Abstract
Anaplastic large cell lymphomas (ALCL) are frequently associated with the t(2;5)(p23;q35). This translocation fuses the nucleophosmin (NPM) gene at 5q35, which encodes a nucleolar protein involved in shuttling ribonucleoproteins from the cytoplasm to the nucleus, to the anaplastic lymphoma kinase (ALK) gene at 2p23, encoding a tyrosine kinase receptor. In this report, we describe a typical case of ALCL whose malignant cells exhibited a novel (1;2)(q25;p23) translocation. These cells expressed ALK protein, but, in contrast to t(2;5)-positive ALCL (which show cytoplasmic, nuclear, and nucleolar staining), labeling was restricted to the malignant cell cytoplasm. Using a polymerase chain reaction (PCR)-based technique to walk on chromosome 2 from the known ALK gene across the breakpoint, we showed that the gene involved at 1q25 is TPM3, encoding a nonmuscular tropomyosin. We subsequently identified, using reverse transcription-PCR analysis of cases showing similar ALK cytoplasm-restricted staining, fusion of the ALK andTPM3 genes in 2 other cases of ALCL. The TPM3 gene has been previously found in papillary thyroid carcinomas as a fusion partner with the TRK kinase gene. We showed that TPM3 is constitutively expressed in lymphoid cell lines, suggesting that, in these t(1;2)-bearing ALCL cases, the TPM3 gene contributes an active promoter for ALK expression. Activation of the ALK catalytic domain probably results from homodimerization of the hybrid protein TPM3-ALK, through the TPM3 protein-protein interaction domain. The present cases of ALCL associated with a novel t(1;2)(q25;p23) demonstrate that at least one fusion partner other than NPM can activate the intracytoplasmic domain of the ALK kinase.
ANAPLASTIC large cell lymphoma (ALCL) is associated with a recurrent translocation, the t(2;5)(p23;q35).1-7 This translocation involves theALK (anaplastic lymphoma kinase) gene at 2p23 and theNPM (nucleophosmin) gene at 5q35. The ALK gene encodes a tyrosine kinase receptor belonging to the insulin growth factor receptor superfamily,8 which is normally expressed in nerve cells but silent in normal lymphoid cells.9 10
The NPM gene is a housekeeping gene encoding a nucleolar phosphoprotein involved in shuttling ribonucleoproteins from the cytoplasm to the nucleus.11,12 It is postulated thatNPM gene promotes the expression of the ALK catalytic domain present in the chimeric NPM-ALK protein, and it has recently been demonstrated that constitutive ALK activity contributes to the malignant transformation of lymphoid cells.13 14
Since the first report of the t(2;5) in ALCL,15 cytogenetic or molecular variants implicating the ALK gene have been described.16-18 The recent report of a complex t(2;5)(q37;q31), which brings the ALK gene into the vicinity of the NPM gene, and of an inv(2)(p23q35), both of which are associated with the expression of the ALK intracytoplasmic domain, suggests that genes other than NPM can promote ALK gene expression.19
In a previous study,5,20 we described a case of ALCL (case GEL) with typical morphologic and phenotypic features whose malignant cells did not show the (2;5) translocation but instead a variant translocation involving a breakpoint at q25 on chromosome 1 and at p23 on chromosome 2. The malignant cells expressed ALK protein strongly, as detected with polyclonal (p80)5,21 and monoclonal (ALK1) antibodies.20 Because both of these antibodies react with the tyrosine kinase domain of the ALK protein, we postulated that a gene located at 1q25 contributed an active promoter for the expression of the ALK catalytic domain in the malignant cells in this case.
To identify the gene on chromosome 1q25 involved in the above-mentioned case, we used a polymerase chain reaction (PCR)-based technique described by Siebert et al22 to walk on chromosome 2 from the ALK gene across the breakpoint to the unknown gene on chromosome 1. This analysis showed that ALK expression probably results from a chromosomal rearrangement fusing the last exons of theALK gene to sequences of the TPM3 gene, encoding nonmuscular tropomyosin.23,24 The latter gene has been reported as fusing to the TRK kinase gene in human thyroid papillary carcinomas.25 Subsequently, we identified 3 other cases of ALCL, on which no cytogenetic data were available, but which also showed ALK staining restricted to the cytoplasm. When investigated by reverse transcription-PCR (RT-PCR), 2 of these cases were also positive for TPM3-ALK transcripts.
MATERIALS AND METHODS
Material
The biopsy from patient GEL, which had been fixed in ethanol-based Bouin’s fluid (Duboscq-Brasil), showed the typical morphologic and immunohistochemical features of common-type ALCL.6,26 However, cytogenetic analysis showed that malignant cells carried a (1;2)(q25;p23) translocation. No frozen material was available, but some of the lymph node biopsy specimen had been fixed and embedded using the ModAMeX method.27
DNA from the JAR cell line (a human choriocarcinoma; ATCC HTB 144) and the CNE2 tumor (a human nasopharyngeal carcinoma28) were used, respectively, for walking steps on the chromosome 1 beyond the breakpoint and for Southern experiments designed to test the PCR fragments obtained from the case GEL.
