Abstract
Abstract 2428
Aberrant DNA methylation repeatedly has been implicated in cancer development. DNA methyltransferase (DNMT) 3A, which mediates de novo DNA methylation, was found to be mutated in 20% of patients with acute myeloid leukemia and 10% of patients with myelodysplastic syndrome. Recently, mutations associated with myeloid malignancies such as DNMT3A and FLT3 have also been uncovered in patients with early T-cell precursor lymphoblastic leukemia (ETP-ALL) (Neumann et al., 2012; Van Vlierberghe et al., 2011; Zaremba et al., 2012). ETP-ALL is a type of very high-risk ALL associated with myeloid/stem cell gene expression signature and myeloid markers. We have demonstrated that Dnmt3a deletion in mouse causes increased self-renewal of hematopoietic stem cells and an impairment of differentiation (Challen et al., 2011). Dnmt3a loss also produces aberrant methylation associated with oncogenes and tumor suppressor genes. Yet, whether aberrant DNA methylation can drive leukemia remains unknown. As Dnmt3a deletion alone was insufficient for malignancy, secondary mutations are likely necessary for leukemic transformation. Because FLT3 internal tandem duplication (ITD) frequently co-exist with DNMT3A mutations in acute leukemias, we hypothesized that Dnmt3a-loss may cooperate with FLT3-ITD to promote leukemic transformation; and we established a mouse model to test this.
Deletion of conditional Dnmt3a with Mx1-cre was induced by injections of pIpC. Subsequently, bone marrow from Dnmt3a-deleted (Dnmt3aKO) donor mice was transduced with MSCV-FLT3-ITD-GFP retrovirus or MSCV-GFP control and transplanted into lethally irradiated recipients. The mice were monitored monthly for development of malignancies by complete blood count and peripheral blood analysis by flow cytometry and followed for disease latency. Moribund mice were sacrificed and analyzed with peripheral blood smears, histology, and immunophenotyping.
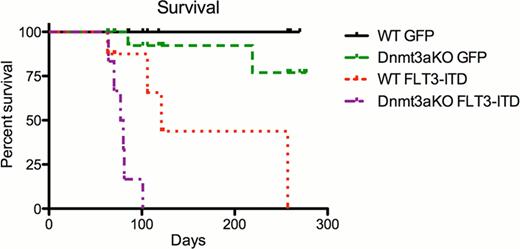
Dnmt3a deletion with overexpression of FLT3-ITD caused rapid onset T-ALL in 6/8 mice (n=6) with a median latency of 78 days compared to 121 days in WT mice (n=4) overexpressing FLT3-ITD (p<0.0001 Log-rank Mantel-Cox Test) (See figure). Mice from both groups exhibited leukocytosis, splenomegaly, and thymomegaly with high GFP expression detected by FACS. Even after we transduced bone marrow cells enriched for myeloid progenitor and stem cells, Dnmt3a deletion again accelerated T-ALL with median survival of 89 days (n=9) versus 110 days in WT-FLT3-ITD (n=10) mice. T-ALL was observed in 2/4 WT-FLT3-ITD mice and 5/6 Dnmt3aKO-FLT3-ITD mice analyzed (p<0.0001 Log-rank Mantel-Cox Test). By flow cytometry, two distinct types of T-ALL were observed in the bone marrow of Dnmt3a deleted leukemic mice: one was characterized by a double positive population (DP) of CD4+CD8+ lympoblasts (1/6) and another early immature T-cell-like type of CD4-CD8-CD44+CD25-CD11bloCD117+ lymphoblasts (4/6). Gene expression analysis by RT-PCR in the early immature T-ALL showed downregulation of Notch-pathway genes (such as Notch1, Notch 3, Deltex, Hes1) and upregulation of stem cell-associated genes Lyl1 and Scl1, suggesting an ETP-like T-ALL. The ETP-like ALL phenotype has not been seen in WT mice overexpressing FLT3-ITD. The opposite gene expression pattern was seen in the DP population with upregulation of Notch-pathway genes. Furthermore, the DP leukemia was transplantable to secondary recipients within 2 weeks. Whether ETP-like ALL can be transplanted is still under investigation. We are also currently studying the changes in global CpG methylation among the leukemias that have Dnmt3a loss, FLT3-ITD overexpression, and control and also anticipate data from transcriptome analysis by RNA-Seq.
These data suggest that stem or progenitor bone marrow cells primed by early loss of Dnmt3a are transformed into DP T-ALL and ETP-like ALL fueled by the overexpression of the oncogene FLT3-ITD. The ETP-like ALL phenotype has not been seen previously in WT mice overexpressing FLT3-ITD, suggesting that Dnmt3a ablation is required. The Dnmt3a-deleted-FLT3-ITD mice with T-ALL is, to our knowledge, the first animal model of human immature T-cell leukemia. This model can enhance our understanding of the pathogenesis of ETP-like ALL with respect to aberrant DNA methylation and will serve as a powerful tool to test novel therapeutic strategies.
No relevant conflicts of interest to declare.
Author notes
Asterisk with author names denotes non-ASH members.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal