Abstract
Classical Hodgkin lymphoma (cHL) and mediastinal large B-cell lymphoma (MLBCL) are lymphoid malignancies with certain shared clinical, histologic, and molecular features. Primary cHLs and MLBCLs include variable numbers of malignant cells within an inflammatory infiltrate, suggesting that these tumors escape immune surveillance. Herein, we integrate high-resolution copy number data with transcriptional profiles and identify the immunoregulatory genes, PD-L1 and PD-L2, as key targets at the 9p24.1 amplification peak in HL and MLBCL cell lines. We extend these findings to laser-capture microdissected primary Hodgkin Reed-Sternberg cells and primary MLBCLs and find that programmed cell death-1 (PD-1) ligand/9p24.1 amplification is restricted to nodular sclerosing HL, the cHL subtype most closely related to MLBCL. Using quantitative immunohistochemical methods, we document the association between 9p24.1 copy number and PD-1 ligand expression in primary tumors. In cHL and MLBCL, the extended 9p24.1 amplification region also included the Janus kinase 2 (JAK2) locus. Of note, JAK2 amplification increased protein expression and activity, specifically induced PD-1 ligand transcription and enhanced sensitivity to JAK2 inhibition. Therefore, 9p24.1 amplification is a disease-specific structural alteration that increases both the gene dosage of PD-1 ligands and their induction by JAK2, defining the PD-1 pathway and JAK2 as complementary rational therapeutic targets.
Introduction
Classical Hodgkin lymphoma (cHL) is a B-cell malignancy that occurs frequently in Western countries and commonly affects young adults.1 These tumors are characterized by small numbers of neoplastic Reed-Sternberg (RS) cells within an extensive inflammatory/immune cell infiltrate. There are 4 subtypes of cHL, 2 of which comprise ≈ 90% of cases: nodular sclerosing Hodgkin lymphoma (NSHL; 60% of cases) and mixed cellularity Hodgkin lymphoma (MCHL; 30% of cases). cHLs lack surface immunoglobulin expression and B-cell receptor–mediated signals and rely on alternative survival pathways, including aberrant nuclear factorκB signaling.1
In previous studies, we and others have defined shared molecular features of cHL and a specific subtype of diffuse large B-cell lymphoma (DLBCL), primary mediastinal large B-cell lymphoma (MLBCL).2,3 Like cHL, MLBCLs have a T-helper cell type 2 (Th2)–skewed cytokine profile, decreased expression of B-cell receptor signaling pathway components, and constitutive activation of nuclear factorκB.2 MLBCL also exhibits certain clinical and histologic similarities to cHL, particularly the NSHL subtype.4,5 For example, both diseases are most common in young adults and often present as an anterior mediastinal or localized nodal mass.2,4,5 In addition, both MLBCLs and NSHLs include bands of sclerotic tissue and immune/inflammatory cell infiltrates.4,5 However, the inflammatory infiltrate is less prominent in MLBCLs, which have a more diffuse growth pattern.4
Although cHLs have an extensive polymorphous inflammatory infiltrate, there is little evidence of an effective host antitumor immune response. In fact, recent studies indicate that Hodgkin RS cells produce certain molecules that limit the efficacy of T cell–mediated antitumor immune responses.1,6 For example, Hodgkin RS cells selectively express the immunoregulatory glycan-binding protein, galectin-1, which fosters a Th2/T regulatory cell–skewed tumor microenvironment.6 Primary HL RS cells also variably express programmed cell death-1 ligand 1 (PD-L1)/B7H1, whereas tumor-infiltrating T cells express the coinhibitory receptor, programmed death-1 (PD-1).7 Similarly, primary MLBCLs are reported to express PD-L2.3
The natural function of PD-1 signaling is to limit certain T cell–mediated immune responses.8 Normal antigen-presenting cells, dendritic cells, and macrophages express PD-1 ligands that engage PD-1 receptors on activated T cells.8,9 On ligand binding, the PD-1 receptor recruits the Src homology 2 domain–containing protein tyrosine phosphatase-2 (SHP2) phosphatase to the immunoreceptor complex, resulting in dephosphorylation of proximal T-cell receptor (TCR) signaling molecules (CD3δ, ζ-associated protein 70 (ZAP70), and protein kinase C θ (PKCθ) and attenuation of TCR signaling.8 In addition, PD-L1 inhibits CD28 costimulation by competitively binding to the CD28 ligand, CD80 (B7-1).10 PD-1 signaling results in “T-cell exhaustion,” a temporary inhibition of activation and proliferation that can be reversed on removal of the PD-1 signal. Furthermore, PD-L1 also promotes the induction and maintenance of PD-1+ T regulatory cells.11
Emerging data suggest that viruses and tumors have developed mechanisms that exploit the PD-1 pathway to evade immune detection. In models of chronic viral infection, engagement of PD-1 receptors triggers T-cell exhaustion and the progressive loss of effector T-cell function and proliferative capacity.8 In murine cancer models, the tumor cell expression of PD-1 ligands inhibits T-cell activation and promotes the apoptosis of tumor-specific T cells.12,13 PD-1 ligands are also expressed and associated with an unfavorable prognosis in multiple human tumors, including malignant melanoma, colon, pancreatic, hepatocellular, and ovarian carcinomas.14-19 Despite the prognostic significance of PD-1 ligand expression and the demonstrated role of PD-1 signaling in tumor immune privilege, structural genetic mechanisms for deregulated PD-1 ligand expression in cancer have not been described.
The PD-1 ligand genes, PD-L1 and PD-L2, are located on chromosome 9p24.1 and separated by only 42 kilobases.8 Of interest, 9p copy gain has been described in both HL and MLBCL with low-resolution techniques such as comparative genomic hybridization.20,21 Several genes residing on 9p have been postulated to play a role in cHL and MLBCL, although the key targets of this genetic alteration3,21-23 remain undefined. Herein, we integrate copy number data from high-density single nucleotide polymorphism (HD SNP) arrays with paired transcriptional profiles and identify the PD-1 ligands as key targets of the 9p24.1 amplification in NSHL and MLBCL. In addition, we characterize a novel regulatory loop in which Janus kinase 2 (JAK2), located 322 kilobases upstream from PD-L1 on 9p24.1, further augments PD-1 ligand expression in these tumors.
