Abstract
Previous studies have established that factor VII gene (F7) polymorphisms (5′F7 and R353Q) contribute about one-third of factor VII (FVII) level variation in plasma. However, F7 genotyping in patients with cardiovascular disease has produced conflicting results. Population and expression studies were used to investigate the role of intron 7 (IVS7 ) polymorphisms, including repeat and sequence variations, in controlling activated FVII (FVIIa) and antigen (FVIIag) levels. Genotype–phenotype studies performed in 438 Italian subjects suggested a positive relation between the IVS7 repeat number and FVII levels. The lowest values were associated with theIVS7 + 7G allele. The screening of 52 patients with mild FVII deficiency showed an 8-fold increase in frequency (8%) of this allele, and among heterozygotes for identical mutations, lower FVII levels were observed in the IVS7 + 7G carriers. This frequent genetic component participates in the phenotypic heterogeneity of FVII deficiency. The evaluation of the individual contribution of polymorphisms was assisted by the expression of each IVS7variant, as a minigene, in eukaryotic cells. The novel quantitative analysis revealed that higher numbers of repeats were associated with higher mRNA expression levels and that the IVS7 + 7Gallele, previously defined as a functionally silent polymorphism, was responsible for the lowest relative mRNA expression. Taken together, these findings indicate that the IVS7 polymorphisms contribute to the plasmatic variance of FVII levels via differential efficiency of mRNA splicing. These studies provide further elements to understand the control of FVII levels, which could be of importance to ensure the hemostatic balance under pathologic conditions.
Factor VII (FVII) is a vitamin K–dependent serine protease glycoprotein with a pivotal role in blood coagulation.1 In association with tissue factor, an integral membrane protein exposed in the vascular lumen following a lesion, activated FVII (FVIIa) initiates coagulation,2,3which makes its levels of importance for the hemostatic balance in normal and pathologic conditions. Decreased FVII levels are associated with a variable bleeding tendency,4 and the FVII gene (F7) knock-out experiments in mice demonstrated that death following birth was due to hemorrhage from normal blood vessels.5 The association between increased FVII levels and cardiovascular disease is strongly debated,6-18 but high levels may contribute, at least in the presence of additional risk factors, to myocardial infarction and should be taken into account when assessing cardiovascular risk.19 High FVII levels might disproportionately enhance the coagulation cascade at the time of plaque rupture, which could explain the apparently differential association with fatal and nonfatal events.6,7,10 19
FVII plasma levels are determined by environmental20,21 and genetic factors22-24 and by their interplay.25-27 A strong contribution of the F7genotype to FVII levels has been demonstrated: different F7genotypes demonstrated up to 5-fold differences in mean FVIIa levels.28 Tightly linked polymorphisms in the 5′ regulatory region are associated with FVII levels and differential promoter activity in vitro.29,30 Regulation by polymorphisms of the low FVII concentration (500 ng/mL) in plasma, the lowest among the vitamin K–dependent coagulation proteins, might be favored by the weak F7 gene promoter, which lacks the TATA and CAAT boxes and shows a low affinity for ubiquitous transcription factors.31-33 Moreover, the R353Q substitution in the catalytic domain, tightly associated with the decanucleotide insertion and with the −401T allele in the promoter region,30 reduces FVII levels by decreasing its secretion.34 An additional polymorphism, located in intron 7 (IVS7 ) of the F7 gene, is characterized35-37 by the presence of a variable number of 37–base pair (bp) repeats. In the first repeat, which contains the IVS7donor splice site, sequence variations have also been characterized.38 Recent findings have suggested a relationship of certain IVS7 genotypes with myocardial infarction,39 and repeat polymorphisms have been associated with human disorders.40F7 gene repeat and sequence variations might play an additional role in modulating FVII levels in plasma.
Understanding the functional role of the different FVII genetic components might be essential to explain previous conflicting results. Using genotype–phenotype studies, the goal of the present study was to define IVS7 polymorphisms in the control of plasma FVII levels. A control population and a cohort of patients with mild FVII deficiency, potentially useful to evidence the contribution of polymorphic alleles, were investigated. To dissect at the molecular level the genetic components indicated by population studies, the in vitro expression in eukaryotic cells of the different polymorphic forms was performed. The use of minigenes, previously exploited by us38 for the study of splicing mutations in FVII deficiency, was extended and modified by quantitative analysis at the mRNA level.
