Abstract
Abstract 2608
HUCB has been described to contain hematopoietic multi-lineage progenitor cells that contribute to the success in treating malignant and non-malignant diseases (Cairo et al, Blood, 1997). We demonstrated that multi-lineage progenitor cells derived from HUCB can differentiate into cells representative of the 3 germ layers in vitro (Vandeven/Cairo et al, Exp Hem, 2007). Zaehres et al (Exp Hem, 2010) have described another population of primitive stem cells, USSCs, as a new potential source to generate iPS cells following retroviral transduction. This suggests that USSCs are a strong candidate as a source for developing patient specific or HLA matched iPSC banks. Since using retroviral iPS cells are challenged with the possibility of oncogene reactivation; gene integration free methods of generating iPS cells such as small molecule treatment to activate ES transcription factor genes are needed. Furthermore, the epigenetic modification of the USSCs at the key ES transcription factors has not been described and this information will provide insights on the differentiation potential of USSCs and their capacity to reprogramming.
To determine the DNA methylation patterns in the core regulatory regions, including both enhancer and promoter of ES transcription factors Oct4 and Nanog in USSCs prior to and following gene transcription effects of DNMT1 inhibition by treatment with 5-azacytidine.
USSCs were derived from HUCB mononuclear cells in 30% FBS and 10-7M dexamethasone (Kogler, J Exp Med, 2004). The USSC population was confirmed by flow cytometry and their fibroblastic morphology. The cells were passaged in the same medium without dexamethasone. Bisulfite sequencing was performed to characterize the CpG island methylation in the regulatory regions of Oct4 (from 2563 bp upstream to 250 bp downstream of Oct4 transcription start site) and Nanog (from 1449 bp to 82 bp upstream of the Nanog transcription start site). RT-PCR and qPCR were conducted to determine the expression levels of Nanog, Oct4 and Sox2, using isoform specific and intron spanning primers. 3mM 5-azacytidine was used to treat USSCs for demethylation studies and the RNA was collected 24 hours following treatment. The results were compared to human embryonic stem cells and human foreskin fibroblasts as positive and negative controls, respectively.
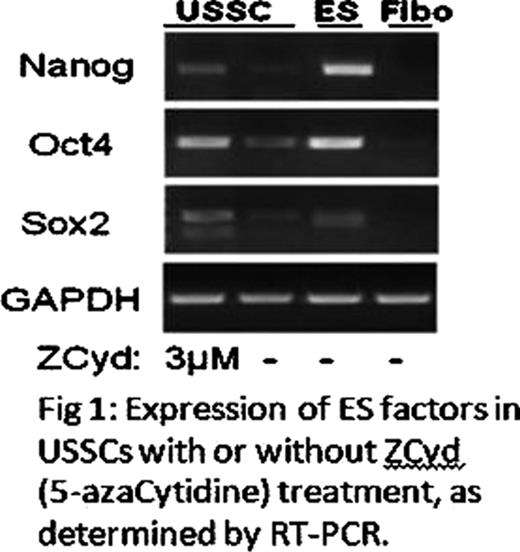
USSCs were derived from 50% (n=25) of HUCB; with 1–10 colonies from each successfully derived cord blood. Flow cytometry indicated that USSCs were lineage negative and express overlapping but distinct cell surface markers with MSCs; positive for CD73, CD90, CD146, CD50, but negative for CD106. Bisulfite sequencing of USSCs demonstrated a mosaic methylation pattern of CpG islands at the regulatory sites of both Oct4 and Nanog. An average of 65% and 47% of the CpGs were unmethylated in the enhancer and promoter regions of Nanog, respectively. 56% were unmethylated at the enhancer of Oct4 while the promoter was heavily methylated, except for the 400 bp region that spans the Oct4 transcription start site, which was 80% unmethylated. This is consistent with the RT-PCR results showing a low but consistent level of Nanog, Oct4 and Sox2 (Figure 1). Based on q-PCR using isoform-specific and intro-spanning primers, we determined that USSCs have about 20-and 400- fold higher levels of Nanog and Oct4 expression, respectively,compared to fibroblasts. Following a 24hr exposure to a DNMT1 inhibitor, 5-azacytidine, to the USSCs, there was a 10-fold increase in the mRNA expression of Oct4, Nanog, and Sox2.
HUCB derived USSCs have a mosaic pattern of CpG island methylations in the distal and proximal regions of key ES transcription factors, Oct4 and Nanog. This is supported by a consistent low expression level of these genes. The mosaic pattern of CpG islands seems to be more malleable by small molecule perturbation; 3mM of 5-azacytidine appeared to significantly increase the Oct4 and Nanog expression. We hypothesize that due to their semi-permissive chromatin structure at the core regulatory regions in key ES transcription factors, HUCB derived USSCs are likely to be a more optimal choice of small molecule derived induced pluripotent stem cells compared to other cell types.
No relevant conflicts of interest to declare.
Author notes
Asterisk with author names denotes non-ASH members.