Methods
Cell Lines
This study was approved by the Institutional Review Board of the Dana-Farber Cancer Institute and Brigham and Women's Hospital. The HL cell lines L428, L1236, and KMH2 and the DLBCL cell lines SUDHL4, OCILy1, KARPAS422, PFEIFFER, and SUDHL6 were grown in RPMI medium supplemented with 10% heat-inactivated fetal bovine serum (FBS). The HL cell lines L-540 and HD-LM-2 and the MLBCL cell line KARPAS1106P were maintained in RPMI with 20% heat-inactivated FBS. The HL cell line SUPHD1 was grown in McCoy 5A with 20% heat-inactivated FBS. Media for all cell lines were supplemented with 10mM HEPES (N-2-hydroxyethylpiperazine-N′-2-ethanesulfonic acid) buffer, 4mM l-glutamine, 50U/mL penicillin, and 50 U/mL streptomycin.
Integrative analysis
Genomic DNAs from 18 DLBCL, 1 MLBCL, and 6 HL cell lines and 21 peripheral blood lymphocyte samples from healthy donors were extracted as previously described and profiled with Affymetrix SNP6.0 microarrays,24 which include > 900 000 SNP probes and ∼ 950 000 additional probes for copy number.
The inference of DNA copy number from .cel files was performed with a previously described GenePattern pipeline.25 Data are available from the Gene Expression Omnibus (National Center for Biotechnology Information, http://www.ncbi.nlm.nih.gov/geo/) under accession no. GSE22208. Data were segmented with the Circular Binary Segmentation algorithm,26,27 and naturally occurring copy number variants were removed before assessment of the significance of copy number alterations across samples with the GISTIC algorithm.28 GISTIC defines regions of interest with associated false discovery rate FDR Q values below a predefined threshold and a smaller peak (or peaks) within each region defined by the set of contiguous markers with the highest Q values.
Transcriptional profiling was performed for all cell lines with the use of Affymetrix U133 A&B microarrays as previously described.6 Integrative analysis, combining DNA copy number and gene transcript data, was performed to assess relationships between DNA copy number change and alteration in gene transcript abundance to refine the list of candidate genes within alteration regions. Genes within the peak (region) of GISTIC-identified alterations were tested for difference in expression between samples with or without each lesion by a 2-group t statistic, and significance was assessed both by permutation test and by asymptotically derived P values. FDR Q values were derived by taking the union of all genes within all the peaks (regions). Genes were considered positive by integrative analysis if differences in transcript abundance attained an FDR ≤ 0.25 by at least 1 of the 2 procedures (permutation-based or asymptotic). Additional detailed integrative analysis methods are included in the supplemental data (available on the Blood Web site; see the Supplemental Materials link at the top of the online article).
Flow cytometry
Flow cytometry was performed by resuspending 1 × 106 cells in 100 μL of PBS, incubating with 500 μg of antibody for 30 minutes at room temperature, washing once in PBS, and resuspending in a volume of 500 μL. Cells were stained with phycoerythrin-conjugated antibodies specific for PD-L1 (Clone 29E.2A3), PD-L2 (Clone 24F.10C12), or the analogous isotype controls (immunoglobulin G2b κ and immunoglobulin G2a κ, respectively; BioLegend). After staining, 20 000 cells were analyzed with a BD FACSCanto flow cytometer (BD Biosciences). FlowJo software (TreeStar) was used for selection of viable cells by forward and side scatter and subsequent generation of histograms and median fluorescence intensities. Fluorescence intensities for each antigen were normalized to their respective isotype controls.
Intracellular phospho-flow was performed by fixing cells with BD CytoFix fixation buffer (BD Biosciences) followed by permeabilization with BD Perm Buffer II (BD Biosciences) and staining with phycoerythrin-conjugated antibodies to signal transducer and activator of transcription 1 (STAT1; pY701; BD Biosciences). Phospho-flow was normalized to fixed, permeabilized, and unstained cells.
Primary tumor specimens
Primary tumor specimens included 23 cHLs and 41 primary MLBCLs. All specimens were identified by clinical criteria and pathologic features and reviewed by expert hematopatholigists (S.J.R. and J.L.K.) to confirm diagnosis. Primary cHLs included 7 of the MCHL subtype and 16 of the NSHL subtype. Primary MLBCLs included 33 previously described tumors2 and 8 additional cases.
Laser-capture microdissection of primary Hodgkin RS cells
To isolate neoplastic RS cells from primary cHLs, specimens were sectioned and immunostained for the RS cell marker, CD30. Rapid immunohistochemistry (IHC) was performed with 5-μm-thick frozen tissue sections. Briefly, slides were fixed in −20°C acetone and air-dried for 10 minutes. All further steps were performed at room temperature in a hydrated chamber. Slides were pretreated with Peroxidase Block (Dako North America Inc) for 5 minutes to quench endogenous peroxidase activity. For CD30, monoclonal mouse anti–human CD30 antibody (Dako North America Inc) was applied ready to use for 10 minutes. Slides were washed in 50mM Tris (tris(hydroxymethyl)aminomethane)–Cl, pH 7.4, and incubated with anti–mouse Envision+ kit (Dako North America Inc) for 5 minutes. After further washing, immunoperoxidase staining was developed with a diaminobenzidine (DAB) chromogen (Dako North America Inc). Slides were then redehydrated by passing through graded alcohols and xylene and air-dried before use in laser-capture microdissection. Immediately after staining, between 250 and 1000 CD30+ cells per section were laser-capture microdissected from 3 sections per tumor with the use of a PixCell II (Arcturus). Normal tissue, free from CD30+ cells, was also isolated from each section as a control. Caps were placed immediately into lysis buffer, and DNA was isolated with PicoPure DNA Extraction kits (Arcturus). The samples were analyzed on a PixCell II laser capture microscope with the objective lens of 40×/0.60 Olympus LCPlanFL (Olympus). The pictures were taken with a Cohu 3122-1000 color CCD camera and analyzed with PixCell II acquistion software (Acturus) and Adobe Photoshop 6.0 (Adobe).