Patients, materials, and methods
Patients and controls
We studied 52 patients with mild FVII deficiency (FVIIc) (range, 17%-55%; mean, 38.7%). We had previously characterized the FVII coagulation phenotypes and the underlying molecular lesions in most of the patients.38 41 Four asymptomatic patients (pedigrees PFVII29-32, Table 1) had not been previously reported. Patients with severe FVII deficiency cannot provide information about the additional contribution of polymorphic alleles to the FVII levels, and as a result, they were excluded from this study. We also studied 438 subjects who were enrolled as controls in an ongoing vascular disease study in Northern Italy.
Coagulation studies
Blood sampling from control subjects was performed essentially as previously described.28 FVIIa was assayed with a kit (STACLOT VIIa-rTF; Diagnostica Stago, Asnières-Sur-Seine, France) on a Behring Coagulation Timer (Behring Diagnostic GmbH, Marburg, Germany). Values were expressed as 30 mU/mL = 1 ng of FVIIa. Recombinant FVIIa (Novo-Nordisk, Bagsvd, Denmark) supplied with the kit was used as a standard. In 102 control subjects the FVIIag was assayed using an enzyme immunoassay as previously described.24Plasma samples suitable for FVIIa assay were not available for the FVII-deficient patients. In the newly reported patients, FVIIc was assayed by a 1-stage method.
Statistical analysis
Distributions of continuous variables in groups were expressed as the mean ± SD. FVIIa levels of individuals with differentF7 genotypes were compared by analysis of variance (ANOVA) using the Tukey test, a post-hoc comparison of the means. ANOVAs were carried out on values of FVIIa and FVIIag after logarithmic transformation. The presence of the Hardy-Weinberg equilibrium for F7 genotype frequencies was analyzed by the χ2 test. All computations were performed by using the SPSS 7.5 statistical package (SPSS, Chicago, IL).
Genomic studies
Normal subjects and patients were characterized for the5′F7, IVS7, and R353Q polymorphisms in the F7 gene as previously described.22,42,43 Among the strongly linked F7 promoter polymorphisms, the deletion (A1)/easily detectable decanucleotide insertion (A2) (A1/A2) marker was chosen for this study. The IVS7 allelic forms defined in the original reports36,43 as c, b, a, and d are now reported asH5, H6, H7, and H8, respectively, depending on the presence of 5, 6, 7, and 8 repeats. The9726 + 7A→G transition, which abolishes an RsaI restriction site,43 is reported as 7G. Direct sequencing was used to confirm the identity of this mutation. The search for mutations in the F7 gene of patients not previously characterized was carried out by polymerase chain reaction (PCR) amplification and direct sequencing of coding regions and intron–exon junctions, as previously reported.41
Cloning
Genomic fragments containing different polymorphic alleles were obtained by PCR amplification using the Expand high-fidelity PCR system (Boehringer Mannheim, Mannheim, Germany). The genomic region spanning nucleotides 9568-10 908 (3′ of exon 7, intron 7, and 5′ of exon 8) and containing 5 or 8 repeats was cloned in the pUC18 vector and then subcloned in the pcDNA3 vector as previously described for alleles H6, H7, IVS7 + 5A, andIVS7 + 7G.38 The complete insert of each construct was sequenced by primers located in exon 7 (CCCTGATC AACACCATCTGG [nucleotides 9642-9663]) and intron 7 (GATGTCTGTCTG TCTGTGGA [nucleotides 10 009-9990]; TCCACAGACAGACAGACATC [nucleotides 9990-10 009]; and GAGGTGGCAGGTGGTGGAAA [nucleotides 10 495-10 514]). Plasmids containing the different allelic forms are reported as pIVS7H5, pIVS7H6, pIVS7H7, pIVS7H8, pIVS7 + 5A (9726 + 5A38), and pIVS7 + 7G (9726 + 7G38).