Three other ALCL cases, which were negative for NPM-ALKtranscripts and which showed ALK staining restricted to the cytoplasm similar to that observed in the case GEL, were also investigated. One case (case no. 1) was categorized as the lymphohistiocytic variant of ALCL,6,29 and the 2 other cases (cases no. 2 and 3) showed morphological and immunohistochemical features of common-type ALCL.6
The detection of TPM3 transcripts was performed by RT-PCR on mRNA from four lymphoid cell lines—CEM, Jurkat, HSB-2 (all acute T leukemia; ATCC CCL 119, ATCC TIB 153, and ATCC CCL 120.1, respectively) and SU-DHL-1 [established from a t(2;5)-positive anaplastic lymphoma; kindly provided by Dr M.L. Cleary, Stanford University Medical Center, Stanford, CA]—and also from thyroid frozen sections (used as positive control, because the TPM3 gene is constitutively active in this tissue25).
Immunostaining
Polyclonal anti-ALK (antibody p80)21 and monoclonal anti-ALK (antibody ALK1)20 were used, as previously described,5,20 diluted 1/100 and as undiluted culture supernatant, respectively. All cases were also immunostained with a panel of monoclonal antibodies detecting B and T antigens and for CD30 (Dako-Ber-H2, Copenhagen, Denmark), for EMA (Dako-EMA/E29), and with antibody BNH9,30 which is known to react with ALCL.
Cytogenetics
Karyotyping was performed on cells from the diagnostic lymph node from patient GEL after 24 hours of culture. Reverse-Heating-Giemsa (RHG) banding was performed using hot phosphate buffer, and chromosomal abnormalities were described according to the ISCN 95 nomenclature.30a
DNA Extraction and Digestion
DNA from case GEL was extracted from 10 deparaffinized ModAMeX-processed sections (10-μm thick) by proteinase K digestion, phenol-chloroform extraction, and ethanol precipitation.31A small quantity (70 μg) of genomic DNA could be extracted from this material, but this DNA was of poor quality, with a maximum size of 6.5 kb when analyzed by agarose gel electrophoresis, precluding the construction and the screening of any genomic library for cloning the breakpoint.
DNA from the CNE2 tumor and the JAR cell line was extracted after solubilization of cell pellets in 5 mol/L guanidium isothiocyanate (GITC), 50 mmol/L Tris, pH 8, and 1% mercaptoethanol by gentle agitation at room temperature. The lysate was then dialysed three times using 100 vol of 10 mmol/L Tris, pH 8, 1 mmol/L EDTA, and 10 mmol/L NaCl (buffer A) before being treated at 37°C successively with 100 μg/mL RNase (Sigma-Aldrich Chimie, L’Isle d’Abeau Chesnes, France) for 2 hours and with 100 μg/mL proteinase K (Merck, Chelles, France) for 2 to 4 hours. The solution was then extracted several times with equal volumes of phenol and chloroform-isoamyl alcohol (24:1) and again dialysed against buffer A.
Aliquots of DNA (5 μg) from GEL, CNE 2, and JAR were digested withDra I, EcoRV, Pvu II, Sca I, andSsp I enzymes (New England Biolabs, Montigny-Le-Bretonneux, France), following the manufacturer’s recommendations.
PCR-Based Technique for Walking on Genomic DNA
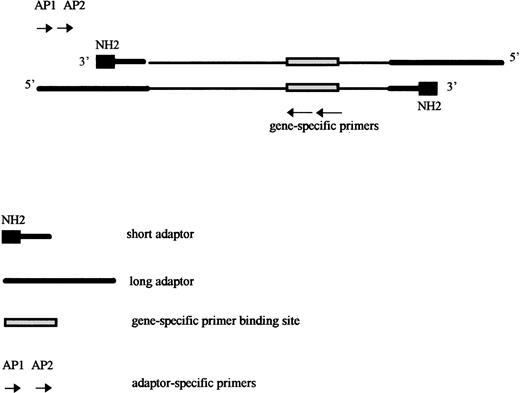
This technique, described by Siebert et al,22 involves a PCR between a gene-specific primer and a primer annealing to an adaptor sequence previously ligated to the ends of the genomic DNA fragments. The adaptor is designed in such a way that its specific primer cannot anneal to the native adaptor, because the target sequence is only created by the elongation of the gene-specific primer after the first PCR cycle.22 Nonspecific amplifications between adaptors are thus avoided and specific amplification occurs despite the use of only a single gene-specific primer (Fig 1). To increase sensitivity and specificity, the first amplification is followed by a second nested amplification.
Long-range amplification on digested and adaptor-ligated DNA. Genomic DNA is digested with restriction enzymes and the blunt ends of DNA fragments are ligated to an excess of adaptors, made of two complementary oligonucleotides, one long and one short. This technique consists of an amplification between a gene-specific primer and a primer annealing in the adaptor sequence previously ligated to the ends of genomic DNA fragments. The adaptor is designed in such a way that its specific primer cannot anneal on the native adaptor, and the target sequence for the adaptor specific primer (AP1) is therefore only created by elongation from the gene-specific primer. The 3′ extremity of the short oligonucleotide is blocked by an amine group, avoiding its elongation by the DNA polymerase. To increase sensitivity and specificity, a nested amplification is performed using a nested adaptor specific primer (AP2) and a nested gene-specific primer.