Quantitative polymerase chain reaction analysis of DNA copy number and transcript abundance
PD-L1 DNA copy number was assessed with TaqMan DNA Copy Number Assay (Hs03704252_cn; Applied Biosystems) and TaqMan Copy Number Reference Assay RNase P (Applied Biosystems). Polymerase chain reactions (PCRs) were performed with TaqMan Universal Genotyping Master Mix (Applied Biosystems) according to the manufacturer's protocol on an ABI 7300 real-time PCR machine (Applied Biosystems). Before interrogation of primary specimens, the assay was validated with data obtained from HD SNP array analysis of HL cell lines. First, the RNase P gene was determined not to be in a region of DNA copy number alteration. Second, quantitative PCR (qPCR)–based DNA copy number calls were found to tightly correlate with DNA copy number inferences from the HD SNP array analysis (Pearson correlation coefficient = 0.896; P = .003). RS cell PD-L1 DNA copy number was inferred from fold change in DNA with reference to tumor-matched normal DNA as calculated by the 2−ΔΔCT method.29 Significant differences between the ΔCT values of RS cells and normal tissue from a given tumor were determined with a one-tailed paired samples t test. In primary MLBCL samples, copy number was inferred by normalization to data obtained from 5 normal samples.
In primary MLBCLs, complementary DNA was also synthesized by reverse transcription of 500 ng of total RNA with the use of the Superscript III First Strand Synthesis System for RT-PCR (Invitrogen). Gene transcript abundance was assessed by quantitative real-time PCR (qRT-PCR) with the use of commercially available TaqMan Gene Expression Assays (Applied Biosystems) for PD-L1 (Hs01125299_m1) and PD-L2 (Hs01057775_m1) relative to the internal reference gene HPRT1 (Hs99999909_m1). Reactions were prepared with TaqMan Gene Expression Master Mix (Applied Biosystems) according to the manufacturer's protocol.
Quantitative IHC
IHC was performed with 4-μm thick formalin-fixed, paraffin-embedded tissue sections. Briefly, slides were soaked in xylene, passed through graded alcohols, and put in distilled water. All further steps were performed at room temperature in a hydrated chamber. Slides were pretreated with Peroxidase Block (Dako North America Inc) for 5 minutes to quench endogenous peroxidase activity. Slides were washed in 50mM Tris-Cl, pH 7.4, and then blocked with an Avidin/Biotin Blocking Kit (Vector) according to the manufacturer's instructions. For PD-L1, monoclonal mouse anti–human PD-L1 antibody (BioLegend) was applied 1:1000 in DAKO diluent for 1 hour. For PD-L2, monoclonal mouse anti–human PD-L2 antibody (BioLegend) was applied at 1:250 in DAKO diluent for 1 hour. Slides were washed and detected with anti–mouse Envision+ kit (Dako North America Inc) according to the manufacturer's instructions. After another wash, the slides were treated with a Tyramide kit (PerkinElmer) at 1:250 for 10 minutes. Slides were then thoroughly washed and treated with LSAB2 Streptavidin-horseradish peroxidase (Dako North America Inc) according to the manufacturer's instruction. After further washing, immunoperoxidase staining was developed with a DAB chromogen (Dako North America Inc) and counterstained with hematoxylin. For cHL tumors, PD-L1 protein expression was quantified in 150 individual RS cells/slide with the use of an Aperio Scan Scope XT workstation, ImageScope software, and the Aperio Color Deconvolution v9 immunohistochemical analysis algorithm that provided the optical density of DAB staining per RS cell. For MLBCL tumors, PD-L2 staining was quantified in 10 tumor-involved nonsclerotic regions per specimen with the use of the same Aperio Color Deconvolution algorithm and the average optical density per analyzed tumor region. The samples were analyzed using an Olympus BX41 microscope with the objective lens of 40×/0.75 Olympus UPlanFL (Olympus). The pictures were taken using Olympus QColor5 and analyzed with acquistion software QCapture Pro 6.0 (QImaging) and Adobe Photoshop 6.0 (Adobe).
Chemical JAK2 inhibition
To assess the effects of JAK2 inhibition on PD-1 ligand expression, 4 × 106 cells from representative cHL lines were treated in triplicates with either dimethylsulfoxide (vehicle control) or 2.5-10.0μM of the specific JAK2 inhibitor SD-1029 (Calbiochem)30 for 24 hours. Thereafter, cells were harvested to isolate total RNA or to prepare whole-cell extracts for Western blot. After purification of total RNA with the use of an RNeasy Mini Kit (QIAGEN), PD-L1 and PD-L2 transcript abundance were assessed by qPCR. Abundance of active phospho-JAK2 and total JAK2 protein were assessed by Western blot with the use of anti-pY1007/1008-JAK2 (Cell Signaling Technology) and anti-JAK2 (Cell Signaling Technology) antibodies. Anti–glyceraldehyde-3-phosphate dehydrogenase (Abcam) was used as a loading control.
Antiproliferative effects of JAK2 inhibitors were assessed by MTS (3-(4,5-dimethylthiazol-2-yl)5-(3-carboxymethoxyphnyl)-2-(4-sulfophenyl)-2H-tetrazolium) assay. Briefly, 5000 cells per well were exposed to a 2-fold serial dilution series of the specific JAK2 inhibitors, SD-1029 and Z3 (Calbiochem)31 between the concentrations of 0.625 and 20.0μM. After 48 hours of incubation, cellular respiration was measured with the CellTiter 96 Aqueous One Solution Cell Proliferation Assay (Promega). Median effective concentrations (EC50s) were calculated by fitting sigmoidal dose response to triplicate measurements of each concentration with the use of GraphPad Prism and averaged over 3 independent experiments.
PD-L1 promoter region analysis and luciferase assays
Computational analysis of PD-L1 genomic sequences was performed with the USCS Genome Browser (http://www.genome.ucsc.edu), the publicly available MatInspector module of Genomatix suite (http://www.genomatix.de), and the open-source TOUCAN software.32 Briefly, putative promoter regions were identified by high interspecies sequence conservation and high PolI I binding peaks with the use of the Yale TFBS tracks within the UCSC Genome Browser. Sequences were then interrogated for transcription factor binding sites and transcription factor modules with the use of MatInspector and TOUCAN, respectively.
The candidate PD-L1 promoter sequence (−281 to +43 base pair [bp] relative to the transcription start site) was PCR-amplified from the L428 cHL line and cloned into the promoterless pGL3 luciferase vector (Promega). The cHL cell lines L428 and SUPHD1 were grown to approximately 80% confluence, and 4 × 106 cells each were cotransfected with 1.0 μg/well of pGL3 luciferase construct (empty vector or pGL3-PD-L1p) and 0.5 μg/well pRL-TK (Promega). After 24 hours of incubation, cells were treated with 10μM SD-1029 or the equivalent volume of dimethylsulfoxide. Importantly, this inhibitor maintains specificity for JAK2 at 10μM. After an additional 24 hours of incubation, cells were lysed, and luciferase activities were determined by chemiluminescence assay with the use of the Dual Luciferase Assay kit (Promega) and Luminoskan Ascent luminometer (Thermo Lab Systems).
Results
Chromosome 9p24.1 amplification and increased expression of the PD-1 ligands in cHL and MLBCL cell lines
We first performed genomewide DNA copy number analyses of a large panel of cHL and DLBCL cell lines and an additional MLBCL cell line with the use of a HD SNP array with > 900 000 SNP probes and an additional ≈ 950 000 probes for copy number variation.28 Copy number alterations were assessed with the GISTIC method that computes separate scores for amplification and deletion for each probe, taking into account both the frequency and average amplitude of the observed alteration.28 This approach identified highly significant amplification of chromosome 9p24.1 (Q value, 0.002) in the lymphoma cell line panel. The amplified 9p24.1 segment includes 977 genes within a 22-Mbp region (chr9:1-21944952) and 7 genes within the 177-Kbp amplification peak (chr9:5404875-5581849; Figure 1A). The 9p24.1 amplification was present in 100% (6 of 6) of HL cell lines but only 22% (4 of 18) of DLBCL lines (P = .001; FDR, 0.024); notably, the single MLBCL cell line also had amplification of this region.
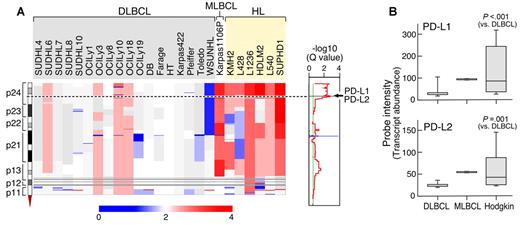
Chromosome 9p24.1 amplification and increased expression of PD-1 ligands in HL and MLBCL cell lines. (A left) Smoothed chromosome 9p gene copy number estimates for each DLBCL, MLBCL, and HL cell line. The color scale ranges from blue (deletion) to red (amplification). (Right) GISTIC Q values (for all cell lines) for the 9p24.1 amplification that includes the PD-L1/PD-L2 loci. (B) PD-L1 and PD-L2 transcript abundance in the DLBCL, MLBCL, and HL cell lines was assessed by transcriptional profiling and represented in box plots.
Chromosome 9p24.1 amplification and increased expression of PD-1 ligands in HL and MLBCL cell lines. (A left) Smoothed chromosome 9p gene copy number estimates for each DLBCL, MLBCL, and HL cell line. The color scale ranges from blue (deletion) to red (amplification). (Right) GISTIC Q values (for all cell lines) for the 9p24.1 amplification that includes the PD-L1/PD-L2 loci. (B) PD-L1 and PD-L2 transcript abundance in the DLBCL, MLBCL, and HL cell lines was assessed by transcriptional profiling and represented in box plots.
Integrative analyses identified C9ORF46 (FDR = 0.0062), PD-L1 (CD274/B7H1; FDR = 0.0399), and PD-L2 (PDCD1LG2/CD273/B7-DC; FDR = 0.0870)8 as the genes with the most significant association between DNA copy number and transcript abundance in the amplification peak (supplemental data). Consistent with these findings, PD-L1 and PD-L2 transcripts were significantly more abundant in HL cell lines than in DLBCL lines and high in the single MLBCL cell line (Figure 1B).
We next assessed the potential association between PD-1 ligand gene copy number, transcript abundance, and cell-surface protein expression in the lymphoma cell line panel. In flow cytometric analysis, the DLBCL cell lines largely lacked cell-surface expression of PD-1 ligands, PD-L1 and PD-L2 (Figure 2). In marked contrast, the HL cell lines with increased copies of 9p24.1 had significantly higher cell-surface expression of the PD-L1 and PD-L2 proteins (Figure 2). The single MLBCL cell line also had high expression of cell-surface PD-1 ligand (Figure 2). Taken together, these data indicate that 9p24.1 amplification in HL and MLBCL cell lines targets the PD-1 ligand genes and increases the cell-surface expression of PD-L1 and PD-L2 (Figures 1–2).
9p24.1 amplification and PD-1 ligand cell-surface expression in HL and MLBCL cell lines. Flow cytometric analysis of cell-surface PD-1 ligand (PD-L1 and PD-L2) expression in DLBCL, MLBCL, and HL cell lines (PD-L1 or PD-L2, open black lines; isotype controls, solid gray histograms). DLBCL and HL cell lines are arranged according to PD-1 ligand copy number (normal to high, left to right).
9p24.1 amplification and PD-1 ligand cell-surface expression in HL and MLBCL cell lines. Flow cytometric analysis of cell-surface PD-1 ligand (PD-L1 and PD-L2) expression in DLBCL, MLBCL, and HL cell lines (PD-L1 or PD-L2, open black lines; isotype controls, solid gray histograms). DLBCL and HL cell lines are arranged according to PD-1 ligand copy number (normal to high, left to right).
9p24.1 amplification and increased PD-L1 expression in primary HL RS cells
Given the identification of PD-1 ligands as key targets of the 9p24.1 amplification in HL cell lines, we next evaluated the frequency of PD-1 ligand gene amplification and overexpression in primary cHL RS cells. For these studies, primary Hodgkin RS cells were laser-capture microdissected from a series of primary NSHLs (n = 16) and MCHLs (n = 7; Figure 3A). Thereafter, PD-L1 gene copy number in primary HL RS cells was assessed by qPCR relative to matched normal tissue (Figure 3B). Of note, LCM Hodgkin RS cell specimens included a small amount of surrounding normal tissue (Figure 3A), probably causing an underestimation of PD-L1 copy number in samples with 9p24.1 amplification (Figure 3B). With the use of the statistically significant differences in ΔCT values between tumor and matched normal cells (corresponding to tumor DNA copy numbers > 2.2), 38% (6 of 16) of primary NSHLs had PD-L1/9p24.1 amplification. In marked contrast, none of the primary MCHLs had PD-L1/9p24.1 amplification (Figure 3B NSHL vs MCHL 9p24.1 amplification; P = .032). These data indicate that 9p24.1 amplification is restricted to the cHL subtype, NSHL, most closely related to MLBCL.