Cell culture and transfection
We transfected 7 × 105 baby hamster kidney (BHK) cells, grown as described,38 into a 10-cm petri dish using the calcium-phosphate method or a nonliposomal blend of lipids (FuGENE6 Transfection Reagent; Boehringer Mannheim). For each transfection, 6 μL FuGENE6 was incubated with 94 μL serum-free medium for 5 minutes at room temperature, added dropwise to the DNA, and then reincubated for 15 minutes at room temperature. The reagent/DNA mixture was added dropwise to the cell medium and then incubated at 37°C with 5% carbon dioxide (CO2).
A constant amount of 3 μg pIVS7 + 5A was mixed with decreasing amounts of each construct (pIVS7H5, pIVS7H6, pIVS7H7, pIVS7H8, and pIVS7 + 7G) at DNA concentration ratios of 1:1, 1:0.3, 1:0.15, and 1:0.1 and transfected into BHK cells. The optimal ratios, 1:0.15 and 1:0.1, were exploited for subsequent cotransfections.
RNA studies
Twenty-two hours after transfection, total RNA was extracted using RNAfast (Molecular Systems, San Diego, CA) and evaluated by reverse transcriptase–PCR (RT-PCR) using 40 units avian myeloblastosis virus RT and 10 N reverse oligonucleotide (CAGCGCGATGTCGTGGTT, exon 8 [nucleotides 10 652-10 635]). We amplified one-tenth of the RT reaction mixture with forward oligonucleotides (ACCCTGATCAACACCATCTGG, exon 7 [nucleotides 9642-9663]) and reverse oligonucleotides.
Normal and mutated cDNA fragments were separated on 2.5% agarose or 10% polyacrylamide (PAA) gels and evaluated after ethidium bromide staining. Quantitative analysis was performed by densitometric scanning (GS-700 Imagin Densitometer; Bio-Rad, Hercules, CA) of bands, and the intensity ratio between each investigated construct and pIVS7 + 5A was calculated.
Results