Long-range amplification on digested and adaptor-ligated DNA. Genomic DNA is digested with restriction enzymes and the blunt ends of DNA fragments are ligated to an excess of adaptors, made of two complementary oligonucleotides, one long and one short. This technique consists of an amplification between a gene-specific primer and a primer annealing in the adaptor sequence previously ligated to the ends of genomic DNA fragments. The adaptor is designed in such a way that its specific primer cannot anneal on the native adaptor, and the target sequence for the adaptor specific primer (AP1) is therefore only created by elongation from the gene-specific primer. The 3′ extremity of the short oligonucleotide is blocked by an amine group, avoiding its elongation by the DNA polymerase. To increase sensitivity and specificity, a nested amplification is performed using a nested adaptor specific primer (AP2) and a nested gene-specific primer.
Adaptor ligation.
The blunt ends of DNA fragments generated by digestion with DraI, EcoRV, Pvu II, Sca I, and Ssp I enzymes were ligated to an excess of adaptors, using T4 DNA ligase (New England Biolabs), following the manufacturer’s recommendations. The adaptor was made of two complementary oligonucleotides, one long and one short (Fig 1). Once the two complementary oligonucleotides annealed, the adaptor presented with one blunt end that was able to ligate to any blunt end fragment of genomic DNA. The 3′ extremity of the short oligonucleotide was blocked by an amine group, preventing its elongation by DNA polymerase (Fig 1).
DNA amplification.
Long-range PCR was performed with the Expand Long Template PCR system (Boehringer Mannheim, France SA, Meylan, France) using a unique enzyme mix containing thermostable Taq and Pwo DNA polymerases. This was performed using GEL DNA fragments for the three first steps and JAR DNA fragments for the fourth (because of the limited amount of DNA from case GEL; Fig 2). Primary PCR reactions were conducted in 50 μL volumes containing 0.5 μg of ligated DNA, 0.3 μmol/L adaptor primer AP122 and ALKgene-specific primer, 1.75 mmol/L MgCl2, 0.35 mmol/L dNTP, and 6.125 U of DNA polymerase. An initial denaturation step at 94°C for 2 minutes was followed by 30 cycles of denaturation at 94°C for 30 seconds, annealing-extension at 68°C for 5 minutes, and a final extension time of 7 minutes at 65°C. The second (nested) PCR reaction was performed under the same conditions on 1 μL of the primary PCR, using adaptor primer AP222 and a nestedALK gene-specific primer. The adaptor primers AP1 and AP2 and the different ALK gene-specific primers are listed in Table 1.
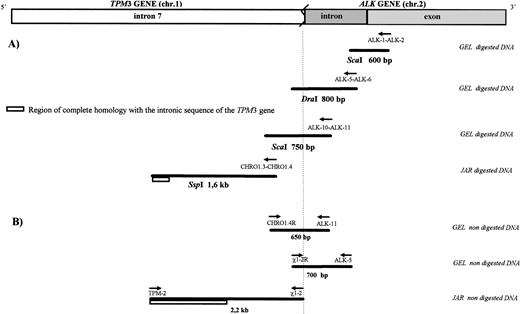
Long-range amplification on digested and nondigested DNA. (A) The first three steps were performed using a PCR-based technique after adaptor ligation to digested DNA, with an adaptor primer and anALK gene-specific primer, to determine the chromosome 1 gene involved in the (1;2) translocation. By walking on the chromosome from the ALK gene across the breakpoint to this unknown gene, we demonstrated that this chromosomal rearrangement fused the ALKgene on chromosome 2p23 to the TPM3 gene on chromosome 1q25. The 1.6-kb Ssp I fragment was only partially sequenced to yield the first 170 bp of this fragment at the 5′ end, allowing preparation of primers for the next step. This 170-bp sequence showed a complete identity to the sequence of intron 7 of the TPM3 gene. (B) The next three PCR steps were performed on undigested DNA and confirmed that all PCR fragments previously generated were colinear. The longer 2.2-kb PCR fragment obtained on undigested DNA was entirely sequenced and showed 99.9% identity with a 1,146-bp sequence in part of intron 7 of the TPM3 gene deposited in the GenBank database25 (accession no. X79910).
Long-range amplification on digested and nondigested DNA. (A) The first three steps were performed using a PCR-based technique after adaptor ligation to digested DNA, with an adaptor primer and anALK gene-specific primer, to determine the chromosome 1 gene involved in the (1;2) translocation. By walking on the chromosome from the ALK gene across the breakpoint to this unknown gene, we demonstrated that this chromosomal rearrangement fused the ALKgene on chromosome 2p23 to the TPM3 gene on chromosome 1q25. The 1.6-kb Ssp I fragment was only partially sequenced to yield the first 170 bp of this fragment at the 5′ end, allowing preparation of primers for the next step. This 170-bp sequence showed a complete identity to the sequence of intron 7 of the TPM3 gene. (B) The next three PCR steps were performed on undigested DNA and confirmed that all PCR fragments previously generated were colinear. The longer 2.2-kb PCR fragment obtained on undigested DNA was entirely sequenced and showed 99.9% identity with a 1,146-bp sequence in part of intron 7 of the TPM3 gene deposited in the GenBank database25 (accession no. X79910).