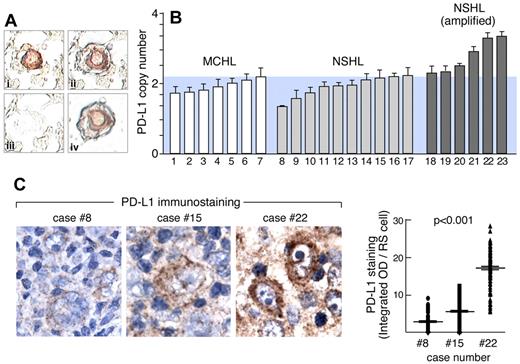
PD-1 ligand amplification and overexpression in primary HL. (A) Laser-capture microdissection (LCM) of primary HL Reed-Sternberg RS cells. (Ai) RS cells were identified by CD30 staining, (Aii) selected by laser (Arcturus PixCell II), (Aiii) removed from surrounding nonneoplastic tissue, resulting in (Aiv) highly enriched RS cell specimens. (B) qPCR-based DNA copy number analysis of PD-L1 in LCM primary RS cells isolated from 7 MCHLs and 16 NSHLs. Data are means (± SD) of triplicate qPCR reactions performed on pooled DNA samples from RS cells or normal tissue from each slide, repeated for 3 slides per sample. (C left) Quantitative IHC of PD-L1 in 3 representative cases (8, 15, and 22) from panel B. (C right) Quantitative analysis of PD-L1 immunostaining in 150 individual RS cells from each of these cases (8, 15, and 22).
PD-1 ligand amplification and overexpression in primary HL. (A) Laser-capture microdissection (LCM) of primary HL Reed-Sternberg RS cells. (Ai) RS cells were identified by CD30 staining, (Aii) selected by laser (Arcturus PixCell II), (Aiii) removed from surrounding nonneoplastic tissue, resulting in (Aiv) highly enriched RS cell specimens. (B) qPCR-based DNA copy number analysis of PD-L1 in LCM primary RS cells isolated from 7 MCHLs and 16 NSHLs. Data are means (± SD) of triplicate qPCR reactions performed on pooled DNA samples from RS cells or normal tissue from each slide, repeated for 3 slides per sample. (C left) Quantitative IHC of PD-L1 in 3 representative cases (8, 15, and 22) from panel B. (C right) Quantitative analysis of PD-L1 immunostaining in 150 individual RS cells from each of these cases (8, 15, and 22).
In representative primary NSHLs with known PD-L1 copy numbers, we also performed quantitative IHC and determined PD-L1 protein expression in 150 RS cells/tumor (Figure 3C-D). With this sensitive IHC method, PD-L1 copy number and protein expression were tightly correlated in primary NSHLs (Figure 3C-D; P < .001).
PD-1 ligand amplification and overexpression in primary MLBCLs
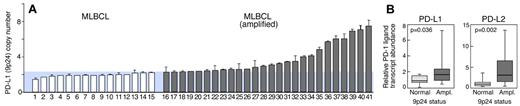
The previously identified similarities between NSHL and primary MLBCL,2 the selective amplification of PD-1 ligands in NSHL (Figure 3), and the observed PD-1 ligand gene amplification and cell-surface expression in a MLBCL cell line (Figures 1–2) prompted us to assess PD-1 ligand copy numbers in a series of 41 primary MLBCLs by qPCR (Figure 4). PD-L1 (9p24.1) amplification was detected in 63% (26 of 41) of primary MLBCLs (Figure 4A). Thereafter, PD-1 ligand transcript abundance was evaluated by qRT-PCR and found to be significantly higher in primary MLBCLs with the 9p24.1 amplification (Figure 4B).
PD-1 ligand amplification and overexpression in primary MLBCL. (A) qPCR-based DNA copy number analysis of PD-L1 in 41 primary MLBCLs. (B) RT-qPCR analysis of PD-L1 and PD-L2 transcript abundance in the same series of primary MLBCLs. Transcript abundance in tumors with normal or increased 9p24.1 is represented in box blots.
PD-1 ligand amplification and overexpression in primary MLBCL. (A) qPCR-based DNA copy number analysis of PD-L1 in 41 primary MLBCLs. (B) RT-qPCR analysis of PD-L1 and PD-L2 transcript abundance in the same series of primary MLBCLs. Transcript abundance in tumors with normal or increased 9p24.1 is represented in box blots.
Because there were larger changes in PD-L2 than in PD-L1 transcript abundance in primary MLBCLs with 9p24.1 amplification (Figure 4B), we analyzed PD-L2 protein expression by quantitative IHC in representative MLBCLs from this series. In these tumors, 9p24.1 amplification was associated with increased PD-L2 protein expression (supplemental data). Taken together, these data confirm an association between amplification of 9p24.1 and increased expression of PD-1 ligands, particularly PD-L2, in primary MLBCLs.
9p24.1 amplification, JAK2 overexpression, and increased activity in HL and MLBCL cell lines
In normal immune cells, PD-1 ligands are induced by cytokine-mediated activation of JAK2/STAT1 signaling. These observations were of particular interest because JAK2 is located on chromosome 9p24.1 (chr9:4985245-5128183). In our pilot series of lymphoma cell lines, JAK2 was coamplified with the PD-L1 and PD-L2 loci (chr9:5450559-5468477 and chr9:5510545-5571282, respectively) as part of the broader 9p24.1 amplification region in HL and MLBCL cell lines (Figure 5A). Consistent with these findings, HL and MLBCL cell lines had significantly higher JAK2 transcript levels than DLBCL cell lines (Figure 5B). In addition, primary MLBCLs with 9p24.1 (PD-L1) amplification had significantly higher JAK2 transcript levels than primary MLBCLs with normal 9p24.1 copy numbers (Figures 4,5B). There was also a close association between JAK2 copy numbers, transcript abundance, and total and active (phosphorylated) JAK2 in HL and MLBCL cell lines (Figure 5C).