Population studies
The allelic forms of IVS7 and the 5′F7insertion/deletion polymorphisms were studied in 438 subjects characterized for plasma FVIIa levels. Out of 438 patients, 102 were also characterized for FVIIag levels, and association between variables was observed (correlation coefficient FVIIa/FVIIag, 0.69). The observed allelic and genotype frequencies were similar to those previously reported in the Italian population28 and were within the Hardy-Weinberg equilibrium.
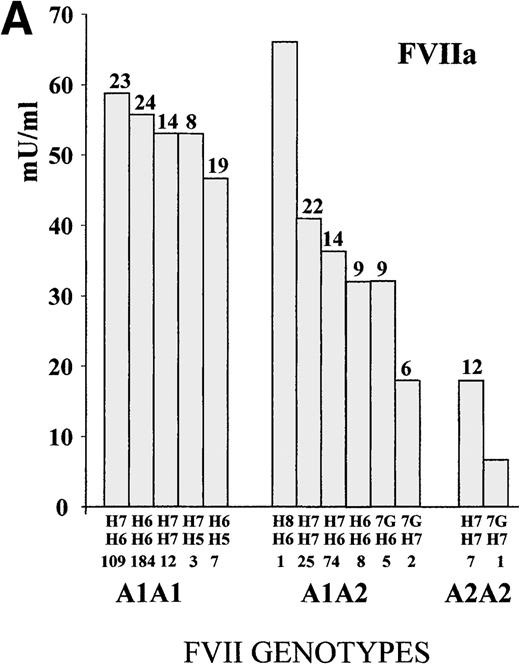
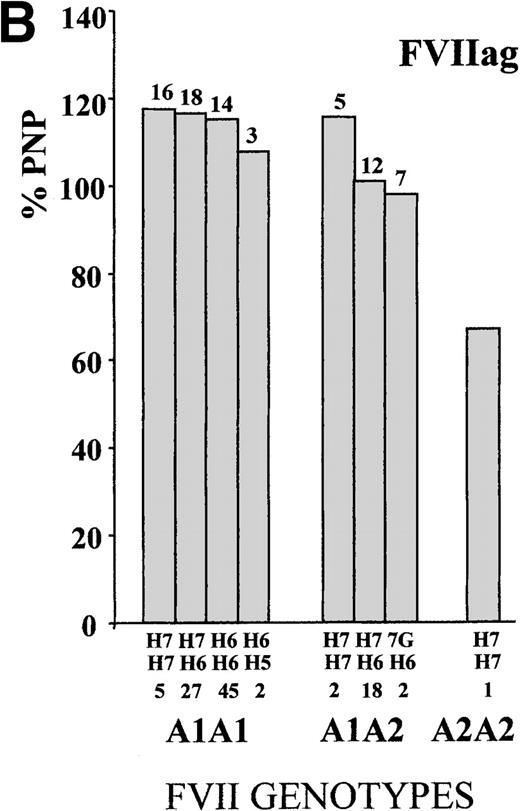
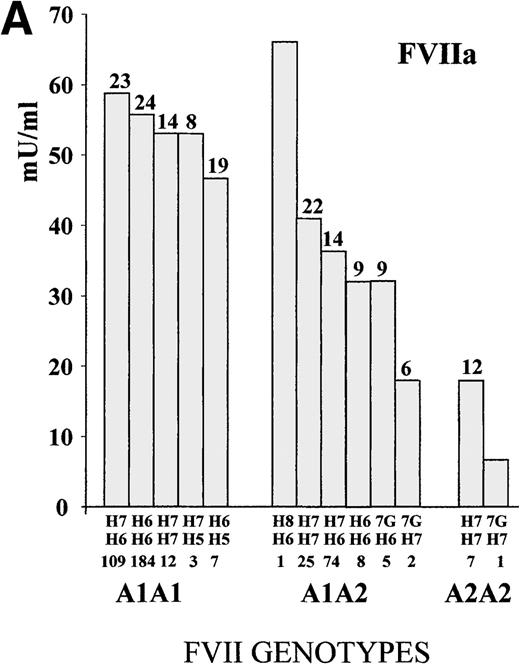
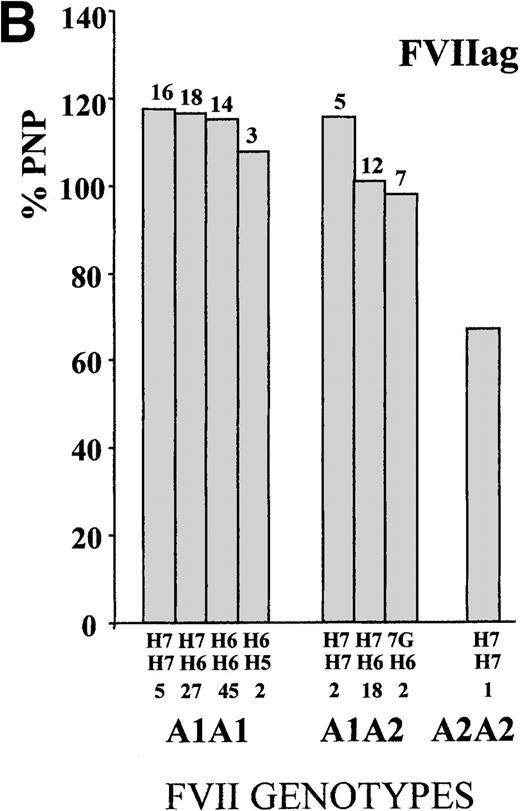
To define the A1/A2 and IVS7 genotype–phenotype relationship, FVII values were studied in subjects grouped by combined genotypes (Figure1). The ANOVA for FVIIa and FVIIag resulted in highly significant differences (FVIIa, P < .001, and FVIIag, P = .007; F values, 14.4 and 3.1, respectively). The heterozygous or homozygous condition for the A2 allele (5′F7 insertion) was associated with a FVIIa mean reduction of 34.9% and 68.2%, respectively, which confirmed previously obtained results.28 Mean FVIIag levels showed a reduction of 13.2% in the A2 carriers. All significant differences (Tukey post-hoc comparison of the means) were found between genotypes including a different number of A2 alleles. Frequent additional small differences were observed within A1A1 orA1A2 genotypes grouped by IVS7 polymorphisms. For example, the FVIIa levels associated with the A1A1/H6H7genotype were 5.3% higher than those found in subjects characterized by the genotype A1A1/H6H6, and genotype A1A2/H7H7 was associated with a 12.5% increase in FVIIa values as compared with genotype A1A2/H6H7.
FVII levels and genotypes.
Mean (A) FVIIa and (B) FVIIag levels in control subjects are grouped by5′F7 and IVS7 genotypes. The 5′F7 A2 allele contains the decanucleotide insertion.7G is the result of the9726 + 7A→G transition. The number of subjects is reported below the IVS7 genotypes, and the SD is shown above each column. PNP indicates pooled normal plasma.
FVII levels and genotypes.
Mean (A) FVIIa and (B) FVIIag levels in control subjects are grouped by5′F7 and IVS7 genotypes. The 5′F7 A2 allele contains the decanucleotide insertion.7G is the result of the9726 + 7A→G transition. The number of subjects is reported below the IVS7 genotypes, and the SD is shown above each column. PNP indicates pooled normal plasma.
The evaluation of infrequent genotypes provided additional information. The IVS7 allele H5 was present in 10 (2.3%) subjects; the H8 allele in 1 (0.2%) subject; and the 9726 + 7Gallele (7G) in 8 (1.8%) subjects. Taken together, infrequentIVS7 alleles were present in more than 4% (19/438) of the subjects. Only A1 homozygote subjects carried theH5 allelic form, and FVIIa values determined in subjects with the A1A1/H5 genotype (mean FVIIa, 48.6 mU/mL ± 16.4 SD) were 14.3% lower than those obtained in the other A1A1subjects (n = 305; mean FVIIa, 56.7 mU/mL ± 23.5 SD). Among theA1A2 heterozygotes, carriers of the 7G allele (n = 7; mean FVIIa, 28.7 mU/mL ± 9.9 SD) showed a mean reduction of 23% in FVIIa levels as compared with the other A1A2 subjects (n = 108; mean FVIIa, 37.3 mU/mL ± 16.1 SD). A similar contribution of the 7G allele in lowering FVIIa levels was observed among the A2 homozygotes. When all carriers of the7G alleles (n = 8) were grouped and compared with the otherA2 heterozygote or homozygote subjects (n = 115), they differed significantly (P < .05). The only subject carrying an H8 allele showed a high FVIIa level.
Although FVIIag differences among genotype groups (Figure 1) were smaller than those observed for FVIIa, genotypes ranked by FVIIag or FVIIa displayed a very similar pattern. In particular, carriers of the H5 or 7G alleles had the lowest levels in both distributions.
Patient studies
The contribution of polymorphic alleles to FVII levels was further investigated in 52 patients selected for mild FVII deficiency. Most of them were already characterized for heterozygosity of molecular lesions affecting the F7 gene.38,41 The rationale for this choice was the assumption that the presence of molecular lesions impairing the expression of 1 allele would facilitate the detection of the contribution of polymorphisms linked to the functional allele. Using 5′F7 and IVS7 polymorphism analysis, frequencies of the 7G allele (8/52, 8%) and the A2allele (39.4%) were increased 8-fold and 2-fold, respectively, in patients compared with controls. The finding of 5 A2homozygotes41 among the 7G carriers enabled us to establish an almost complete linkage disequilibrium between these alleles, which complicates the evaluation of the individual contribution of the 7G allele.
The coagulation phenotypes of patients (Table 1) heterozygous for identical FVII missense mutations were compared. The observation of lower FVII levels in carriers of the 7G allele (PFVII29 and PFVII31) further suggested the functional effect of this allele. In the 4 newly reported heterozygous patients (Table 1), sequencing of theF7 gene revealed the presence of the C310F andM298I mutations, which have previously been described37 41 as responsible for CRM+ FVII deficiency in Italian patients.
Expression studies
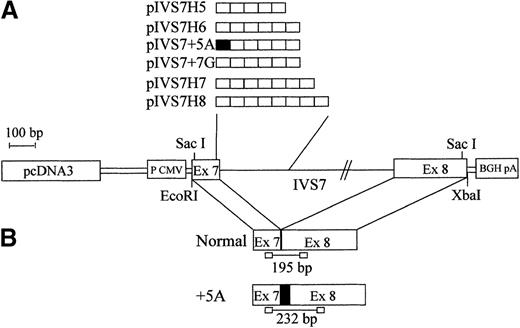
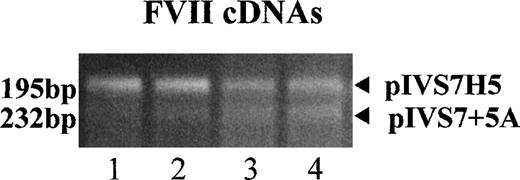
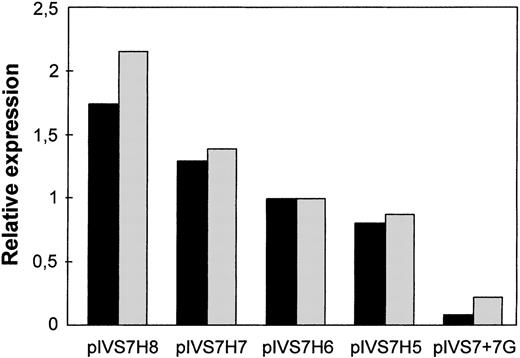
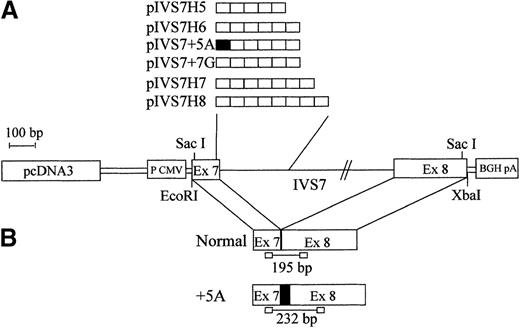
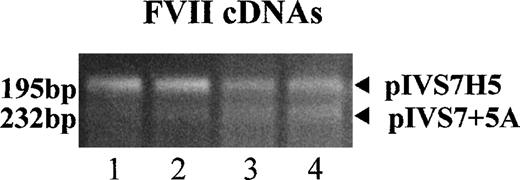
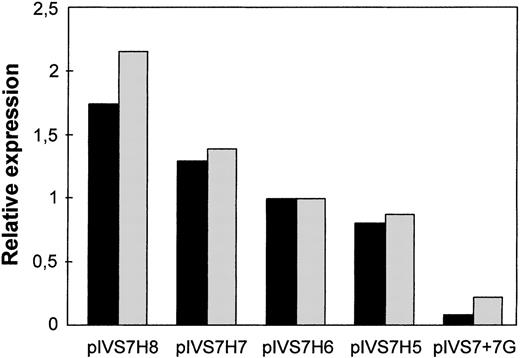
To study the expression of the IVS7 alleles at the messenger RNA (mRNA) level, we made minigene constructs containing the various polymorphic forms (Figure 2), and the presence of the correct sequences was confirmed by sequencing. RT-PCR evaluation of mRNA produced by constructs pIVS7H5, pIVS7H6, pIVS7H7, pIVS7H8, and pIVS7 + 7G and transfected into mammalian cells showed that all of the constructs had the presence of cDNAs with the normal expected size, 195 bp (Figure 2 and Figure3). pIVS7 + 5A, bearing a mutation that impairs splicing at the IVS7 donor splice site,38showed only an abnormal transcript of 232 bp (Figure 2 and Figure 3), which was chosen as an internal control for quantitative analysis. Competition between constructs was demonstrated in cotransfection experiments obtained with constant amounts of pIVS7 + 5A and decreasing amounts of pIVS7H5 (Figure 3). A quantitative analysis of transcripts was performed by the densitometric scanning of bands, and the relative expression of each construct was estimated. The expression level of pIVS7H6, bearing the most frequent repeat number in the general population, was considered as a reference point (ratio = 1) (Figure 4). Data obtained from repeated transfection experiments (Figure 4) indicated a parallel decrease of the IVS7 repeat number and mRNA relative expression. In these quantitative studies, construct pIVS7 + 7G resulted in the lowest expression level.
Schematic diagram of the F7 gene and mRNA regions expressed in mammalian cells.