Sequences of the Adaptors and the Different Primers Used
| Adaptors |
| AD-L-1: 5′ CTA ATA CGA CTC ACT ATA GGG CTC GAG CGG CCG CCC GGG CAG GT 3′ |
| AD-S-1-NH2: 5′ ACC TGC CC-NH2 3′ |
| Adaptor primers |
| AP-1: 5′ CTA ATA CGA CTC ACT ATA GGG CTC GAG 3′ |
| AP-2: 5′ ATA GGG CTC GAG CGG CCG CCC 3′ |
| ALK andTPM3 gene-specific primers used on genomic DNA |
| ALK-1: 5′ GCC AGC AAA GCA GTA GTT GGG GTT G 3′ |
| ALK-2: 5′ GTC GAG GTG CGG AGC TTG CTC AGC 3′ |
| ALK-5: 5′ GAA ATG GGG AGG GAG CCA GGG AAG G 3′ |
| ALK-6: 5′ GGC TGG GTG AAC CAG CAG ACT GTG TT 3′ |
| ALK-10: 5′ GTA GAT TCT GTG TGT AAA GCC CAG CCC 3′ |
| ALK-11: 5′ TTG AAA TGT GAA ATT GCC GAG C 3′ |
| χ 1-2: 5′ GCC AAC TTC TGC CCA GGA TCT GAA TA 3′ |
| χ 1-2R: 5′ TAT TCA GAT CCT GGG CAG AAG TTG GC 3′ |
| CHRO-1-3: 5′ CCA CTA GAA AGC ATC CAA CCC GGA AC 3′ |
| CHRO-1-4: 5′ TCA ACC GTC TTC TTT CCC TCC AAT CG 3′ |
| CHRO-1-4R: 5′ CGA TTG GAG GGA AAG AAG ACG GTT GA 3′ |
| TPM 2: 5′ GCT TCT CGA GTT CAA GCG ATT CTC CTG T 3′ |
| Tropomyosin-specific primers used for RT-PCR to detect TPM3 and TPM3-ALKtranscripts |
| CTPM-1: 5′ CGA GAA GTT GAG GGA GAA AGG 3′ |
| CTPM-2: 5′ CAA CTT ACG AGC CAC CTC TTC 3′ |
| CTPM-3: 5′ CTG GCA GAG TCC CGT TGC C 3′ |
| CTPM-4: 5′ ACT GGG CGT TCT ACA TCT CAT T 3′ |
| Adaptors |
| AD-L-1: 5′ CTA ATA CGA CTC ACT ATA GGG CTC GAG CGG CCG CCC GGG CAG GT 3′ |
| AD-S-1-NH2: 5′ ACC TGC CC-NH2 3′ |
| Adaptor primers |
| AP-1: 5′ CTA ATA CGA CTC ACT ATA GGG CTC GAG 3′ |
| AP-2: 5′ ATA GGG CTC GAG CGG CCG CCC 3′ |
| ALK andTPM3 gene-specific primers used on genomic DNA |
| ALK-1: 5′ GCC AGC AAA GCA GTA GTT GGG GTT G 3′ |
| ALK-2: 5′ GTC GAG GTG CGG AGC TTG CTC AGC 3′ |
| ALK-5: 5′ GAA ATG GGG AGG GAG CCA GGG AAG G 3′ |
| ALK-6: 5′ GGC TGG GTG AAC CAG CAG ACT GTG TT 3′ |
| ALK-10: 5′ GTA GAT TCT GTG TGT AAA GCC CAG CCC 3′ |
| ALK-11: 5′ TTG AAA TGT GAA ATT GCC GAG C 3′ |
| χ 1-2: 5′ GCC AAC TTC TGC CCA GGA TCT GAA TA 3′ |
| χ 1-2R: 5′ TAT TCA GAT CCT GGG CAG AAG TTG GC 3′ |
| CHRO-1-3: 5′ CCA CTA GAA AGC ATC CAA CCC GGA AC 3′ |
| CHRO-1-4: 5′ TCA ACC GTC TTC TTT CCC TCC AAT CG 3′ |
| CHRO-1-4R: 5′ CGA TTG GAG GGA AAG AAG ACG GTT GA 3′ |
| TPM 2: 5′ GCT TCT CGA GTT CAA GCG ATT CTC CTG T 3′ |
| Tropomyosin-specific primers used for RT-PCR to detect TPM3 and TPM3-ALKtranscripts |
| CTPM-1: 5′ CGA GAA GTT GAG GGA GAA AGG 3′ |
| CTPM-2: 5′ CAA CTT ACG AGC CAC CTC TTC 3′ |
| CTPM-3: 5′ CTG GCA GAG TCC CGT TGC C 3′ |
| CTPM-4: 5′ ACT GGG CGT TCT ACA TCT CAT T 3′ |
Identification of the ALK Gene Breakpoint
To determine whether a given PCR fragment contained the breakpoint of interest, the fragment was purified, labeled, and used as a probe in Southern blot experiments on an unrelated DNA (CNE2) digested with the same panel of restriction enzymes: Dra I, EcoRV,Pvu II, Sca I, and Ssp I. We considered that a PCR fragment generated from the GEL DNA (following digestion with a single specific enzyme) that annealed with two different fragments of the unrelated CNE2 DNA digested with the same enzyme probably contained the ALK gene breakpoint.