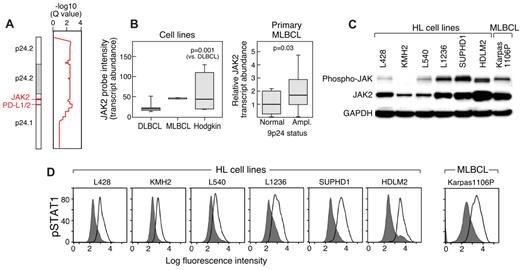
JAK2 amplification, expression, and activity. (A) GISTIC analysis (for all cell lines) of 9p24.1, PD-L1 and PD-L2, in the amplification peak and JAK2 in the broader amplification region. (B) JAK2 transcript abundance in HL and DLBCL cell lines and the single MLBCL cell line (left) and primary MLBCLs with 9p24.1 (PD-L1) amplification or normal copy number (right). (C) Western blot analysis of phospho- and total JAK2 in HL cell lines arranged according to 9p24.1 copy number (normal to high, left to right, as in Figure 2) and the single MLBCL cell line. (D) Intracellular phosphoflow cytometric analysis of phosphorylated STAT1 in HL and MLBCL cell lines in panel C.
JAK2 amplification, expression, and activity. (A) GISTIC analysis (for all cell lines) of 9p24.1, PD-L1 and PD-L2, in the amplification peak and JAK2 in the broader amplification region. (B) JAK2 transcript abundance in HL and DLBCL cell lines and the single MLBCL cell line (left) and primary MLBCLs with 9p24.1 (PD-L1) amplification or normal copy number (right). (C) Western blot analysis of phospho- and total JAK2 in HL cell lines arranged according to 9p24.1 copy number (normal to high, left to right, as in Figure 2) and the single MLBCL cell line. (D) Intracellular phosphoflow cytometric analysis of phosphorylated STAT1 in HL and MLBCL cell lines in panel C.
To evaluate the functional consequences of JAK2 amplification, we assessed the abundance of phosphorylated STAT1 (pSTAT1) in the HL and MLBCL cell lines with intracellular phospho-specific flow cytometry. HL and MLBCL cell lines with increased copies of 9p24.1 and JAK2 had higher levels of intracellular phosphorylated STAT1, directly associating JAK2 amplification with increased JAK2 activity (Figure 5D).
JAK2-associated induction of PD-L1 in HLs
Given the increased JAK2/STAT1 activity in HL and MLBCL cell lines with 9p24.1 amplification, we postulated that this genetic alteration would augment JAK2/STAT1 induction of PD-1 ligands. For this reason, we treated HL cell lines with low- or high-level 9p24.1 amplification (L428 and SUPHD1, respectively) with a chemical inhibitor of JAK2, SD-1029, and assessed associated changes in PD-L1 transcript abundance (Figure 6; supplemental data). In both cell lines, SD-1029 treatment resulted in a dose-dependent decrease in phosphoJAK2 and PD-L1 transcript abundance (Figure 6A-B). Of note, JAK2 inhibition decreased PD-L1 transcripts to a greater extent in SUPHD1, which had higher JAK2 copy numbers and activity (Figures 5C-D,6A-B).
Chemical inhibition of JAK2 decreases PD-1 ligand transcription. (A) Western analysis of phosphoJAK2 in HL cell lines (L428 and SUPHD1) treated with the increasing doses (2.5-10μm) of the specific JAK2 inhibitor, SD-1029. (B) RT-qPCR analysis of PD-L1 transcript abundance in the cell lines treated with SD-1029. Data are means (± SD) of triplicate measurements from the representative experiment shown in panel A. (C) The PD-L1 promoter regulatory module. STAT-responsive (ISRE) element and additional degenerate STAT-binding sites are indicated. This region was cloned into a pGL-luciferase vector (pGL3-PD-L1p). (D) Analysis of pGL3-PD-L1p luciferase activity in L428 and SUPHD1 HL cells treated with SD-1029 or vehicle. Data are means (± SD) of triplicate measurements from a representative experiment. Experiments in panels B and D were performed 3 times.
Chemical inhibition of JAK2 decreases PD-1 ligand transcription. (A) Western analysis of phosphoJAK2 in HL cell lines (L428 and SUPHD1) treated with the increasing doses (2.5-10μm) of the specific JAK2 inhibitor, SD-1029. (B) RT-qPCR analysis of PD-L1 transcript abundance in the cell lines treated with SD-1029. Data are means (± SD) of triplicate measurements from the representative experiment shown in panel A. (C) The PD-L1 promoter regulatory module. STAT-responsive (ISRE) element and additional degenerate STAT-binding sites are indicated. This region was cloned into a pGL-luciferase vector (pGL3-PD-L1p). (D) Analysis of pGL3-PD-L1p luciferase activity in L428 and SUPHD1 HL cells treated with SD-1029 or vehicle. Data are means (± SD) of triplicate measurements from a representative experiment. Experiments in panels B and D were performed 3 times.
We further characterized the role of JAK2/STAT1 signaling in PD-1 ligand induction by directly evaluating the PD-L1 upstream regulatory region. The 5′ PD-L1 sequence (−281 bp upstream to +43 bp downstream of the transcription start site) includes an interferon (IFN)–stimulated regulatory element/IFN-regulatory factor 1 (ISRE/IRF1) module (−125 bp to −188 bp) and several degenerate STAT binding sites (Figure 6C). To assess the effect of JAK2 on transcription mediated by this regulatory element, the 5′ PD-L1 sequence was cloned into the pGL3 luciferase vector and transfected into HL cell lines with low (L428) or high (SUPHD1) 9p24.1 copy numbers. PD-L1 promoter-driven luciferase activity was increased in both L428 and SUPHD1 (Figure 6D empty vector [control] vs pGL3-PD-L1 [control]). Of note, SUPHD1 exhibited significantly higher PD-L1 promoter-driven luciferase expression than L428 (Figure 6D SUPHD1 pGL-3PD-L1p [control], compare y-axes). In addition, treatment with the specific chemical JAK2 inhibitor, SD-1029, resulted in a highly significant decrease in PD-L1 prometer–driven luciferase expression in SUPHD1 and a more modest decrease in L428 (Figure 6D compare pGL3-PD-L1p [control] vs pg3-PD-L1;[SD-1029]). Similar, more modest results were observed after small interfering RNA–mediated knock-down of JAK2 (supplemental data). Taken together, these studies confirm a direct association between 9p24.1 amplification, JAK2 abundance and activity, and PD-L1 up-regulation in HL (Figures 5–6).
