPK, IGF1R, NF-κB and cell survival (e.g. BCL2, BCLXL) signaling are important pathways mediating MM ex vivo drug response. This matched with genomic and transcriptomic data, which identified alterations of genes involved in these pathways. Although additional work is needed to correlate ex vivo drug sensitivity with in vivo treatment response, our initial results suggest the possibility that MM patients could be subjected to stratified treatment based on combined ex vivo drug testing and molecular profiling. In addition, these results highlight the multiple signaling pathways active in MM and emphasize the need for improved combination strategies for treatment.
Subgrouping of MM patient samples (I-IV) based on selective drug response profiles. H/D/R denotes healthy, diagnostic and relapse, respectively.
Subgrouping of MM patient samples (I-IV) based on selective drug response profiles. H/D/R denotes healthy, diagnostic and relapse, respectively.
Silvennoinen:Research Funding of Finland Government, Research Funding from Janssen-cilag, research funding from Celgene: Research Funding; Janssen-Cilag, Sanofi, Celgene: Honoraria. Wennerberg:Pfizer: Research Funding. Kallioniemi:Medisapiens: Consultancy, Membership on an entity's Board of Directors or advisory committees. Porkka:Bristol-Myers Squibb: Honoraria, Research Funding; Novartis: Honoraria, Research Funding. Heckman:Celgene: Research Funding.
Author notes
Asterisk with author names denotes non-ASH members.
This icon denotes a clinically relevant abstract

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal