Follicular lymphoma (FL) originates from a single B cell that has rearranged one copy of its BCL2 gene on chromosome 18 to the Ig locus on chromosome 14 and in addition has acquired a mutation in a histone modifying gene such as CREBBP or KMTD2. By the time the disease is diagnosed the progeny of this original cell harbors additional mutations and is usually found at multiple lymphoid sites throughout the body. At each of these sites the malignant cells are accompanied by a rich network of follicular dendritic cells, T cells and other immune cells. This tumor microenvironment (TME) is clearly an important feature of the biology of FL and can impact the clinical behavior of the disease (Dave et al., NEJM, 2004). It remains unknown whether tumor clonal heterogeneity and the composition of the TME differ between various lymphoma sites within the same patient. Single cell RNA sequencing facilitates a detailed and unbiased view of both the tumor clone and the complex TME. To profile the TME and explore FL tumor evolution, we obtained fine needle aspirates (FNAs) at 2 different sites in the body and peripheral blood specimens all on the same day and subjected these samples to single cell RNA sequencing and immune repertoire analysis.
These biopsies were taken prior to therapy from patients entering immunotherapy clinical trials (NCT02927964, NCT03410901). Single cell RNA sequencing of FNA and blood samples was performed using the 10X Genomics platform to an average targeted depth of 50,000 reads/cell. We have prepared sequencing libraries from 15 tumor FNA and peripheral blood samples from 5 patients thus far. Typically, 3,000-10,000 cells have been sequenced per sample, with excellent sequencing quality metrics.
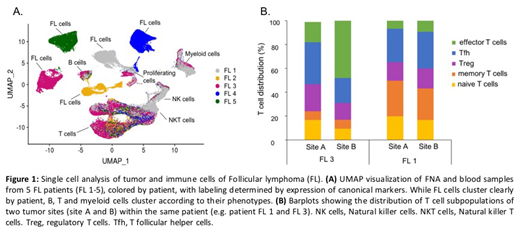
By applying Uniform Manifold Approximation and Projection (UMAP), a dimensionality reduction algorithm, we found the TME of these FL patients to be richly populated by many phenotypically discrete non-malignant cells, including many subpopulations of T cells, B-cells, myeloid cells, NK cells and dendritic cells. Evaluating the combined dataset containing all tumor samples for all 5 patients, we found that malignant B cells from different patients clearly clustered apart from each other, a feature not dependent on immunoglobulin clonality or HLA type. Each patient's tumor population contained 3-5 distinct subpopulations, presumably a result of multiclonal tumor evolution. Nonetheless, we were able to define several malignant B-cell sub-phenotypes common to all patients. Intriguingly, compared to malignant B cells, infiltrating non-malignant B cells showed higher MHC I expression, activation markers, and an enrichment in interferon-induced genes. Of note, we could also detect circulating tumor cells in peripheral blood samples of several patients, and these exhibited a distinct gene expression profile compared to their counterparts within lymph nodes. Analysis of the diverse T cell subpopulations within tumors revealed distinct functional states. For example, in regulatory and T follicular helper cells, we identified activated clusters (CD27, BATF, TNFRSF4) and putative resting clusters (SELL, KLF2, IL7R), while effector T cells resided in separate cytotoxic (GZMA, GZMB, GNLY) and exhausted (TIGIT, CXCL13, LAG3) clusters.
Tumor B cell gene expression and composition of the TME from site to site within the same patient were similar in some cases and remarkably divergent in others. For example, we detected a significant upregulation of interferon signaling pathways in the tumor B cells and an enrichment of effector T cells in only one of the two sites within one patient. Analysis of B cell and T cell antigen receptor sequences to evaluate tumor subclonality and TCR clonotype diversity are ongoing.
To the best of our knowledge, this is the first study to compare different sites of FL in the same patients at the single cell level. Our analyses characterize inter- and intra-patient heterogeneity in malignant and immune cell subsets and provide a baseline for eventual comparison of alterations occurring over time as these patients receive experimental immunotherapy interventions.
Levy:XCella: Membership on an entity's Board of Directors or advisory committees; Immunocore: Membership on an entity's Board of Directors or advisory committees; Walking Fish: Membership on an entity's Board of Directors or advisory committees; Five Prime: Membership on an entity's Board of Directors or advisory committees; Corvus: Membership on an entity's Board of Directors or advisory committees; Quadriga: Membership on an entity's Board of Directors or advisory committees; BeiGene: Membership on an entity's Board of Directors or advisory committees; GigaGen: Membership on an entity's Board of Directors or advisory committees; Teneobio: Membership on an entity's Board of Directors or advisory committees; Sutro: Membership on an entity's Board of Directors or advisory committees; Checkmate: Membership on an entity's Board of Directors or advisory committees; Nurix: Membership on an entity's Board of Directors or advisory committees; Dragonfly: Membership on an entity's Board of Directors or advisory committees; Innate Pharma: Membership on an entity's Board of Directors or advisory committees; Abpro: Membership on an entity's Board of Directors or advisory committees; Apexigen: Membership on an entity's Board of Directors or advisory committees; Nohla: Membership on an entity's Board of Directors or advisory committees; Spotlight: Membership on an entity's Board of Directors or advisory committees; 47 Inc: Membership on an entity's Board of Directors or advisory committees.
Author notes
Asterisk with author names denotes non-ASH members.