After digestion of the CNE2 DNA by the same restriction enzymes (Dra I, EcoRV, Pvu II, Sca I, andSsp I), 5 μg of each digested DNA was loaded on a 0.7% agarose gel, electrophoresed for 16 hours at 1.5 V/cm in TBE, and transferred onto nylon membrane (Genscreen Plus; Dupont, Boston, MA) using 0.4 mol/L NaOH as transfer solution. The membrane was then rinsed, dried, and prehybridized in 10% dextran sulfate, 1 mol/L NaCl, and 1% sodium dodecyl sulfate (SDS) for 30 minutes at 65°C. Long-range PCR fragments were purified after agarose gel electrophoresis using the GeanClean III Kit (Bio 101, Ozyme, Saint Quentin Yvelines, France) and used as probes for the Southern blot experiments. Fragments (∼25 ng) were labeled with 25 μCi of α-32P-dCTP using the NEBlot kit (New England Biolabs). After the addition of the denatured labeled probe, filters were hybridized overnight at 65°C, washed according to the manufacturer’s recommendations, and exposed to Kodak X-Omat AR films (Eastman Kodak, Rochester, NY) using intensifying screens at −80°C.
Direct Cycle Sequencing of PCR Products
Long-range PCR fragments and RT-PCR fragments were sequenced using an original protocol32 with DeepVent(exo−) DNA polymerase (New England Biolabs) and α-35S-dATP. For each PCR fragment, 5′ sequences (located close to the adaptor) were first determined to prepare primers for the next PCR step.
Total RNA Extraction
Total RNA was extracted from frozen sections of normal thyroid and lymph nodes involved by ALCL (cases no. 1, 2, and 3) and from the Jurkat, CEM, HSB-2, and SU-DHL-1 cryopreserved cell pellets (10 × 106) using the RNeasy Midi Kit (Qiagen, Courtaboeuf, France).
Detection of Tropomyosin (TPM3) Transcripts by RT-PCR
Two TPM primers pairs, C-TPM1/C-TPM2 (positions 149-169/451-431) and C-TPM3/C-TPM4 (positions 497-515/809-788; see Table 1), were chosen according to the cDNA sequence of TPM3 published by MacLeod et al,33 yielding RT-PCR products of 303 and 313 bp, respectively. Synthesis of the first cDNA strand and PCR amplification were performed in a single tube with the RT-PCR Access Kit (Promega France, Charbonnières, France), following the manufacturer’s recommendations. The first step consisted of a gene-specific reverse transcription at 48°C for 45 minutes, followed by 30 cycles comprising a denaturation step at 94°C for 45 seconds, an annealing step at 58°C for 45 seconds, and an elongation step at 72°C for 35 seconds.
Detection of Hybrid TPM3-ALK Transcripts by RT-PCR
This study was performed in 3 ALCL cases that were negative forNPM-ALK transcripts and that showed ALK staining restricted to the cytoplasm, resembling that observed in the case GEL. The first round of RT-PCR was performed as described above, using C-TPM1 (position 149-169) and ALK-1 (position 4211-4187 on the cDNAALK sequence) primers. The second round was performed on a 1-μL aliquot from the first amplification, using nested primers C-TPM3 (position 497-515) and ALK-2 (position 4171-4148), yielding a RT-PCR product of 307 bp (see Table 1 for primers sequence).
RESULTS
Immunomorphologic Features
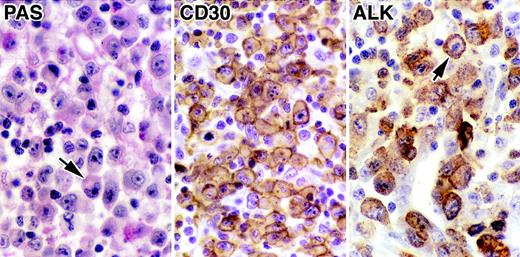
The tumor from patient GEL and the 3 other cases of ALCL investigated subsequently all contained large neoplastic cells with horseshoe or kidney-shaped eccentric nuclei (Fig 3). Their phenotype was also typical of systemic t(2;5)-positive ALCL in that they were of null (cases no. 1 and 2) or T phenotype (case no. 3) and coexpressed CD30 (Fig 3) and epithelial membrane antigen (EMA). Malignant cells were also positive with the antibody BNH.9. All cells stained for ALK protein as detected by p80 and ALK1 antibodies. However, in contrast to most ALCL, which show labeling of both the cytoplasm and the nucleus,14 20 ALK staining in these cases was limited to the cytoplasm (Fig 3).
Immunomorphologic features of the ALCL (GEL) carrying the (1;2) translocation. The morphology in a PAS-stained section is typical of the common-type of ALCL, and a hallmark cell,6 with an eccentric horseshoe-shaped nuclei, is arrowed. Staining for CD30 shows typical cytoplasmic and membrane-associated labeling of malignant cells. Labeling for ALK protein (antibody ALK1) is confined to the cell cytoplasm.
Immunomorphologic features of the ALCL (GEL) carrying the (1;2) translocation. The morphology in a PAS-stained section is typical of the common-type of ALCL, and a hallmark cell,6 with an eccentric horseshoe-shaped nuclei, is arrowed. Staining for CD30 shows typical cytoplasmic and membrane-associated labeling of malignant cells. Labeling for ALK protein (antibody ALK1) is confined to the cell cytoplasm.
Cytogenetics
Cytogenetic analysis of the diagnostic biopsy from patient GEL showed only abnormal metaphases, with the karyotype 46,XY,t(1;2)(q25;p23),der(22)t(1;22)(q12;p11)t(1;2)(q25;p23)[20].
Cloning of the t(1;2) Breakpoint
Long-range PCR was performed on DNA from the biopsy of patient GEL and from the JAR cell line, after digestion with different restriction enzymes and ligation to adaptors, as described in Materials and Methods. Each PCR step was realized with a couple of nested gene-specific primers and the nested adaptor-specific primers, AP1 and AP2. A PCR walk on chromosome 2 was performed using ALK-1 and ALK-2 gene-specific primers, located in the exon adjacent to the intron where the classical t(2;5) breakpoint occurs (Fig 2).
This first step yielded two fragments of approximately 600 bp on theSca I-digested GEL DNA. After sequencing, these fragments appeared to be closely related to and probably products of the two chromosomes 2. Nested ALK-5 and ALK-6 primers were chosen in a sequence common to both chromosomes at the 5′ extremity of these fragments and were used for a second PCR step with primers AP1 and AP2 (Fig 2). A new 800-bp fragment was obtained from Dra I-digested GEL DNA. After purification, this fragment was used in a Southern blot experiment to probe DNA from CNE2 digested with Dra I,EcoRV, Pvu II, Sca I, and Ssp I.
The GEL Dra I fragment showed two bands of differing intensity in the Dra I-digested CNE2 DNA, suggesting that it was composed of two different sequences that were not colinear in CNE2 DNA and, thus, likely contained the t(1;2) breakpoint. Comparison of its sequence with the sequence of the ALK intron (own unpublished results) confirmed that the 5′ last 80 bp of this fragment were not of ALK origin and putatively of chromosome 1 origin. However, this 80-bp sequence was not found to be homologous to any known sequence present in the GenBank database.
We performed a third PCR step using ALK-10 and ALK-11 gene-specific primers, chosen from the ALK intronic region of the 800-bpDra I fragment bearing the breakpoint (Fig 2). This amplification yielded a 750-bp fragment from Sca I-digested GEL DNA, but its sequence also showed no homology with sequences in the GenBank database.
Because no more digested and adaptor-ligated DNA from case GEL was available, we performed the next PCR step on DNA from the JAR cell line prepared under the same conditions using gene-specific primers (CHRO-1-3 and CHRO-1-4) chosen from the 5′ end of the previous fragment (Fig 2). This amplification yielded a large 1.6-kb fragment from the Ssp I-digested JAR DNA. The 170-bp sequence at the 5′ end of this fragment showed complete homology in the GenBank database with the intron 7 of the TPM3 (tropomyosin 3) gene.25
Because our PCR protocol was based on an adaptor-ligated DNA strategy, which can artefactually ligate unrelated fragments, we performed control PCRs (summarized in Fig 2) on undigested DNA from patient GEL using primers defined during our walking through this region. The first control PCR, between ALK-11 and CHRO1-4R, yielded a 650-bp fragment. Because CHRO-1-4R has the same sequence (but in an opposite orientation) as CHRO-1-4, which allowed the amplification of the last 1.6-kb fragment from JAR DNA, this was considered as the first definitive evidence that ALK-11 and the TPM3 gene were colinear in the DNA from case GEL.
The second control comprised another amplification across the breakpoint, between the ALK-5 and χ1-2R primers, with the latter primer being chosen from the chromosome 1-specific 80 bp of the GELDra I fragment. This amplification yielded the expected 700-bp fragment.
Finally, we performed a longer amplification on undigested JAR DNA between primers χ1-2 and TPM-2 chosen from the 5′ end of the 1.6-kb Ssp I fragment. A 2.2-kb fragment was obtained and a 1,146-bp sequence from this fragment was 99.9% identical to part of intron 7 from the TPM3 gene25 deposited in the Genbank database (Fig 2).
Because ALK-5 and χ1-2, on the one hand, and χ1-2 and TPM-2, on the other, were colinear in DNA from the case GEL, we concluded that ALK-5 and TPM-2 were colinear and that the (1;2)(q25;p23) translocation had juxtaposed the TPM3 gene on chromosome 1 to the ALKgene on chromosome 2 (the accession number of a 3,394-bp fragment of the TPM3-ALK genomic sequence submitted to GenBank is AF 112866). Five Alu sequences have been found in the TPM3intron, but none in the partial sequence of the ALK intron, and the breakpoint is located at the beginning of an Alu sequence in the TPM3 intron (data not shown).
Detection of Hybrid TPM3-ALK Transcripts and Direct Cycle Sequencing of RT-PCR Products
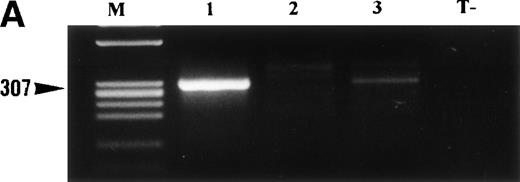
The 3 cases of ALCL that showed a cytoplasm-restricted staining pattern for ALK (from which no cytogenetic data were available) were analyzed for the presence of TPM3-ALK transcripts. A 307-bp band was found in cases no. 1 and 3 (Fig 4A), and sequencing of nested RT-PCR products confirmed the presence of hybridTPM3-ALK transcripts (Fig 4B).
(A) RT-PCR analysis of TPM3-ALK RNA in 3 cases of ALCL showing ALK-staining restricted to the cytoplasm. RNA obtained from the SU-DHL-1 cell line was used as negative control (T−). RT-PCR was performed using two rounds of PCR. The first round involved C-TPM1 and ALK-1 primers, and the second was a nested amplification using C-TPM3 and ALK-2 primers. PCR products were electrophoresed on 2% agarose gel and visualized with ethidium bromide staining. Two of these three cases of ALCL (cases no. 1 and 3) show a band of 307 bp, indicating the presence of hybrid TPM3-ALK RNA. M, size marker. (B) Nucleotide sequence of hybrid TPM3-ALK transcripts detected by nested RT-PCR using C-TPM3 and ALK2 primers. Nucleotides are numbered from the deoxyadenosine residue of the TPM3 initiator ATG codon.
(A) RT-PCR analysis of TPM3-ALK RNA in 3 cases of ALCL showing ALK-staining restricted to the cytoplasm. RNA obtained from the SU-DHL-1 cell line was used as negative control (T−). RT-PCR was performed using two rounds of PCR. The first round involved C-TPM1 and ALK-1 primers, and the second was a nested amplification using C-TPM3 and ALK-2 primers. PCR products were electrophoresed on 2% agarose gel and visualized with ethidium bromide staining. Two of these three cases of ALCL (cases no. 1 and 3) show a band of 307 bp, indicating the presence of hybrid TPM3-ALK RNA. M, size marker. (B) Nucleotide sequence of hybrid TPM3-ALK transcripts detected by nested RT-PCR using C-TPM3 and ALK2 primers. Nucleotides are numbered from the deoxyadenosine residue of the TPM3 initiator ATG codon.
Detection of Tropomyosin Transcripts
The tyrosine kinase domain of ALK is constitutively expressed in ALCL carrying the classical t(2;5), under control of the ubiquitously expressed NPM gene promoter. We therefore wished to know whether the TPM3 gene is also active in lymphoid cells. RT-PCR assays of RNA extracted from the Jurkat, CEM, HSB-2, and SU-DHL-1 T-cell lines (and also from thyroid tissue) demonstrated that theTPM3 gene was constitutionally active,34 because bands of the expected size (303 and 313 bp) were detected with C-TPM1/C-TPM2 and C-TPM3/C-TPM4 primer pairs, respectively (data not shown).
DISCUSSION
Recurrent chromosomal translocations play an important role in many human lymphoid tumors and are often responsible for the deregulation of genes involved in the control of cell proliferation.35 A number of recent studies have investigated the (2;5)(p23;q35) translocation, an anomaly associated with ALK-positive ALCL.3-7 The t(2;5) fuses the nucleophosmin gene (NPM) with the ALK receptor tyrosine kinase gene (anaplastic lymphoma kinase) to produce a chimeric protein in which the 116 aminoterminal residues of NPM are linked to the intracytoplasmic portion of ALK.8 The NPM oligomerization domain present in NPM-ALK leads to homodimerization, thus mimicking ligand binding and leading to activation of the ALK catalytic domain.13,14Furthermore, normal NPM contains two potential nuclear-localization signals located between aminoacids 152-157 and 191-197.11,12,36 These sequences are lost from the chimeric NPM-ALK protein, but the oligomerization domain located in the aminoterminal portion of NPM is retained11,13,37 and heterodimers can thus form between NPM-ALK and normal NPM. These translocate to the nucleus, accounting for the characteristic combination of cytoplasmic and nuclear staining observed with the anti-ALK antibodies.14 20
On the basis of the cytogenetic results in patient GEL, which showed the presence of a (1;2)(q25;p23) translocation, we postulated that, in this case, a gene on chromosome 1 was fused to part of the ALKgene and provided a promoter for its expression, comparable to theNPM promoter in cases carrying the (2;5) translocation. However, in contrast to t(2;5)-positive ALCL, ALK protein expression in the case GEL was restricted to the cytoplasm of malignant cells, suggesting that the aberrant fusion protein produced by the t(1;2) lacks motifs responsible for nuclear localization. This finding is in keeping with evidence that a recombinant hybrid ALK protein in which TPR replaces NPM remains confined to the cytoplasm but possesses transforming activity.13 14
Using a PCR-based chromosome-walking method, we demonstrated that the gene involved on chromosome 1 in the case GEL was the TPM3gene, which encodes a nonmuscular tropomyosin protein. Furthermore, we demonstrated TPM3-ALK transcripts in 2 other cases of ALCL exhibiting ALK staining restricted to the cell cytoplasm, suggesting that the t(1;2) is present in a significant minority of cases of ALCL.
