Abstract
Despite the high cure rate of pediatric acute lymphoblastic leukemia (ALL), relapse and associated therapy resistance remains a significant problem. Recent studies have identified genomic lesions that predict poor outcome (for example: loss of IKZF1 or P53). Because the effects on cell biology of most of these alterations are unknown, rational design of alternative therapy protocols is difficult.
We used a CRISPR/Cas9 based knockout screen in the (cytogenetically normal) B cell progenitor-ALL cell line Nalm6 to identify novel genes involved in resistance towards asparaginase, a key component of current ALL treatment protocols. As a proof-of-principle experiment, we introduced loss-of-function mutations by transiently expressing the Cas9 nuclease in Nalm6 cells transduced with a guideRNA (gRNA) library that targets 507 of the human kinases, each with 10 unique gRNAs (Wang et al. Science 2014;343 (6166):80-4). We treated these cells in the absence or presence of an IC50 dose of asparaginase for two weeks. Subsequently, genomic DNA from treated and untreated control cells was isolated and subjected to the Illumina HiSeq platform for paired-end sequencing. We retrieved 84% of all gRNA sequences with a median of 997 reads per gRNA in the sample before treatment. Furthermore, 454 out of the 507 genes were targeted by 6 or more gRNAs. This indicates that the library complexity was sufficiently maintained during the transduction and culture procedure to study dynamics during treatment.
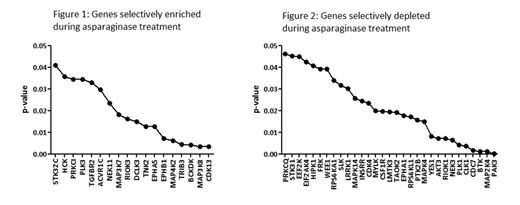
The frequency of each gRNA before and after treatment was compared to determine the effect of loss of function of individual kinases on sensitivity towards asparaginase treatment. The MAGeCK algorithm (Li et al. Genome Biology 2014, 15:554) was used to the prioritize genes of which the gRNAs were selectively enriched or depleted during treatment. This algorithm ranks genes by comparing the performance of each gRNA that targets a specific gene. This analysis yielded 18 genes that were found to be associated with resistance (figure 1), as illustrated by the enrichment of multiple gRNAs targeting these genes. To the converse, 31 (out of 454 evaluable) genes were selectively depleted during treatment (figure 2), suggesting that loss of these genes enhances sensitivity to asparaginase treatment. These gRNAs frequently target pro-survival kinases, including components of B cell receptor signaling (BTK, MAPK, AKT3, and Yes1), suggesting that inhibiting these kinases may be used to enhance treatment response.
The apoptosis inducing effects of Asparaginase treatment impinge on changes in cell metabolism as a result of amino acid starvation. In line with this, our dataset revealed enrichment of gRNAs targeting genes either directly involved in the amino acid response route (TRIB3) or other metabolic pathways (BCKDK). For an initial validation step, we used RNAi to suppress expression of one of these genes, Tribbles homologue 3 (TRIB3), a pseudokinase acting as a pro-apoptotic protein in the amino acid response pathway. Indeed, shRNA mediated knockdown of TRIB3 was sufficient to render these cells refractory to the apoptosis inducing effects of asparaginase.
We conclude from these results that our CRISPR/Cas9 based screens can be used to (i) delineate pathways that contribute therapy resistance and (ii) identify protein (kinase) targets that can be selectively inhibited to improve therapy response.
No relevant conflicts of interest to declare.
Author notes
Asterisk with author names denotes non-ASH members.