We also investigated the role of JAK2 in PD-L2 induction. Like PD-L1, the PD-L2 5′ regulatory region includes an ISRE/IRF1 module and degenerate STAT binding sites (supplemental data). However, the spacing of ISRE and IRF1 elements in the predicted module was larger than that seen in other transcription regulatory elements,33 including the 5′ PD-L1 sequence, and there were fewer degenerate STAT binding sites (supplemental data). Consistent with these findings, JAK2 inhibition decreased PD-L2 transcript abundance and PD-L2 promoter-driven luciferase expression less than PD-L1 (supplemental data).
Together, these results implicate JAK2 in the induction of PD-L1 and, to a lesser extent, PD-L2 gene expression. Therefore, coamplification of JAK2 and PD-1 ligand genes increases both PD-1 ligand gene dosage and the abundance and activity of the PD-1 ligand inducer, JAK2.
JAK2 inhibition and HL cellular proliferation
Given the pleiotropic effects of JAK2/STAT1 signaling and the additional known STAT1 targets, we also assessed the effects of 2 different chemical inhibitors of JAK2 (SD1029 and Z3) on proliferation of HL and MLBCL cell lines at 48 hours. Both of the JAK2 chemical inhibitors decreased the proliferation of HL and MLBCL lines with similar EC50s. In addition, there was an inverse correlation between JAK2 (9p24.1) copy number and the EC50 of both JAK2 inhibitors in HL cells (Table 1).
EC50 of JAK2 inhibitors in HL and MLBCL cell lines
| . | JAK2 copy number . | SD-1029 . | Z3 . | ||
|---|---|---|---|---|---|
| EC50 . | 95% CI . | EC50 . | 95% CI . | ||
| L428 | 2.46 | 10.7 | 10.4-10.9 | 10.1 | 9.9-10.3 |
| L540 | 2.84 | 4.7 | 4.5-4.9 | 5.2 | 5.1-5.3 |
| KMH2 | 2.84 | 4.7 | 4.6-4.9 | 4.9 | 4.6-5.1 |
| L1236 | 3.60 | 4.3 | 3.9-4.7 | 2.5 | 2.1-2.8 |
| SUPHD1 | 4.83 | 3.1 | 3.0-3.3 | 1.7 | 1.4-1.9 |
| HDLM2 | 7.36 | 3.0 | 2.9-3.0 | 2.7 | 2.3-3.1 |
| Karpas-1106P | 3.82 | 1.7 | 1.6-1.7 | 3.2 | 2.9-3.5 |
| . | JAK2 copy number . | SD-1029 . | Z3 . | ||
|---|---|---|---|---|---|
| EC50 . | 95% CI . | EC50 . | 95% CI . | ||
| L428 | 2.46 | 10.7 | 10.4-10.9 | 10.1 | 9.9-10.3 |
| L540 | 2.84 | 4.7 | 4.5-4.9 | 5.2 | 5.1-5.3 |
| KMH2 | 2.84 | 4.7 | 4.6-4.9 | 4.9 | 4.6-5.1 |
| L1236 | 3.60 | 4.3 | 3.9-4.7 | 2.5 | 2.1-2.8 |
| SUPHD1 | 4.83 | 3.1 | 3.0-3.3 | 1.7 | 1.4-1.9 |
| HDLM2 | 7.36 | 3.0 | 2.9-3.0 | 2.7 | 2.3-3.1 |
| Karpas-1106P | 3.82 | 1.7 | 1.6-1.7 | 3.2 | 2.9-3.5 |
Values are reported as the median and 95% confidence interval of the concentration (μM) of inhibitor required to elicit a 50% reduction in cellular proliferation/respiration.
Discussion
By integrating high-resolution copy number data with transcriptional profiles, we identified the immunoregulatory genes PD-L1 and PD-L2 as key targets of the 9p24.1 amplification in HL and MLBCL cell lines. We also extended these findings to primary tumors and found that PD-1 ligand/9p24.1 amplification was restricted to the HL subtype most closely related to primary MLBCL, NSHL. Using quantitative immunohistochemical methods, we demonstrated that PD-1 ligand gene amplification was associated with increased protein expression in primary tumors. In HLs and MLBCLs, the extended 9p24.1 amplification region included the JAK2 locus. JAK2 amplification increased JAK2 protein expression and activity, specifically induced PD-1 ligand transcription, and enhanced sensitivity to JAK2 inhibition. These findings define 9p24.1 amplification as a disease-specific structural alteration that increases both the gene dosage of PD-1 ligands and their induction by JAK2 (Figure 7).
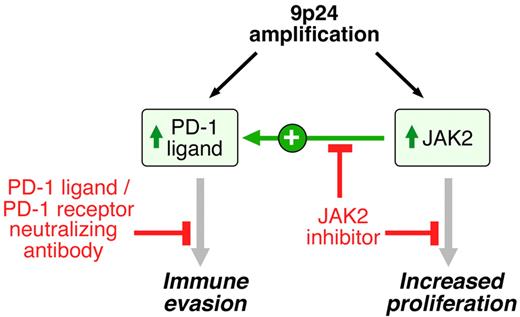
9p24.1 amplification targets, consequences, and associated treatment options. 9p24.1 amplification increases PD-1 ligand (PD-L1 and PD-L2) and JAK2 copy numbers, augments JAK2/STAT1 activity, induces PD-1 ligand expression, and stimulates HL proliferation.
9p24.1 amplification targets, consequences, and associated treatment options. 9p24.1 amplification increases PD-1 ligand (PD-L1 and PD-L2) and JAK2 copy numbers, augments JAK2/STAT1 activity, induces PD-1 ligand expression, and stimulates HL proliferation.
With high-resolution HD SNP array data and GISTIC analysis, it was possible to define the boundaries of the 9p24.1 amplification (13 Mbp, 799 genes) and further delineate the amplification peak (177 Kbp, 7 genes). By integrating these data with concurrent transcriptional profiles, we identified the genes with the closest association between transcript abundance and copy number. With this approach, we built on the prior observations of 9p gain in HL and MLBCL, localized the PD-1 ligand genes to the 9p24.1 amplification peak, and confirmed the highly significant association between PD-L1 and PD-L2 copy number and transcript abundance.