Tropomyosins are actin-binding proteins that were first isolated from muscle38 but then found in nonmuscle cells,39where they constitute a major component of cytoskeletal microfilaments.40 In skeletal muscle, they mediate the effect of Ca2+ on the myosin-actin interaction,41 but their role in smooth muscle and nonmuscle tissues is unclear. More than 15 isoforms of tropomyosin have been identified. However, they are encoded by only four genes, and this diversity is generated by tissue-specific alternative processing of RNA. The same gene can therefore encode both muscle and nonmuscle tropomyosin proteins.
The TPM3 gene spans at least 42 kb42 and contains 13 exons. It was first localized at 1q3143 but later assigned to 1q22-23,44 and in the present case (GEL) the breakpoint on chromosome 1 was found at 1q25. The TPM3 gene was found to be fused, in a colonic carcinoma,23 to theNTRK1 tyrosine kinase receptor proto-oncogene and was subsequently found to be involved in an intrachromosomal inversion creating a fusion gene with NTRK1, also referred to asTRK (tropomyosin receptor kinase), in 3 cases of human papillary thyroid carcinomas.25 The replacement of the extracellular portion of NTRK1 by the sequence of theTPM3 gene encoding the 221 aminoterminal residues results in oncogenic activation of NTRK1.25
The breakpoint in the TPM3-NTRK1 oncogene occurs in the 12-kb TPM3 intron located between exons 7 and 8,25and this was also the site of the TPM3 breakpoint in the (1;2) translocation described in this report. Furthermore, the ALKintron involved in this case is also the intron involved in the classical (2;5) translocation.
It has been suggested that translocations can occur in lymphoid tumors through recombinogenic elements such as χ sequences45 or alternating purine-pyrimidine tracts, which form negative supercoiled regions (Z-DNA) making DNA more accessible to the enzymatic machinery.46 We have not found any of these known recombinogenic elements in the regions sequenced around the breakpoint.Alu sequences can also be implicated in recombinations and translocations, such as the t(9;22), where both the Abl andbcr introns contain Alu sequences in an opposite orientation.47 In the case under study (GEL), sequence analysis demonstrated the presence of numerous Alu sequences in the TPM3 intron, the breakpoint itself being located at the beginning of an Alu sequence, but none was found in the disrupted ALK gene intron. Thus, the lack of recombinogenic elements, including Alu sequences near the breakpoint in theALK intron, suggests that novel mechanisms are involved in the (1;2) translocation.
Whatever the mechanism of the (1;2) translocation, the TPM3gene promoter appears to drive the expression of the ALK tyrosine kinase domain, because we have demonstrated that this promoter is active in lymphoid T-cell lines. In addition, tropomyosins are known to form dimeric α-coiled-coil structures41,48 that could induce homodimerization of the chimeric TPM3-ALK protein, a mechanism also involved in the activation of NPM-ALK in the classical (2;5) translocation.13 However, further experiments are needed to assess this hypothesis.
The present cases of ALCL, associated with a novel TPM3-ALKhybrid gene, indicate that partners other than NPM can promote oncogenic expression of the ALK intracytoplasmic domain. Moreover, this study demonstrates the usefulness of anti-ALK antibodies for detecting cases containing possible new translocations in ALCL, because when staining is restricted to the cell cytoplasm and not present in the nucleus, a partner other than NPM is likely to be present.16,17 19
ACKNOWLEDGMENT
The authors gratefully acknowledge Prof David Y. Mason for helpful suggestions and critical reading of the manuscript.
Supported by the “Projet Hospitalier de Recherche Clinique (PHRC98),” “Ligue Nationale Contre le Cancer,” the “Groupe d’Etude des Lymphomes de l’Adulte (GELA),” and the Leukemia Research Fund, UK.
The publication costs of this article were defrayed in part by page charge payment. This article must therefore be hereby marked “advertisement” in accordance with 18 U.S.C. section 1734 solely to indicate this fact.
REFERENCES
Author notes
Address reprint requests to Georges Delsol, MD, Laboratoire d’Anatomie Pathologique, CHU Purpan, Place du Dr Baylac, 31059, Toulouse Cedex, France.


This feature is available to Subscribers Only
Sign In or Create an Account Close Modal