(A) pcDNA3 plasmid (pIVS7H5, pIVS7H6, pIVS7H7, pIVS7H8, pIVS7 + 5A, and pIVS7 + 7G) containing the different allelic forms ordered by repeat number. (B) Spliced transcription products from plasmids and the size of RT-PCR–amplified fragments. pIVS7 + 5A, used as an internal control, produces the altered transcript (+5A), which contains an additional 37-bp sequence (indicated by filled square).
Schematic diagram of the F7 gene and mRNA regions expressed in mammalian cells.
(A) pcDNA3 plasmid (pIVS7H5, pIVS7H6, pIVS7H7, pIVS7H8, pIVS7 + 5A, and pIVS7 + 7G) containing the different allelic forms ordered by repeat number. (B) Spliced transcription products from plasmids and the size of RT-PCR–amplified fragments. pIVS7 + 5A, used as an internal control, produces the altered transcript (+5A), which contains an additional 37-bp sequence (indicated by filled square).
mRNA expression studies with RT-PCR–amplified fragments of FVII mRNA expressed in BHK cells cotransfected with pIVS7H5 and pIVS7 + 5A (internal control).
The following pIVS7 + 5A/pIVS7H5 DNA concentration ratios were used in lanes 1-4: 1:1, 1:0.3, 1:0.15, and 1:0.1, respectively.
mRNA expression studies with RT-PCR–amplified fragments of FVII mRNA expressed in BHK cells cotransfected with pIVS7H5 and pIVS7 + 5A (internal control).
The following pIVS7 + 5A/pIVS7H5 DNA concentration ratios were used in lanes 1-4: 1:1, 1:0.3, 1:0.15, and 1:0.1, respectively.
Relative expression at the mRNA level of the differentIVS7 allelic forms.
Cotransfections with pIVS7 + 5A and investigated constructs at concentration ratios of 1:0.1 (black columns) and 1:0.15 (gray columns) were carried out.
Relative expression at the mRNA level of the differentIVS7 allelic forms.
Cotransfections with pIVS7 + 5A and investigated constructs at concentration ratios of 1:0.1 (black columns) and 1:0.15 (gray columns) were carried out.
Discussion
The FVII genotype/phenotype relationship confirmed a major role of the 5′F7 and R353Q polymorphisms in lowering FVII levels28 and suggested a parallel decrease of theIVS7 repeat number and the FVII levels. Differences associated with repeat numbers were more pronounced in the A1A2 subjects than in the A1 homozygote subjects, which could suggest that the influence of the IVS7 polymorphisms on FVII levels might be partially masked by the higher promoter activity associated with theA1 allele.29,30 As observed in a previous population study,28 FVIIa differences associated withF7 genotypes (5′F7 and IVS7polymorphisms) were larger than those obtained for the antigen levels. FVIIa is indeed the first and longest living active protease in the coagulation pathway. Its level variations may modify the hemostatic balance both in bleeding and prothrombotic conditions, and the amplified differences in FVIIa values, predicted by genetic components, could have interesting clinical and biologic implications. A correlation between repeat number and plasma protein levels has also been reported for a polymorphism in intron 4 of the endothelial constitutive nitric oxide synthase gene.44
The estimate of the independent contribution of IVS7polymorphisms to FVII variance is complicated by several genetic factors, among them the low frequency of 7G, H5, andH8 alleles, which is always found in the heterozygous condition, and the linkage between the 7G and A2alleles. To substantiate the data obtained in the control population and in FVII-deficient patients, we expressed all the IVS7variants (H5, H6, H7, H8, and7G) into eukaryotic cells. Due to the intronic localization of these polymorphisms, the study was conducted at the mRNA level. Although this approach does not enable the evaluation at the protein level, it is conceivable that the amount of protein reflects the mRNA levels. In fact, the mature FVII mRNAs spliced from the different alleles are identical in sequence because of the absence of the polymorphic intronic region, thus excluding in principle any difference in mRNA stability or translation efficiency. The use of minigene constructs, lacking the 5′F7 and R353Qpolymorphic regions, enabled us to evaluate only the influence of theIVS7 polymorphisms. Moreover, pIVS7 + 5A, used as an internal control in double transfections, provided an original model for quantitative analysis of the expressed products.
The relative mRNA expression levels positively correlated with the number of intronic repeats in different sets of experiments, and this relation was similar to that observed for FVII levels in the population study (Figure 4). However, differences in FVII levels among genotype groups were smaller than those indicated by the in vitro mRNA expression studies. This amplification, observed in other in vitro expression studies,45 might depend on the use of a cell line that is neither human nor hepatic and the use of minigene constructs, which purposely contain only a portion of the F7gene. Thus, further control levels in the RNA processing pathway, which are present in sequence elements of the complete gene,46-48would be excluded. Moreover, in vitro studies do not provide additional physiologic components, which have been demonstrated to influence FVII levels in vivo.49
The effect of the 7G allele in lowering FVII levels was detectable in the 2 independently selected populations, namely in the control population and in the cohort of patients with mild FVII deficiency. Particularly, among both A1A2heterozygous subjects and patients heterozygous for identical missense mutations, 7G carriers had the lowest FVIIa levels. These findings, as well as the high frequency of the 7G allele (approximately 8%) among patients with mild FVII deficiency, indicate that this allele contributes to decreased FVII levels. The in vitro results, which showed the lowest relative mRNA expression for construct pIVS7 + 7G, paralleled those obtained in vivo. Previous experiments, which did not find any qualitative difference between mRNA transcripts from this construct and from the normal control, led us to define the transition 9726 + 7A→G as a functionally silent polymorphism.38 The novel quantitative approach used in the present study reveals the functional role of this mutation, which is consistent with the in vivo findings. For the evaluation of expression studies, this observation points out the importance of appropriately designed control constructs.
These data characterize an additional genetic determinant of FVII levels, which is frequently found in FVII deficiency. In particular, the 7G allele lowers FVII levels and, using standard coagulation screenings, favors the detection of the heterozygous FVII deficiencies as well as the homozygous condition for the A2and 353Q (M2) alleles. The in vitro results, which paralleled the in vivo studies, strengthen the functional role of the variable number of IVS7 tandem repeats in modulating FVII expression. Although the repeated IVS7 sequences could affect different steps of gene expression, a differential splicing efficiency is suggested by the presence in each repeat of 2 G triplets, which may act as an intron splicing enhancer.50,51 The increase of the G triplet number would increase splicing factor binding, thus explaining the positive relationship between the repeat number and mRNA expression. The A→G transition at position +7 does not alter either the splicing consensus sequence52 or the GGG content. It should modify a different molecular interaction that is important for the splicing process. A hemophilic mutation affecting position +7 in the FIX gene53further highlights the functional role of this nucleotide.
The influence of IVS7 polymorphisms on FVII levels could play an important role in ensuring the hemostatic balance under pathologic conditions, as suggested by the recent finding of the association between certain IVS7 genotypes and myocardial infarction.39 When FVII levels associated with the most frequent genotype groups are compared with those obtained by Iacoviello et al,39 a similar relationship between repeat number and FVII levels is detectable. Particularly genotypes RRH6H6 andRQH6H6 showed39 lower FVII levels than genotypes RRH6H7 and RQH6H7, respectively. However, for these combined genotypes, the increase in FVII levels is not associated with a parallel increase of the relative risk of myocardial infarction.39 In the absence of specific screening for the7G allele in patients,39 a more informative comparison between the present data and that previously reported is not feasible.
Taken together, the present findings indicate the independent contribution of the IVS7 polymorphisms to the plasmatic variance of FVII levels, mediated by differential mRNA splicing efficiency. These studies cast light on the complex physiologic control of the FVII level and help the molecular dissection of the phenotypic heterogeneity of FVII deficiency.
Supported in part by a grant from the CNR Biotech Target Project (no. 9900295PF49); a grant from Telethon Italy (no. E.0675); and Cariverona-Progetto Sanità, Verona, Italy. Università-Azienda Ospedaliera, Ferrara, Italy, provided a fellowship to P.F. (Ricerca Biomedica Project).
Submitted August 5, 1999; accepted January 27, 1999.
Reprints:Francesco Bernardi, Dipartimento di Biochimica e Biologia Molecolare, Università di Ferrara, Via L. Borsari 46, 44100 Ferrara, Italy; e-mail: ber@dns.unife.it.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 U.S.C. section 1734.