Recent studies highlight the emerging role of PD-1 ligand/receptor-mediated immune escape in cancer.8 Although PD-1 ligand expression is associated with adverse outcomes in multiple solid tumors, the genetic mechanisms for PD-1 ligand overexpression or induction or both are largely undefined. In previous in vitro analyses of multiple myeloma or lung cancer cell lines, IFNγ treatment induced PD-1 ligand expression.34,35 More recently, loss of the phosphatase and tensin homolog tumor suppressor (PTEN) was associated with increased PD-L1 expression in malignant gliomas36 but not in other solid tumors.35,37 In additional studies of anaplastic large cell lymphomas, the chimeric fusion protein, nucleophosmin/anaplastic lymphoma kinase, was reported to induce PD-L1 expression by a STAT3-dependent mechanism.37
Herein, we demonstrate that structural amplification of chromosome 9p24.1 increases the abundance of both PD-1 ligands and their inducer, JAK2, in the related diseases of NSHL and MLBCL. In HL cell lines with 9p24.1 amplification, cell-surface PD-1 ligand expression was greater than would be predicted by a simple gene-dosage effect. Furthermore, PD-L1 promoter-driven luciferase activity was significantly higher than predicted by gene copy number alone in HL cell lines. Consistent with these findings, JAK2 signaling further augmented PD-1 ligand expression in cell lines with 9p24.1 amplification. In addition, JAK2 inhibition had a greater effect on PD-L1 promoter–driven luciferase activity in the HLs with high-level 9p24.1 amplification. There was a similar, although less striking, association between 9p24.1 copy number and PD-L2 transcript abundance in HL.
The relative differences in JAK2/STAT induction of PD-L1 and PD-L2 may be due, in part, to features of the genes' 5′ regulatory regions, including spacing of the ISRE/IRF1 module and numbers of candidate STAT binding sites. Of note, PD-L1 was more abundant than PD-L2 in HL cell lines with 9p24.1 amplification; in contrast, MLBCLs with 9p24.1 had greater induction of PD-L2 than PD-L1. Previous studies highlight subtle differences in the expression, regulation, and potential function of the 2 PD-1 ligands.10,38,39
Given the association between PD-1 ligand expression and outcome in other cancers, we speculate that 9p24.1 amplification may be an important genetic prognostic factor in HL and MLBCL. Furthermore, the basal and amplification-associated PD-1 ligand expression in NSHL and MLBCLs suggests that these tumors may be particularly responsive to PD-1 ligand/PD-1 receptor blockade (Figure 7). In recent studies, tumor-infiltrating T cells from primary HLs expressed PD-1 and responded to PD-1 blockade in vitro.7,40,41 Because both PD-L1 and PD-L2 engage the PD-1 receptor, PD-1 receptor blockade may represent a promising therapeutic strategy in HL and, possibly, MLBCL. Of note, humanized PD-1 receptor-blocking monoclonal antibodies augment antitumor activities of effector T cells and natural killer cells and increase anticancer immune responses in murine tumor models.42 In addition, PD-1–neutralizing antibodies appear to be well tolerated in initial phase 1 clinical trials.42
Given the copy number–dependent JAK2 activity in HLs, JAK2 may be another rational therapeutic target alone or in combination with PD-1 blockade (Figure 7). In our in vitro assays that used the commercially available JAK2 inhibitors, SD1029 and Z3,30,31 there was an excellent correlation between the doses required to inhibit phosphoJAK2, decrease PD-L1 transcription, and reduce the proliferation of HL cell lines. Of note, these SD1029 and Z3 doses were comparable with those required to inhibit phosphoJAK2 and cellular proliferation in hematologic malignancies with activating JAK2 mutations (JAK2-V617F or TEL-JAK2).30,31 These observations suggest that additional clinical evaluation of more potent clinical-grade JAK2 inhibitors43 is warranted in HL and MLBCL. Consistent with the probable importance of JAK2 amplification in HL and MLBCL, additional genetic mechanisms, including SOCS-1 mutation and miR135a loss also increase JAK2 activity.44-47
The current studies, which highlight the role of 9p24.1-driven PD-1 ligand expression in HL and MLBCL, add to emerging data about complementary mechanisms of tumor immune privilege in these diseases.6 In addition to the PD-1 ligand, cHL RS cells also produce the immunoregulatory protein, galectin-1, and the immunosuppressive cytokine, interleukin-10 (IL-10).6,48 IL-10 suppresses the activation of naive T cells, and PD-L1 inhibits the TCR signaling of activated T cells.48 Furthermore, IL-10 and PD-L1 promote the differentiation of immunosuppressive T-regulatory cells by different mechanisms,49 and galectin-1 supports the expansion of T-regulatory cells and the selective deletion of Th1 and Th17 cells.50 These mechanisms of immune escape probably act in concert within the tumor microenvironment to foster immune privilege.
In summary, our current studies define a disease-specific genetic abnormality, 9p24.1 amplification, with probable prognostic significance and associated complementary rational therapeutic targets, PD-1 ligand and JAK2 (Figure 7). Furthermore, we describe a quantitative method for determining PD-1 ligand expression in primary tumors, an important adjunct to personalized approaches to therapy. These data support further evaluation of PD-1 blockade and JAK2 inhibition, alone and in combination, in patients with HL and MLBCL characterized for 9p24.1 and the associated targets.
The online version of this article contains a data supplement.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.
Acknowledgments
This work was supported by funding from the National Institutes of Health (PO1 CA092625) and the Miller Family Research Fund.
National Institutes of Health
Authorship
Contribution: M.R.G. designed research, performed research, analyzed data, and wrote the paper; S.M., S.J.R., and P.J. performed research and analyzed data; T.C., E.O., and K.T. performed research; B.C., D.N., and T.R.G. analyzed data; J.L.K. designed research and analyzed data; and M.A.S. designed research, analyzed data, and wrote the paper.
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Correspondence: Margaret A. Shipp, Dana-Farber Cancer Institute, 44 Binney St, Boston, MA 02115; e-mail: margaret_shipp@dfci.harvard.edu.



This feature is available to Subscribers Only
Sign In or Create an Account Close Modal